Abstract
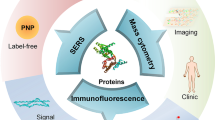
Proteins have been considered key building blocks of life. In particular, the protein content of an organism and a cell offers significant information for the in-depth understanding of the disease and biological processes. Single-molecule protein detection/sequencing tools will revolutionize clinical (proteomics) research, offering ultrasensitivity for low-abundance biomarker (protein) detection, which is important for the realization of early-stage disease diagnosis and single-cell proteomics. This improved detection/measurement capability delivers new sets of techniques to explore new frontiers and address important challenges in various interdisciplinary areas including nanostructured materials, molecular medicine, molecular biology, and chemistry. Importantly, fluorescence-based methods have emerged as indispensable tools for single protein detection/sequencing studies, providing a higher signal-to-noise ratio (SNR). Improvements in fluorescent dyes/probes and detector capabilities coupled with advanced (image) analysis strategies have fueled current developments for single protein biomarker detections. For example, in comparison to conventional ELISA (i.e., based on ensembled measurements), single-molecule fluorescence detection is more sensitive, faster, and more accurate with reduced background, high-throughput, and so on. In comparison to MS sequencing, fluorescence-based single-molecule protein sequencing can achieve the sequencing of peptides themselves with higher sensitivity. This review summarizes various typical single-molecule detection technologies including their methodology (modes of operation), detection limits, advantages and drawbacks, and current challenges with recent examples. We describe the fluorescence-based single-molecule protein sequencing/detection based on five kinds of technologies such as fluorosequencing, N-terminal amino acid binder, nanopore light sensing, and DNA nanotechnology. Finally, we present our perspective for developing high-performance fluorescence-based sequencing/detection techniques.




Similar content being viewed by others
References
Wu C, Garden PM, Walt DR. Ultrasensitive Detection of Attomolar Protein Concentrations by Dropcast Single Molecule Assays. J Am Chem Soc. 2020;142(28):12314–23.
Zhou F, Wang M, Yuan L, Cheng Z, Wu Z, Chen H. Sensitive sandwich ELISA based on a gold nanoparticle layer for cancer detection. Analyst. 2012;137(8):1779–84.
Chatterjee T, Knappik A, Sandford E, Tewari M, Choi SW, Strong WB, et al. Direct kinetic fingerprinting and digital counting of single protein molecules. Proc Natl Acad Sci. 2020;117(37):22815–22.
Mao C-P, Wang S-C, Su Y-P, Tseng S-H, He L, Wu AA, et al. Protein detection in blood with single-molecule imaging. Science Advances. 2021;7(33):eabg6522.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66(1):7–30.
Thaxton CS, Elghanian R, Thomas AD, Stoeva SI, Lee JS, Smith ND, et al. Nanoparticle-based bio-barcode assay redefines “undetectable” PSA and biochemical recurrence after radical prostatectomy. Proc Natl Acad Sci U S A. 2009;106(44):18437–42.
Blennow K, de Leon MJ, Zetterberg H. Alzheimer’s disease. The Lancet. 2006;368(9533):387–403.
Zecca C, Tortelli R, Panza F, Arcuti S, Piccininni M, Capozzo R, et al. Plasma β-amyloid1–42 reference values in cognitively normal subjects. J Neurol Sci. 2018;391:120–6.
Tatebe H, Kasai T, Ohmichi T, Kishi Y, Kakeya T, Waragai M, et al. Quantification of plasma phosphorylated tau to use as a biomarker for brain Alzheimer pathology: pilot case-control studies including patients with Alzheimer’s disease and down syndrome. Mol Neurodegener. 2017;12(1):63.
Zetterberg H, Wilson D, Andreasson U, Minthon L, Blennow K, Randall J, et al. Plasma tau levels in Alzheimer’s disease. Alzheimer’s Research & Therapy. 2013;5(2):9.
Akama K, Shirai K, Suzuki S. Droplet-Free Digital Enzyme-Linked Immunosorbent Assay Based on a Tyramide Signal Amplification System. Anal Chem. 2016;88(14):7123–9.
