Abstract
High-quality mass spectral libraries have become crucial in mass spectrometry-based metabolomics. Here, we investigate a workflow to generate accurate mass discrete and composite spectral libraries for metabolite identification and for SWATH mass spectrometry data processing. Discrete collision energy (5–100 eV) accurate mass spectra were collected for 532 metabolites from the human metabolome database (HMDB) by flow injection analysis and compiled into composite spectra over a large collision energy range (e.g., 10–70 eV). Full scan response factors were also calculated. Software tools based on accurate mass and predictive fragmentation were specially developed and found to be essential for construction and quality control of the spectral library. First, elemental compositions constrained by the elemental composition of the precursor ion were calculated for all fragments. Secondly, all possible fragments were generated from the compound structure and were filtered based on their elemental compositions. From the discrete spectra, it was possible to analyze the specific fragment form at each collision energy and it was found that a relatively large collision energy range (10–70 eV) gives informative MS/MS spectra for library searches. From the composite spectra, it was possible to characterize specific neutral losses as radical losses using in silico fragmentation. Radical losses (generating radical cations) were found to be more prominent than expected. From 532 metabolites, 489 provided a signal in positive mode [M+H]+ and 483 in negative mode [M-H]−. MS/MS spectra were obtained for 399 compounds in positive mode and for 462 in negative mode; 329 metabolites generated suitable spectra in both modes. Using the spectral library, LC retention time, response factors to analyze data-independent LC-SWATH-MS data allowed the identification of 39 (positive mode) and 72 (negative mode) metabolites in a plasma pool sample (total 92 metabolites) where 81 previously were reported in HMDB to be found in plasma.

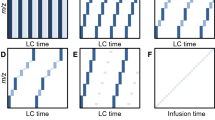
Library generation workflow for LC-SWATH MS, using collision energy spread, accurate mass, and fragment annotation.




Similar content being viewed by others
References
Wishart DS, Jewison T, Guo AC, Wilson M, Knox C, Liu YF, et al. HMDB 3.0—the Human Metabolome Database in 2013. Nucleic Acids Res. 2013;41(D1):D801–D7. https://doi.org/10.1093/nar/gks1065.
Rainville PD, Theodoridis G, Plumb RS, Wilson ID. Advances in liquid chromatography coupled to mass spectrometry for metabolic phenotyping. TRAC-Trend Anal Chem. 2014;61:181–91. https://doi.org/10.1016/j.trac.2014.06.005.
Brenton AG, Godfrey AR. Accurate mass measurement: terminology and treatment of data. J Am Soc Mass Spectrom. 2010;21(11):1821–35. https://doi.org/10.1016/j.jasms.2010.06.006.
Kind T, Tsugawa H, Cajka T, Ma Y, Lai Z, Mehta SS, et al. Identification of small molecules using accurate mass MS/MS search. Mass Spectrom Rev. 2017; https://doi.org/10.1002/mas.21535.
Ruttkies C, Schymanski EL, Wolf S, Hollender J, Neumann S. MetFrag relaunched: incorporating strategies beyond in silico fragmentation. J Cheminformatics. 2016;8. https://doi.org/10.1186/s13321-016-0115-9.
Ojanpera I, Kolmonen M, Pelander A. Current use of high-resolution mass spectrometry in drug screening relevant to clinical and forensic toxicology and doping control. Anal Bioanal Chem. 2012;403(5):1203–20. https://doi.org/10.1007/s00216-012-5726-z.
Stravs MA, Schymanski EL, Singer HP, Hollender J. Automatic recalibration and processing of tandem mass spectra using formula annotation. J Mass Spectrom. 2013;48(1):89–99. https://doi.org/10.1002/jms.3131.
Gillet LC, Navarro P, Tate S, Rost H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics. 2012;11(6):O111 016717. https://doi.org/10.1074/mcp.O111.016717.
Hopfgartner G, Tonoli D, Varesio E. High-resolution mass spectrometry for integrated qualitative and quantitative analysis of pharmaceuticals in biological matrices. Anal Bioanal Chem. 2012;402(8):2587–96. https://doi.org/10.1007/s00216-011-5641-8.
Siegel D, Meinema AC, Permentier H, Hopfgartner G, Bischoff R. Integrated quantification and identification of aldehydes and ketones in biological samples. Anal Chem. 2014;86(10):5089–100. https://doi.org/10.1021/ac500810r.
Scheidweiler KB, Jarvis MJ, Huestis MA. Nontargeted SWATH acquisition for identifying 47 synthetic cannabinoid metabolites in human urine by liquid chromatography-high-resolution tandem mass spectrometry. Anal Bioanal Chem. 2015;407(3):883–97. https://doi.org/10.1007/s00216-014-8118-8.
Roemmelt AT, Steuer AE, Kraemer T. Liquid chromatography, in combination with a quadrupole time-of-flight instrument, with sequential window acquisition of all theoretical fragment-ion spectra acquisition: validated quantification of 39 antidepressants in whole blood as part of a simultaneous screening and quantification procedure. Anal Chem. 2015;87(18):9294–301. https://doi.org/10.1021/acs.analchem.5b02031.
Go EP. Database resources in metabolomics: an overview. J Neuroimmune Pharm. 2010;5(1):18–30. https://doi.org/10.1007/s11481-009-9157-3.
Scheubert K, Hufsky F, Bocker S. Computational mass spectrometry for small molecules. J Cheminformatics. 2013;5(1):12. https://doi.org/10.1186/1758-2946-5-12.
