Abstract
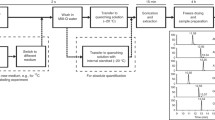
As microfluidic cell culture progresses, the need for robust and reproducible intracellular analyses grows. In particular, intracellular metabolites are subject to perturbation and degradation during the lysing process. The reliability of intracellular metabolomic analysis in microfluidic devices depends on the preservation of metabolite integrity during sample preparation and storage. Described here is a novel automated microfluidic system exhibiting the necessary rapid cellular lysis and quenching of enzymatic activity. Quenching efficiency was assessed using a novel ratiometric MALDI-MS-based assay of exogenous isotopic adenosine triphosphate (ATP) hydrolysis to isotopic adenosine diphosphate (ADP) as a marker of metabolite degradation. The lysis system of the microfluidic device was enhanced using a Peltier cooler to chill the lysate and quench aberrant enzymatic activity. Parameter optimization (flow rate, collection time, and temperature control) improved the endogenous and exogenous ADP/ATP ratios by 44.9% and 39.8% respectively consistent with traditional quenching techniques. The effects of chilling/quenching on metabolism were evaluated resulting in over 500 significant features compared to non-chilled from untargeted capillary LC-MS metabolomic analyses. These include increased levels of tryptophan, histidine, and pyruvate as well as decreased levels in UDP-N-acetylglucosamine. The results illustrate the need for both rapid lysis and quenching in microfluidic cell culture platforms.

Graphical abstract







Similar content being viewed by others
Abbreviations
- ATP:
-
Adenosine triphosphate
- ADP:
-
Adenosine diphosphate
- AMP:
-
Adenosine monophosphate
- EC:
-
Endothelial cell
- DMEM:
-
Dulbecco’s modified eagle medium
- NAD+:
-
Nicotinamide adenine dinucleotide
- UDP:
-
Uridine diphosphate
- MeOH:
-
Methanol
- ACN:
-
Acetonitrile
References
Ussher JR, Elmariah S, Gerszten RE, Dyck JRB. The emerging role of metabolomics in the diagnosis and prognosis of cardiovascular disease. J Am Coll Cardiol. 2016;68(25):2850.
Filla LA, Edwards JL. Metabolomics in diabetic complications. Mol BioSyst. 2016;12(4):1090–105. https://doi.org/10.1039/C6MB00014B.
Dunn WB, Ellis DI. Metabolomics: current analytical platforms and methodologies. TrAC Trends Anal Chem. 2005;24(4):285–94. https://doi.org/10.1016/j.trac.2004.11.021.
Zhang A, Sun H, Xu H, Qiu S, Wang X. Cell metabolomics. OMICS : a J Integr Biol. 2013;17(10):495–501. https://doi.org/10.1089/omi.2012.0090.
Lorenz MA, Burant CF, Kennedy RT. Reducing time and increasing sensitivity in sample preparation for adherent mammalian cell metabolomics. Anal Chem. 2011;83(9):3406–14. https://doi.org/10.1021/ac103313x.
Yanes O, Tautenhahn R, Patti GJ, Siuzdak G. Expanding coverage of the metabolome for global metabolite profiling. Anal Chem. 2011;83(6):2152–61. https://doi.org/10.1021/ac102981k.
Kimball E, Rabinowitz JD. Identifying decomposition products in extracts of cellular metabolites. Anal Biochem. 2006;358(2):273–80. https://doi.org/10.1016/j.ab.2006.07.038.
Shehadul Islam M, Aryasomayajula A, Selvaganapathy RP. A review on macroscale and microscale cell lysis methods. Micromachines. 2017;8(3). https://doi.org/10.3390/mi8030083.
Alshammari TM, Al-Hassan AA, Hadda TB, Aljofan M. Comparison of different serum sample extraction methods and their suitability for mass spectrometry analysis. Saudi Pharm J. 2015;23(6):689–97. https://doi.org/10.1016/j.jsps.2015.01.023.
Pinu RF, Villas-Boas GS, Aggio R. Analysis of intracellular metabolites from microorganisms: quenching and extraction protocols. Metabolites. 2017;7(4). https://doi.org/10.3390/metabo7040053.
Burns B, Mendz G, Hazell S. Methods for the measurement of a bacterial enzyme activity in cell lysates and extracts. Biological Procedures Online. 1998;1:17–26. https://doi.org/10.1251/bpo5.