Zhang L, Fan W, Jia D, Feng Q, Ren W, Liu C. Microchamber-Free Digital Flow Cytometric Analysis of T4 Polynucleotide Kinase Phosphatase Based on Single-Enzyme-to-Single-Bead Space-Confined Reaction. Anal Chem. 2021;93(44):14828–36.
Shashkova S, Leake MC. Single-molecule fluorescence microscopy review: shedding new light on old problems. Biosci Rep. 2017;37(4):BSR20170031.
Pecker LH, Lanzkron S. In the Clinic Sickle Cell Disease. Ann Int Med. 2021;174(1):ITC1–16.
Timp W, Timp G. Beyond mass spectrometry, the next step in proteomics. Sci Adv. 2020;6(2):eaax8978.
Angel TE, Aryal UK, Hengel SM, Baker ES, Kelly RT, Robinson EW, et al. Mass spectrometry-based proteomics: existing capabilities and future directions. Chem Soc Rev. 2012;41(10):3912–28.
Swaminathan J, Boulgakov AA, Hernandez ET, Bardo AM, Bachman JL, Marotta J, et al. Highly parallel single-molecule identification of proteins in zeptomole-scale mixtures. Nat Biotechnol. 2018;36(11):1076–82.
Chait BT. Mass spectrometry: bottom-up or top-down? Science. 2006;314(5796):65–6.
Rissin DM, Kan CW, Campbell TG, Howes SC, Fournier DR, Song L, et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat Biotechnol. 2010;28(6):595–9.
Moerner WE, Kador L. Optical detection and spectroscopy of single molecules in a solid. Phys Rev Lett. 1989;62(21):2535–8.
Wilson DH, Rissin DM, Kan CW, Fournier DR, Piech T, Campbell TG, et al. The Simoa HD-1 Analyzer: A Novel Fully Automated Digital Immunoassay Analyzer with Single-Molecule Sensitivity and Multiplexing. SLAS Technology. 2016;21(4):533–47.
Chen Y, Shimoni O, Huang G, Wen S, Liao J, Duong HTT, et al. Upconversion nanoparticle-assisted single-molecule assay for detecting circulating antigens of aggressive prostate cancer. Cytometry A. 2022;101(5):400–10.
Farka Z, Mickert MJ, Hlavacek A, Skladal P, Gorris HH. Single Molecule Upconversion-Linked Immunosorbent Assay with Extended Dynamic Range for the Sensitive Detection of Diagnostic Biomarkers. Anal Chem. 2017;89(21):11825–30.
Cohen L, Cui N, Cai Y, Garden PM, Li X, Weitz DA, et al. Single Molecule Protein Detection with Attomolar Sensitivity Using Droplet Digital Enzyme-Linked Immunosorbent Assay. ACS Nano. 2020;14(8):9491–501.
Wu C, Dougan TJ, Walt DR. High-Throughput, High-Multiplex Digital Protein Detection with Attomolar Sensitivity. ACS Nano. 2022;16(1):1025–35.
Hao N, Zhang JXJ. Microfluidic Screening of Circulating Tumor Biomarkers toward Liquid Biopsy. Sep Purif Rev. 2018;47(1):19–48.
Shang L, Cheng Y, Zhao Y. Emerging Droplet Microfluidics. Chem Rev. 2017;117(12):7964–8040.
Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, et al. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods. 2013;10(10):1003–5.
Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, et al. High-Throughput Droplet Digital PCR System for Absolute Quantitation of DNA Copy Number. Anal Chem. 2011;83(22):8604–10.
Mazutis L, Gilbert J, Ung WL, Weitz DA, Griffiths AD, Heyman JA. Single-cell analysis and sorting using droplet-based microfluidics. Nat Protoc. 2013;8(5):870–91.
Prakadan SM, Shalek AK, Weitz DA. Scaling by shrinking: empowering single-cell “omics” with microfluidic devices. Nat Rev Genet. 2017;18(6):345–61.
Shahi P, Kim SC, Haliburton JR, Gartner ZJ, Abate AR. Abseq: Ultrahigh-throughput single cell protein profiling with droplet microfluidic barcoding. Sci Rep. 2017;7(1):44447.
Cui N, Zhang H, Schneider N, Tao Y, Asahara H, Sun Z, et al. A mix-and-read drop-based in vitro two-hybrid method for screening high-affinity peptide binders. Sci Rep. 2016;6(1):22575.
