Abstract
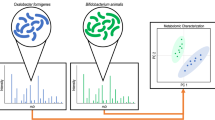
Diseases of oxalate, such as nephrolithiasis and primary hyperoxaluria, affect a significant portion of the US population and have limited treatment options. Oxalobacter formigenes, an obligate oxalotrophic bacterium in the mammalian intestine, has generated great interest as a potential probiotic or therapeutic treatment for oxalate-related conditions due to its ability to degrade both exogenous (dietary) and endogenous (metabolic) oxalate, lowering the risk of hyperoxaluria/hyperoxalemia. Although all oxalotrophs degrade dietary oxalate, Oxalobacter formigenes is the only species shown to initiate intestinal oxalate secretion to draw upon endogenous, circulating oxalate for consumption. Evidence suggests that Oxalobacter regulates oxalate transport proteins in the intestinal epithelium using an unidentified secreted bioactive compound, but the mechanism of this function remains elusive. It is essential to gain an understanding of the biochemical relationship between Oxalobacter and the host intestinal epithelium for this microbe to progress as a potential remedy for oxalate diseases. This investigation includes the first profiling of the metabolome and lipidome of Oxalobacter formigenes, specifically the human strain HC1 and rat strain OxWR, the only two strains shown thus far to initiate net intestinal oxalate secretion across native gut epithelia. This study was performed using untargeted and targeted metabolomics and lipidomics methodologies utilizing ultra-high-performance liquid chromatography-mass spectrometry. We report our findings that the metabolic profiles of these strains, although largely conserved, show significant differences in their expression of many compounds. Several strain-specific features were also detected. Discussed are trends in the whole metabolic profile as well as in individual features, both identified and unidentified.

ᅟ





Similar content being viewed by others
Abbreviations
- BMP:
-
Bis(monoacylglycero)phosphate
- CER:
-
Ceramide
- CL:
-
Cardiolipin
- CoQ10:
-
Coenzyme Q10
- DG:
-
Diacylglycerol
- DMPE:
-
Dimethylphosphatidylethanolamine
- FA:
-
Formic acid
- LPC:
-
Lysophosphatidylcholine
- LPE:
-
Lysophosphatidylethanolamine
- LSM:
-
Lysosphingomyelin
- OxPE:
-
Oxidized phosphatidylethanolamine
- PAzePC:
-
1-Palmitoyl-2-azelaoyl-sn-glycero-3-phosphocholine
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- PG:
-
Phosphatidylglycerol
- PI:
-
Phosphatidylinositol
- PS:
-
Phosphatidylserine
- SM:
-
Sphingomyelin
- SO:
-
Sphingosine
- TG:
-
Triacylglycerol
References
Scales CD, Smith AC, Hanley JM, Saigal CS, Project UDiA. Prevalence of kidney stones in the United States. Eur Urol. 2012;62(1):160–5. https://doi.org/10.1016/j.eururo.2012.03.052.
Worcester EM, Coe FL. Nephrolithiasis. Prim Care. 2008;35(2):369–91, vii. https://doi.org/10.1016/j.pop.2008.01.005.
Holmes RP, Goodman HO, Assimos DG. Contribution of dietary oxalate to urinary oxalate excretion. Kidney Int. 2001;59(1):270–6. https://doi.org/10.1046/j.1523-1755.2001.00488.x.
Holmes RP, Assimos DG. Glyoxylate synthesis, and its modulation and influence on oxalate synthesis. J Urol. 1998;160(5):1617–24.
Khan SR, Pearle MS, Robertson WG, Gambaro G, Canales BK, Doizi S, et al. Kidney stones. Nat Rev Dis Primers. 2017;3:17001. https://doi.org/10.1038/nrdp.2017.1.
Bouzidi H, Majdoub A, Daudon M, Najjar MF. Primary hyperoxaluria: a review. Nephrol Ther. 2016;12(6):431–6. https://doi.org/10.1016/j.nephro.2016.03.005.
