Abstract
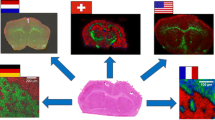
A standardized workflow for matrix-assisted laser desorption/ionization imaging mass spectrometry (MALDI imaging MS) is a prerequisite for the routine use of this promising technology in clinical applications. We present an approach to develop standard operating procedures for MALDI imaging MS sample preparation of formalin-fixed and paraffin-embedded (FFPE) tissue sections based on a novel quantitative measure of dataset quality. To cover many parts of the complex workflow and simultaneously test several parameters, experiments were planned according to a fractional factorial design of experiments (DoE). The effect of ten different experiment parameters was investigated in two distinct DoE sets, each consisting of eight experiments. FFPE rat brain sections were used as standard material because of low biological variance. The mean peak intensity and a recently proposed spatial complexity measure were calculated for a list of 26 predefined peptides obtained by in silico digestion of five different proteins and served as quality criteria. A five-way analysis of variance (ANOVA) was applied on the final scores to retrieve a ranking of experiment parameters with increasing impact on data variance.

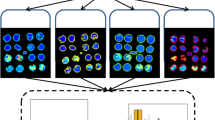
MALDI imaging experiments were planned according to fractional factorial design of experiments for the parameters under study. Selected peptide images were evaluated by the chosen quality metric (structure and intensity for a given peak list), and the calculated values were used as an input for the ANOVA. The parameters with the highest impact on the quality were deduced and SOPs recommended.




Similar content being viewed by others
References
Stoeckli M, Chaurand P, Hallahan DE, Caprioli RM. Imaging mass spectrometry: a new technology for the analysis of protein expression in mammalian tissues. Nat Med. 2001;7:493–6.
Caprioli RM, Farmer TB, Gile J. Molecular imaging of biological samples: localization of peptides and proteins using MALDI-TOF MS. Anal Chem. 1997;69:4751–60.
McDonnell LA, Corthals GL, Willems SM, van Remoortere A, van Zeijl RJM, et al. Peptide and protein imaging mass spectrometry in cancer research. J Proteomics. 2010;73:1921–44.
OuYang C, Liang Z, Li L. Mass spectrometric analysis of spatio-temporal dynamics of crustacean neuropeptides. Biochim Biophys Acta Proteins Proteomics. 2015;1854:798–811.
Burnum KE, Cornett DS, Puolitaival SM, Milne SB, Myers DS, et al. Spatial and temporal alterations of phospholipids determined by mass spectrometry during mouse embryo implantation. J Lipid Res. 2009;50:2290–8.
Sun N, Fernandez IE, Wei M, Wu Y, Aichler M, et al. Pharmacokinetic and pharmacometabolomic study of pirfenidone in normal mouse tissues using high mass resolution MALDI-FTICR-mass spectrometry imaging. Histochem Cell Biol. 2016;145:201–11.
Reyzer ML, Caprioli RM. MALDI-MS-based imaging of small molecules and proteins in tissues. Curr Opin Chem Biol. 2007;11:29–35.
Aichler M, Walch A. MALDI Imaging mass spectrometry: current frontiers and perspectives in pathology research and practice. Lab Invest. 2015;95:422–31.
Elsner M, Rauser S, Maier S, Schӧne C, Balluff B, Meding S, et al. MALDI imaging mass spectrometry reveals COX7A2, TAGLN2 and S100-A10 as novel prognostic markers in Barrett’s adenocarcinoma. J Proteomics. 2012;75:4693–703.
Rauser S, Marquardt C, Balluff B, Deininger S-O, Albers C, et al. Classification of HER2 receptor status in breast cancer tissues by MALDI imaging mass spectrometry. J Proteome Res. 2010;9:1854–63.
Poté N, Alexandrov T, Le Faouder J, Laouirem S, Léger T, Mebarki M, et al. Imaging mass spectrometry reveals modified forms of histone H4 as new biomarkers of microvascular invasion in hepatocellular carcinomas. Hepatolog. 2013;58:983–94.
Veselkov KA, Mirnezami R, Strittmatter N, Goldin RD, Kinross J, Speller AV, et al. Chemo-informatic strategy for imaging mass spectrometry-based hyperspectral profiling of lipid signatures in colorectal cancer. PNAS. 2014;111:1216–21.
Cazares LH, Troyer D, Mendrinos S, Lance RA, Nyalwidhe JO, Beydoun HA, et al. Imaging mass spectrometry of a specific fragment of mitogen-activated protein kinase/extracellular signal-regulated kinase kinase kinase 2 discriminates cancer from uninvolved prostate tissue. Clin Cancer Res. 2009;15:5541–51.
Casadonte R, Kriegsmann M, Zweynert F, Friedrich K, Bretton G, Otto M, et al. Imaging mass spectrometry to discriminate breast from pancreatic cancer metastasis in formalin-fixed paraffin-embedded tissues. Proteomics. 2014;14:956–64.
Casadonte R, Caprioli RM. Proteomic analysis of formalin-fixed paraffin-embedded tissue by MALDI imaging mass spectrometry. Nat Protoc. 2011;6:1695–709.
Lemaire R, Desmons A, Tabet J, Day R, Salzet M, Fournier I. Direct analysis and MALDI imaging of formalin-fixed, paraffin-embedded tissue sections. J Proteome Res. 2007;6:1295–305.
Thavarajah R, Mudimbaimannar VK, Elizabeth J, Rao UK, Ranganathan K. Chemical and physical basics of routine formaldehyde fixation. J Oral Maxillofac Pathol. 2012;16:400–5.