Abbaspourrad A, Zhang H, Tao Y, Cui N, Asahara H, Zhou Y, et al. Label-free single-cell protein quantification using a drop-based mix-and-read system. Sci Rep. 2015;5(1):12756.
Huang Q, Li N, Zhang H, Che C, Sun F, Xiong Y, et al. Critical Review: digital resolution biomolecular sensing for diagnostics and life science research. Lab Chip. 2020;20(16):2816–40.
Fan Y, Hao R, Han C, Zhang B. Counting Single Redox Molecules in a Nanoscale Electrochemical Cell. Anal Chem. 2018;90(23):13837–41.
Barulin A, Claude J-B, Patra S, Bonod N, Wenger J. Deep Ultraviolet Plasmonic Enhancement of Single Protein Autofluorescence in Zero-Mode Waveguides. Nano Lett. 2019;19(10):7434–42.
Fan S, Webb JEA, Yang Y, Nieves DJ, Goncales VR, Tran J, et al. Observing the Reversible Single Molecule Electrochemistry of Alexa Fluor 647 Dyes by Total Internal Reflection Fluorescence Microscopy. Angew Chem Int Ed Engl. 2019;58(41):14495–8.
Vukojevic V, Heidkamp M, Ming Y, Johansson B, Terenius L, Rigler R. Quantitative single-molecule imaging by confocal laser scanning microscopy. Proc Natl Acad Sci U S A. 2008;105(47):18176–81.
Macdonald PJ, Ruan Q, Tetin SY. Direct single-molecule counting for immunoassay applications. Anal Biochem. 2019;566:139–45.
Stroock AD, Dertinger SK, Ajdari A, Mezic I, Stone HA, Whitesides GM. Chaotic mixer for microchannels. Science. 2002;295(5555):647–51.
Axelrod D, T P Burghardt a, Thompson NL. Total Internal Reflection Fluorescence. Ann Rev Biophys Bioeng. 1984;13(1):247–68.
Khorasaninejad M, Chen WT, Devlin RC, Oh J, Zhu AY, Capasso F. Metalenses at visible wavelengths: Diffraction-limited focusing and subwavelength resolution imaging. Science. 2016;352(6290):1190–4.
Zhang C-Y, Yeh H-C, Kuroki MT, Wang T-H. Single-quantum-dot-based DNA nanosensor. Nat Mater. 2005;4(11):826–31.
Mickert MJ, Farka Z, Kostiv U, Hlaváček A, Horák D, Skládal P, et al. Measurement of Sub-femtomolar Concentrations of Prostate-Specific Antigen through Single-Molecule Counting with an Upconversion-Linked Immunosorbent Assay. Anal Chem. 2019;91(15):9435–41.
Aebersold R, Mann M. Mass-spectrometric exploration of proteome structure and function. Nature. 2016;537(7620):347–55.
Shendure J, Balasubramanian S, Church GM, Gilbert W, Rogers J, Schloss JA, et al. DNA sequencing at 40: past, present and future. Nature. 2017;550(7676):345–53.
Abascal F, Juan D, Jungreis I, Kellis M, Martinez L, Rigau M, et al. Loose ends: almost one in five human genes still have unresolved coding status. Nucleic Acids Res. 2018;46(14):7070–84.
Hu ZL, Huo MZ, Ying YL, Long YT. Biological Nanopore Approach for Single-Molecule Protein Sequencing. Angewandte Chemie-International Edition. 2021;60(27):14738–49.
Reed BD, Meyer MJ, Abramzon V, Ad O, Ad O, Adcock P, et al. Real-time dynamic single-molecule protein sequencing on an integrated semiconductor device. Science. 2022;378(6616):186–92.
Wang R, Gilboa T, Song J, Huttner D, Grinstaff MW, Meller A. Single-Molecule Discrimination of Labeled DNAs and Polypeptides Using Photoluminescent-Free TiO2 Nanopores. ACS Nano. 2018;12(11):11648–56.
Dai M, Yin P. Methods and compositions relating to super-resolution imaging and modification. US patent 10006917. 2018.
De Lannoy CV, Filius M, van Wee R, Joo C, de Ridder D. Evaluation of FRET X for single-molecule protein fingerprinting. iSci. 2021;24(11):103239.