Hatch M. Gut microbiota and oxalate homeostasis. Ann Transl Med. 2017;5(2):36. https://doi.org/10.21037/atm.2016.12.70.
Oppici E, Montioli R, Cellini B. Liver peroxisomal alanine:glyoxylate aminotransferase and the effects of mutations associated with primary hyperoxaluria type I: an overview. Biochim Biophys Acta. 2015;1854(9):1212–9. https://doi.org/10.1016/j.bbapap.2014.12.029.
Dawson KA, Allison MJ, Hartman PA. Isolation and some characteristics of anaerobic oxalate-degrading bacteria from the rumen. Appl Environ Microbiol. 1980;40(4):833–9.
Allison MJ, Dawson KA, Mayberry WR, Foss JG. Oxalobacter formigenes gen. nov., sp. nov.: oxalate-degrading anaerobes that inhabit the gastrointestinal tract. Arch Microbiol. 1985;141(1):1–7.
Hatch M, Cornelius J, Allison M, Sidhu H, Peck A, Freel RW. Oxalobacter sp. reduces urinary oxalate excretion by promoting enteric oxalate secretion. Kidney Int. 2006;69(4):691–8. https://doi.org/10.1038/sj.ki.5000162.
Hatch M, Gjymishka A, Salido EC, Allison MJ, Freel RW. Enteric oxalate elimination is induced and oxalate is normalized in a mouse model of primary hyperoxaluria following intestinal colonization with Oxalobacter. Am J Physiol Gastrointest Liver Physiol. 2011;300(3):G461–9. https://doi.org/10.1152/ajpgi.00434.2010.
Liu H, Garrett TJ, Tayyari F, Gu L. Profiling the metabolome changes caused by cranberry procyanidins in plasma of female rats using (1) H NMR and UHPLC-Q-Orbitrap-HRMS global metabolomics approaches. Mol Nutr Food Res. 2015;59(11):2107–18. https://doi.org/10.1002/mnfr.201500236.
Folch J, Lees M, Sloane Stanley GH. A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem. 1957;226(1):497–509.
Koelmel JP, Kroeger NM, Gill EL, Ulmer CZ, Bowden JA, Patterson RE, et al. Expanding lipidome coverage using LC-MS/MS data-dependent acquisition with automated exclusion list generation. J Am Soc Mass Spectrom. 2017;28(5):908–17. https://doi.org/10.1007/s13361-017-1608-0.
He L, Diedrich J, Chu YY, Yates JR. Extracting accurate precursor information for tandem mass spectra by RawConverter. Anal Chem. 2015;87(22):11361–7. https://doi.org/10.1021/acs.analchem.5b02721.
Kessner D, Chambers M, Burke R, Agus D, Mallick P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics. 2008;24(21):2534–6. https://doi.org/10.1093/bioinformatics/btn323.
Chambers MC, Maclean B, Burke R, Amodei D, Ruderman DL, Neumann S, et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol. 2012;30(10):918–20. https://doi.org/10.1038/nbt.2377.
Pluskal T, Castillo S, Villar-Briones A, Oresic M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics. 2010;11:395. https://doi.org/10.1186/1471-2105-11-395.
Wehrens R, Hageman JA, van Eeuwijk F, Kooke R, Flood PJ, Wijnker E, et al. Improved batch correction in untargeted MS-based metabolomics. Metabolomics. 2016;12:88. https://doi.org/10.1007/s11306-016-1015-8.
Koelmel JP, Kroeger NM, Ulmer CZ, Bowden JA, Patterson RE, Cochran JA, et al. LipidMatch: an automated workflow for rule-based lipid identification using untargeted high-resolution tandem mass spectrometry data. BMC Bioinformatics. 2017;18(1):331. https://doi.org/10.1186/s12859-017-1744-3.