Kraly JR, Holcomb RE, Guan Q, Henry CS. Review: microfluidic applications in metabolomics and metabolic profiling. Anal Chim Acta. 2009;653(1):23–35. https://doi.org/10.1016/j.aca.2009.08.037.
Filla LA, Sanders KL, Filla RT, Edwards JL. Automated sample preparation in a microfluidic culture device for cellular metabolomics. Analyst. 2016;141(12):3858–65. https://doi.org/10.1039/C6AN00237D.
Fitz JG. Regulation of cellular Atp release. Trans Am Clin Climatol Assoc. 2007;118:199–208.
Sellick CA, Hansen R, Stephens GM, Goodacre R, Dickson AJ. Metabolite extraction from suspension-cultured mammalian cells for global metabolite profiling. Nat Protoc. 2011;6:1241. https://doi.org/10.1038/nprot.2011.366.
Luz JA, Hans E, Zeng A-P. Automated fast filtration and on-filter quenching improve the intracellular metabolite analysis of microorganisms. Eng Life Sci. 2014;14(2):135–42. https://doi.org/10.1002/elsc.201300099.
León Z, García-Cañaveras JC, Donato MT, Lahoz A. Mammalian cell metabolomics: experimental design and sample preparation. ELECTROPHORESIS. 2013;34(19):2762–75. https://doi.org/10.1002/elps.201200605.
Kovarik ML, Allbritton NL. Measuring enzyme activity in single cells. Trends Biotechnol. 2011;29(5):222–30. https://doi.org/10.1016/j.tibtech.2011.01.003.
Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science. 2002;298(5600):1912.
Faijes M, Mars AE, Smid EJ. Comparison of quenching and extraction methodologies for metabolome analysis of lactobacillus plantarum. Microb Cell Factories. 2007;6(1):27. https://doi.org/10.1186/1475-2859-6-27.
Vuckovic D. Current trends and challenges in sample preparation for global metabolomics using liquid chromatography–mass spectrometry. Anal Bioanal Chem. 2012;403(6):1523–48. https://doi.org/10.1007/s00216-012-6039-y.
Kamerlin SCL, Warshel A. On the energetics of ATP hydrolysis in solution. J Phys Chem B. 2009;113(47):15692–8. https://doi.org/10.1021/jp907223t.
Mukhitov N, Yi L, Schrell AM, Roper MG. Optimization of a microfluidic electrophoretic immunoassay using a Peltier cooler. J Chromatogr A. 2014;1367:154–60. https://doi.org/10.1016/j.chroma.2014.09.040.
Wishart DS, Feunang YD, Marcu A, Guo AC, Liang K, Vázquez-Fresno R, et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 2018;46(D1):D608–D17. https://doi.org/10.1093/nar/gkx1089.
Gowda H, Ivanisevic J, Johnson CH, Kurczy ME, Benton HP, Rinehart D, et al. Interactive XCMS online: simplifying advanced metabolomic data processing and subsequent statistical analyses. Anal Chem. 2014;86(14):6931–9. https://doi.org/10.1021/ac500734c.
Groskreutz SR, Horner AR, Weber SG. Development of a 1.0 mm inside diameter temperature-assisted focusing precolumn for use with 2.1 mm inside diameter columns. J Chromatogr A. 2017;1523:193–203. https://doi.org/10.1016/j.chroma.2017.07.015.
Alley WR, Madera M, Mechref Y, Novotny MV. Chip-based reversed-phase liquid chromatography−mass spectrometry of permethylated N-linked glycans: a potential methodology for cancer-biomarker discovery. Anal Chem. 2010;82(12):5095–106. https://doi.org/10.1021/ac100131e.
Acknowledgments
The authors wish to thank Dr. Dave Wood of the Protein Core Facility in the Saint Louis University Department of Biochemistry for his assistance with MALDI analyses.
Funding
This work is supported by the National Institutes of Health (USA): 1R15GM113153 (JLE).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 222 kb)
Rights and permissions
About this article
Cite this article
Filla, L.A., Sanders, K.L., Coulton, J.B. et al. Determination of online quenching efficiency for an automated cellular microfluidic metabolomic platform using mass spectrometry based ATP degradation analysis. Anal Bioanal Chem 411, 6399–6407 (2019). https://doi.org/10.1007/s00216-019-02018-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-02018-3