Swaminathan J, Boulgakov AA, Marcotte EM. A theoretical justification for single molecule peptide sequencing. PLoS Comput Biol. 2015;11(2): e1004080.
Gavrilyuk J, Ban H, Nagano M, Hakamata W, Barbas CF 3rd. Formylbenzene Diazonium Hexafluorophosphate Reagent for Tyrosine-Selective Modification of Proteins and the Introduction of a Bioorthogonal Aldehyde. Bioconjug Chem. 2012;23(12):2321–8.
Ban H, Gavrilyuk J, Barbas CF III. Tyrosine bioconjugation through aqueous ene-type reactions: a click-like reaction for tyrosine. J Am Chem Soc. 2010;132(5):1523–5.
Bach K, Beerkens BLH, Zanon PRA, Hacker SM. Light-Activatable, 2,5-Disubstituted Tetrazoles for the Proteome-wide Profiling of Aspartates and Glutamates in Living Bacteria. ACS Cent Sci. 2020;6(4):546–54.
Hernandez ET, Swaminathan J, Marcotte EM, Anslyn EV. Solution-phase and solid-phase sequential, selective modification of side chains in KDYWEC and KDYWE as models for usage in single-molecule protein sequencing. New J Chem. 2017;41(2):462–9.
Taylor MT, Nelson JE, Suero MG, Gaunt MJ. A protein functionalization platform based on selective reactions at methionine residues. Nature. 2018;562(7728):563–8.
Lin Shixian, Yang Xiaoyu, Jia Shang, Weeks Amy M, MH, Peter S. Lee, Rita V. Nichiporuk ATI, James A. Wells, F. Dean Toste CJC. Redox-based reagents for chemoselective methionine bioconjugation. Sci. 2017;355(6325):597–602.
Christian AH, Jia S, Cao W, Zhang P, Meza AT, Sigman MS, et al. A Physical Organic Approach to Tuning Reagents for Selective and Stable Methionine Bioconjugation. J Am Chem Soc. 2019;141(32):12657–62.
Jia S, He D, Chang CJ. Bioinspired Thiophosphorodichloridate Reagents for Chemoselective Histidine Bioconjugation. J Am Chem Soc. 2019;141(18):7294–301.
Howard CJ, Floyd BM, Bardo AM, Swaminathan J, Marcotte EM, Anslyn EV. Solid-Phase Peptide Capture and Release for Bulk and Single-Molecule Proteomics. ACS Chem Biol. 2020;15(6):1401–7.
Perona JJ, Hadd A. Structural diversity and protein engineering of the aminoacyl-tRNA synthetases. Biochemistry. 2012;51(44):8705–29.
Dougan DA, Reid BG, Horwich AL, Bukau B. ClpS, a Substrate Modulator of the ClpAP Machine. Mol Cell. 2002;9(3):673–83.
Erbse A, Schmidt R, Bornemann T, Schneider-Mergener J, Mogk A, Zahn R, et al. ClpS is an essential component of the N-end rule pathway in Escherichia coli. Nature. 2006;439(7077):753–6.
Roman-Hernandez G, Hou JY, Grant RA, Sauer RT, Baker TA. The ClpS adaptor mediates staged delivery of N-end rule substrates to the AAA+ ClpAP protease. Mol Cell. 2011;43(2):217–28.
Nishimura K, van Wijk KJ. Organization, function and substrates of the essential Clp protease system in plastids. Biochim Biophys Acta. 2015;1847(9):915–30.
Tullman J, Callahan N, Ellington B, Kelman Z, Marino JP. Engineering ClpS for selective and enhanced N-terminal amino acid binding. Appl Microbiol Biotechnol. 2019;103(6):2621–33.
Rodriques SG, Marblestone AH, Boyden ES. A theoretical analysis of single molecule protein sequencing via weak binding spectra. PLoS ONE. 2019;14(3): e0212868.
Alfaro JA, Bohlander P, Dai MJ, Filius M, Howard CJ, van Kooten XF, et al. The emerging landscape of single-molecule protein sequencing technologies. Nat Methods. 2021;18(6):604–17.
Tullman J, Marino JP, Kelman Z. Leveraging nature’s biomolecular designs in next-generation protein sequencing reagent development. Appl Microbiol Biotechnol. 2020;104(17):7261–71.