Chong J, Soufan O, Li C, Caraus I, Li S, Bourque G, et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018;46(W1):W486–W94. https://doi.org/10.1093/nar/gky310.
van den Berg RA, Hoefsloot HC, Westerhuis JA, Smilde AK, van der Werf MJ. Centering, scaling, and transformations: improving the biological information content of metabolomics data. BMC Genomics. 2006;7:142. https://doi.org/10.1186/1471-2164-7-142.
Holm S. A simple sequentially rejective multiple test procedure. Scand J Stat. 1979;6(2):65–70.
Hatch M, Allison MJ, Yu F, Farmerie W. Genome sequence of Oxalobacter formigenes strain HC-1. Genome Announc. 2017;5(27). https://doi.org/10.1128/genomeA.00533-17.
Hatch M, Allison MJ, Yu F, Farmerie W. Genome sequence of Oxalobacter formigenes strain OXCC13. Genome Announc. 2017;5(28). https://doi.org/10.1128/genomeA.00534-17.
Gomelsky M, Galperin MY. Bacterial second messengers, cGMP and c-di-GMP, in a quest for regulatory dominance. EMBO J. 2013;32(18):2421–3. https://doi.org/10.1038/emboj.2013.193.
Gomelsky M. cAMP, c-di-GMP, c-di-AMP and now cGMP: bacteria use them all! Mol Microbiol. 2011;79(3):562–5. https://doi.org/10.1111/j.1365-2958.2010.07514.x.
An SQ, Chin KH, Febrer M, McCarthy Y, Yang JG, Liu CL, et al. A cyclic GMP-dependent signalling pathway regulates bacterial phytopathogenesis. EMBO J. 2013;32(18):2430–8. https://doi.org/10.1038/emboj.2013.165.
Marden JN, Dong Q, Roychowdhury S, Berleman JE, Bauer CE. Cyclic GMP controls Rhodospirillum centenum cyst development. Mol Microbiol. 2011;79(3):600–15. https://doi.org/10.1111/j.1365-2958.2010.07513.x.
Jonas K, Melefors O, Römling U. Regulation of c-di-GMP metabolism in biofilms. Future Microbiol. 2009;4(3):341–58. https://doi.org/10.2217/fmb.09.7.
Allen F, Pon A, Wilson M, Greiner R, Wishart D. CFM-ID: a web server for annotation, spectrum prediction and metabolite identification from tandem mass spectra. Nucleic Acids Research. 2014;42:(W1)W94–W99.
Guijas C, Montenegro-Burke JR, Domingo-Almenara X, Palermo A, Warth B, Hermann G, et al. METLIN: a technology platform for identifying knowns and unknowns. Anal Chem. 2018;90(5):3156–64. https://doi.org/10.1021/acs.analchem.7b04424.
Wishart DS, Feunang YD, Marcu A, Guo AC, Liang K, Vázquez-Fresno R, et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 2018;46(D1):D608–D17. https://doi.org/10.1093/nar/gkx1089.
Gil de la Fuente A, Godzien J, Fernández López M, Rupérez FJ, Barbas C, Otero A. Knowledge-based metabolite annotation tool: CEU Mass Mediator. J Pharm Biomed Anal. 2018;154:138–49. https://doi.org/10.1016/j.jpba.2018.02.046.
Parsons JB, Rock CO. Bacterial lipids: metabolism and membrane homeostasis. Prog Lipid Res. 2013;52(3):249–76. https://doi.org/10.1016/j.plipres.2013.02.002.
Qiu X, Choudhry AE, Janson CA, Grooms M, Daines RA, Lonsdale JT, et al. Crystal structure and substrate specificity of the beta-ketoacyl-acyl carrier protein synthase III (FabH) from Staphylococcus aureus. Protein Sci. 2005;14(8):2087–94. https://doi.org/10.1110/ps.051501605.
Jackowski S, Murphy CM, Cronan JE, Rock CO. Acetoacetyl-acyl carrier protein synthase. A target for the antibiotic thiolactomycin. J Biol Chem. 1989;264(13):7624–9.