Ausloos P, Clifton CL, Lias SG, Mikaya AI, Stein SE, Tchekhovskoi DV, et al. The critical evaluation of a comprehensive mass spectral library. J Am Soc Mass Spectrom. 1999;10(4):287–99.
Oberacher H, Pitterl F, Siapi E, Steele BR, Letzel T, Grosse S, et al. On the inter-instrument and the inter-laboratory transferability of a tandem mass spectral reference library. 3. Focus on ion trap and upfront CID. J Mass Spectrom. 2012;47(2):263–70. https://doi.org/10.1002/jms.2961.
Lei Z, Jing L, Qiu F, Zhang H, Huhman D, Zhou Z, et al. Construction of an ultrahigh pressure liquid chromatography-tandem mass spectral library of plant natural products and comparative spectral analyses. Anal Chem. 2015;87(14):7373–81. https://doi.org/10.1021/acs.analchem.5b01559.
Zhang Y, Bilbao A, Bruderer T, Luban J, Strambio-De-Castillia C, Lisacek F, et al. The use of variable Q1 isolation windows improves selectivity in LC-SWATH-MS acquisition. J Proteome Res. 2015;14(10):4359–71. https://doi.org/10.1021/acs.jproteome.5b00543.
Duchoslav E, Burton L, Varesio E, Bonner R, Hopfgartner G, editors. Towards comprehensive, reliable and accurate mass data repositories 2014; Proceedings of the 69th ASMS Conference on Mass Spectrometry and Allied Topics, Baltimore, MD, 15–19 June, 2014.
Bazso FL, Ozohanics O, Schlosser G, Ludanyi K, Vekey K, Drahos L. Quantitative comparison of tandem mass spectra obtained on various instruments. J Am Soc Mass Spectrom. 2016;27(8):1357–65. https://doi.org/10.1007/s13361-016-1408-y.
Broecker S, Herre S, Wust B, Zweigenbaum J, Pragst F. Development and practical application of a library of CID accurate mass spectra of more than 2,500 toxic compounds for systematic toxicological analysis by LC-QTOF-MS with data-dependent acquisition. Anal Bioanal Chem. 2011;400(1):101–17. https://doi.org/10.1007/s00216-010-4450-9.
Yang X, Neta P, Stein SE. Quality control for building libraries from electrospray ionization tandem mass spectra. Anal Chem. 2014;86(13):6393–400. https://doi.org/10.1021/ac500711m.
Stein S. Mass spectral reference libraries: an ever-expanding resource for chemical identification. Anal Chem. 2012;84(17):7274–82. https://doi.org/10.1021/ac301205z.
Vinaixa M, Schymanski EL, Neumann S, Navarro M, Salek RM, Yanes O. Mass spectral databases for LC/MS- and GC/MS-based metabolomics: state of the field and future prospects. TRAC-Trend Anal Chem. 2016;78(Supplement C):23–35. https://doi.org/10.1016/j.trac.2015.09.005.
Bruderer T, Varesio E, Hopfgartner G. The use of LC predicted retention times to extend metabolites identification with SWATH data acquisition. J Chromatogr B. 2017; https://doi.org/10.1016/j.jchromb.2017.07.016.
Niessen WMA, RA CC. Interpretation of MS-MS mass spectra of drugs and pesticides, Wiley Series on Mass Spectrometry. Hoboken, New Jersey: Wiley; 2017.
Bonner R, Hopfgartner G. SWATH acquisition mode for drug metabolism and metabolomics investigations. Bioanalysis. 2016;8(16):1735–50. https://doi.org/10.4155/bio-2016-0141.
Bocker S. Searching molecular structure databases using tandem MS data: are we there yet? Curr Opin Chem Biol. 2017;36:1–6. https://doi.org/10.1016/j.cbpa.2016.12.010.
Simon-Manso Y, Lowenthal MS, Kilpatrick LE, Sampson ML, Telu KH, Rudnick PA, et al. Metabolite profiling of a NIST standard reference material for human plasma (SRM 1950): GC-MS, LC-MS, NMR, and clinical laboratory analyses, libraries, and web-based resources. Anal Chem. 2013;85(24):11725–31. https://doi.org/10.1021/ac402503m.
Boudah S, Olivier MF, Aros-Calt S, Oliveira L, Fenaille F, Tabet JC, et al. Annotation of the human serum metabolome by coupling three liquid chromatography methods to high-resolution mass spectrometry. J Chromatogr B. 2014; https://doi.org/10.1016/j.jchromb.2014.04.025.
Acknowledgements
GH would like to thank SystemsX and the Swiss National Sciences Foundation for the financial support: Projects 51RTP0_151032 and 206021_170779.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Plasma samples were obtained from healthy voluntary who have consented that their donation or some of its components being used for medical research after final anonymization or in a coded form. The plasma samples were provided by the Centre de Transfusion Sanguine, University Hospital Geneva, Geneva, Switzerland. The Human Research Act (HRA) does not apply for the anonymized pooled plasma samples analyzed in the present work (Art. 2 para. 2 let. b and c).
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 2180 kb)
Rights and permissions
About this article
Cite this article
Bruderer, T., Varesio, E., Hidasi, A.O. et al. Metabolomic spectral libraries for data-independent SWATH liquid chromatography mass spectrometry acquisition. Anal Bioanal Chem 410, 1873–1884 (2018). https://doi.org/10.1007/s00216-018-0860-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-0860-x