Ferro ES, Rioli V, Castro LM, Fricker LD. Intracellular peptides: From discovery to function. EuPA Open Proteom. 2014;3:143–51.
Gonzales T, Robert-Baudouy J. Bacterial aminopeptidases: properties and functions. FEMS Microbiol Rev. 1996;18(4):319–44.
Sanderink G-J, Artur Y, Siest G. Human Aminopeptidases: A Review of the Literature. 1988;26(12):795–808.
Brinkerhoff H, Kang AS, Liu J, Aksimentiev A, Dekker C. Multiple rereads of single proteins at single–amino acid resolution using nanopores. Science. 2021;374(6574):1509–13.
Assad ON, Di Fiori N, Squires AH, Meller A. Two color DNA barcode detection in photoluminescence suppressed silicon nitride nanopores. Nano Lett. 2015;15(1):745–52.
Sawafta F, Clancy B, Carlsen AT, Huber M, Hall AR. Solid-state nanopores and nanopore arrays optimized for optical detection. Nanoscale. 2014;6(12):6991–6.
Dette C, Perez-Osorio MA, Kley CS, Punke P, Patrick CE, Jacobson P, et al. TiO2 anatase with a bandgap in the visible region. Nano Lett. 2014;14(11):6533–8.
Schneider J, Matsuoka M, Takeuchi M, Zhang J, Horiuchi Y, Anpo M, et al. Understanding TiO2 photocatalysis: mechanisms and materials. Chem Rev. 2014;114(19):9919–86.
Tang H, Prasad K, Sanjinès R, Schmid PE, Lévy F. Electrical and optical properties of TiO2anatase thin films. J Appl Phys. 1994;75(4):2042–7.
Chandrasekaran AR, Punnoose JA, Zhou L, Dey P, Dey BK, Halvorsen K. DNA nanotechnology approaches for microRNA detection and diagnosis. Nucleic Acids Res. 2019;47(20):10489–505.
Chen T, Ren L, Liu X, Zhou M, Li L, Xu J, et al. DNA Nanotechnology for Cancer Diagnosis and Therapy. International Journal of Molecular Sciences. 2018;19(6):1671.
Chi Q, Yang Z, Xu K, Wang C, Liang H. DNA Nanostructure as an Efficient Drug Delivery Platform for Immunotherapy. Front Pharmacol. 2019;10:1585.
Hu Q, Li H, Wang L, Gu H, Fan C. DNA Nanotechnology-Enabled Drug Delivery Systems. Chem Rev. 2019;119(10):6459–506.
Schnitzbauer J, Strauss MT, Schlichthaerle T, Schueder F, Jungmann R. Super-resolution microscopy with DNA-PAINT. Nat Protoc. 2017;12(6):1198–228.
Floyd BM, Marcotte EM. Protein Sequencing, One Molecule at a Time. Annu Rev Biophys. 2022;51:181–200.
Dai M, Jungmann R, Yin P. Optical imaging of individual biomolecules in densely packed clusters. Nat Nanotechnol. 2016;11(9):798–807.
Bustamante C, Alexander L, Maciuba K, Kaiser CM. Single-Molecule Studies of Protein Folding with Optical Tweezers. Annu Rev Biochem. 2020;89:443–70.
Filius M, Kim SH, Severins I, Joo C. High-Resolution Single-Molecule FRET via DNA eXchange (FRET X). Nano Lett. 2021;21(7):3295–301.
Funding
This work was supported by the National Natural Science Foundation of China (22150410330, 22074059).
Author information
Authors and Affiliations
Contributions
Haihan He, Chuhong Wu, and Muhammad Saqib contributed equally. The manuscript was written and proofread by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection Young Investigators in (Bio-)Analytical Chemistry 2023 with guest editors Zhi-Yuan Gu, Beatriz Jurado-Sánchez, Thomas H. Linz, Leandro Wang Hantao, Nongnoot Wongkaew, and Peng Wu.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, H., Wu, C., Saqib, M. et al. Single-molecule fluorescence methods for protein biomarker analysis. Anal Bioanal Chem 415, 3655–3669 (2023). https://doi.org/10.1007/s00216-022-04502-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04502-9