Khandekar SS, Gentry DR, Van Aller GS, Warren P, Xiang H, Silverman C, et al. Identification, substrate specificity, and inhibition of the Streptococcus pneumoniae beta-ketoacyl-acyl carrier protein synthase III (FabH). J Biol Chem. 2001;276(32):30024–30. https://doi.org/10.1074/jbc.M101769200.
Reynolds CM, Kalb SR, Cotter RJ, Raetz CR. A phosphoethanolamine transferase specific for the outer 3-deoxy-D-manno-octulosonic acid residue of Escherichia coli lipopolysaccharide. Identification of the eptB gene and Ca2+ hypersensitivity of an eptB deletion mutant. J Biol Chem. 2005;280(22):21202–11. https://doi.org/10.1074/jbc.M500964200.
Choi KH, Heath RJ, Rock CO. beta-ketoacyl-acyl carrier protein synthase III (FabH) is a determining factor in branched-chain fatty acid biosynthesis. J Bacteriol. 2000;182(2):365–70.
He X, Reynolds KA. Purification, characterization, and identification of novel inhibitors of the beta-ketoacyl-acyl carrier protein synthase III (FabH) from Staphylococcus aureus. Antimicrob Agents Chemother. 2002;46(5):1310–8.
Sohlenkamp C, Geiger O. Bacterial membrane lipids: diversity in structures and pathways. FEMS Microbiol Rev. 2016;40(1):133–59. https://doi.org/10.1093/femsre/fuv008.
Koebnik R, Locher KP, Van Gelder P. Structure and function of bacterial outer membrane proteins: barrels in a nutshell. Mol Microbiol. 2000;37(2):239–53.
Epand RM, Epand RF. Lipid domains in bacterial membranes and the action of antimicrobial agents. Biochim Biophys Acta. 2009;1788(1):289–94. https://doi.org/10.1016/j.bbamem.2008.08.023.
Goldfine H. Bacterial-membranes and lipid packing theory. J Lipid Res. 1984;25(13):1501–7.
López-Lara IM, Sohlenkamp C, Geiger O. Membrane lipids in plant-associated bacteria: their biosyntheses and possible functions. Mol Plant-Microbe Interact. 2003;16(7):567–79. https://doi.org/10.1094/MPMI.2003.16.7.567.
Goldfine H. Lipids of Prokaryotes - structure and distribution. Curr Topics Membr Transport. 1982;17:1–43.
Lopez-Lara I, Geiger O. Novel pathway for phosphatidylcholine biosynthesis in bacteria associated with eukaryotes. J Biotechnol. 2001;91(2–3):211–21. https://doi.org/10.1016/S0168-1656(01)00331-5.
Sohlenkamp C, Lopez-Lara I, Geiger O. Biosynthesis of phosphatidylcholine in bacteria. Prog Lipid Res. 2003;42(2):115–62. https://doi.org/10.1016/S0163-7827(02)00050-4.
Acknowledgments
The authors would like to acknowledge Dr. Cory A. Leonard, Department of Pathology, Immunology and Laboratory Medicine, University of Florida, for her assistance with sample generation for this experiment, including media preparation and cell culture, harvest, and lysis. Also to be acknowledged are Dr. Jeremy P. Koelmel, Department of Pathology, Immunology and Laboratory Medicine, University of Florida, and Vanessa Y. Rubio, Department of Chemistry, University of Florida, for their contributions with figure generation for the lipidomics analysis and graphical abstract, respectively.
Funding
This work was funded by the National Institutes of Health grant 2R01DK088892-05A1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection Young Investigators in (Bio-)Analytical Chemistry with guest editors Erin Baker, Kerstin Leopold, Francesco Ricci, and Wei Wang.
Rights and permissions
About this article
Cite this article
Chamberlain, C.A., Hatch, M. & Garrett, T.J. Metabolomic and lipidomic characterization of Oxalobacter formigenes strains HC1 and OxWR by UHPLC-HRMS. Anal Bioanal Chem 411, 4807–4818 (2019). https://doi.org/10.1007/s00216-019-01639-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01639-y