Fowler CB, Evers DL, O’Leary TJ, Mason JT. Antigen Retrieval Causes Protein Unfolding Evidence for a Linear Epitope Model of Recovered Immunoreactivity. J Histochem Cytochem. 2011;59:366–81.
Diehl HC, Beine B, Elm J, Trede D, Ahrens M, Eisenacher M, et al. The challenge of on-tissue digestion for MALDI MSI—a comparison of different protocols to improve imaging experiments. Anal Bioanal Chem. 2015;407:2223–43.
Hecht ES, Oberg AL, Muddiman DC. Optimizing mass spectrometry analyses: a tailored review on the utility of design of experiments. J Am Soc Mass Spectrom. 2016;27:767–85.
Riter LS, Vitek O, Gooding KM, Hodge BD, Julian RK. Statistical design of experiments as a tool in mass spectrometry. J Mass Spectrom. 2005;40:565–79.
Alexandrov T. MALDI imaging mass spectrometry: statistical data analysis and current computational challenges. BMC Bioinf. 2012;13:S11. doi:10.1186/1471-2105-13-S16-S11.
Alexandrov T, Bartels A. Testing for presence of known and unknown molecules in imaging mass spectrometry. Bioinformatics. 2013;29:2335–42.
Palmer A, Ovchinnikova E, Thuné M, Lavigne R, Guével B, Dyatlov A, et al. Using collective expert judgements to evaluate quality measures of mass spectrometry images. Bioinformatics. 2015;31:i375–84.
Montgomery DC. Design and analysis of experiments. John Wiley & Sons; 2008.
Groseclose MR, Andersson M, Hardesty WM, Caprioli RM. Identification of proteins directly from tissue: in situ tryptic digestions coupled with imaging mass spectrometry. J Mass Spectrom. 2007;42:254–62.
Groseclose MR, Massion PP, Chaurand P, Caprioli RM. High-throughput proteomic analysis of formalin-fixed paraffin-embedded tissue microarrays using MALDI imaging mass spectrometry. Proteomics. 2008;8:3715–24.
Gustafsson JO, Oehler MK, McColl SR, Hoffmann P. Citric acid antigen retrieval (CAAR) for tryptic peptide imaging directly on archived formalin-fixed paraffin-embedded tissue. J Proteome Res. 2010;9:4315–28.
Deutskens F, Yang J, Caprioli RM. High spatial resolution imaging mass spectrometry and classical histology on a single tissue section. J Mass Spectrom. 2011;46:568–71.
Schober Y, Schramm T, Spengler B, Rӧmpp A. Protein identification by accurate mass matrix-assisted laser desorption/ionization imaging of tryptic peptides. Rapid Commun Mass Spectrom. 2011;25:2475–83.
Liebeke M, Strittmatter N, Fearn S, Morgan AJ, Kille P, Fuchser J, et al. Unique metabolites protect earthworms against plant polyphenols. Nat Commun. 2015. doi:10.1038/ncomms8869.
Lauzon N, Dufresne M, Chauhan V, Chaurand P. Development of laser desorption imaging mass spectrometry methods to investigate the molecular composition of latent fingermarks. J Am Soc Mass Spectrom. 2015;26:878–86.
Mascini NE, Eijkel GB, ter Brugge P, Jonkers J, Wesseling J, Heeren RM. The use of mass spectrometry imaging to predict treatment response of patient-derived xenograft models of triple-negative breast cancer. J Proteome Res. 2015;14:1069–75.
Kriegsmann J, Kriegsmann M, Casadonte R. MALDI TOF imaging mass spectrometry in clinical pathology: a valuable tool for cancer diagnostics (review). Int J Oncol. 2015;46:893–906.
Balluff B, Elsner M, Kowarsch A, Rauser S, Meding S, Schuhmacher C, et al. Classification of HER2/neu status in gastric cancer using a breast-cancer derived proteome classifier. J Proteome Res. 2010;9:6317–22.
Clough T, Braun S, Fokin V, Ott I, Ragg S, Schadow G, et al. Statistical design and analysis of label-free LC-MS proteomic experiments: a case study of coronary artery disease. Serum/Plasma Proteomics. 2011;293–319.
Havlivs J, Thomas H, Sebela M, Shevchenko A. Fast-response proteomics by accelerated in-gel digestion of proteins. Anal Chem. 2003;75:1300–6.
Acknowledgments
We would like to thank Olga Vitek for valuable discussions regarding the statistical design of experiments. The authors gratefully acknowledge the financial support from the European Commission Seventh Framework Program (project 3D-MASSOMICS, grant 305259), the German Science Foundation (DFG core facility MALDI-MULTI), and the German Central Innovation Program for SMEs of the German Federal Ministry of Economic Affairs and Energy (BMWI-ZIM grant 2443904SB4).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Theodore Alexandrov is the Scientific Director, Peter Maass is on the Advisory Board, and Tobias Boskamp is a consultant for SCiLS GmbH, a company that develops and markets software for imaging mass spectrometry. At the time of analysis, Michael Becker was employed at Bruker Daltonics GmbH, a vendor of mass spectrometry instrumentation. All other authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants. All animal care guidelines according to the German animal protection law were met.
Informed consent
Not applicable
Rights and permissions
About this article
Cite this article
Oetjen, J., Lachmund, D., Palmer, A. et al. An approach to optimize sample preparation for MALDI imaging MS of FFPE sections using fractional factorial design of experiments. Anal Bioanal Chem 408, 6729–6740 (2016). https://doi.org/10.1007/s00216-016-9793-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9793-4