Abstract
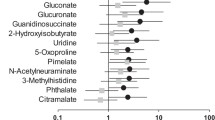
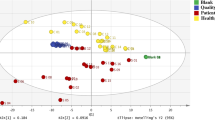
Capillary electrophoresis coupled with time-of-flight mass spectrometry was used to explore new serum biomarkers with high sensitivity and specificity for diabetic nephropathy (DN) diagnosis, through comprehensive analysis of serum metabolites with 78 diabetic patients. Multivariate analyses were used for identification of marker candidates and development of discriminative models. Of the 289 profiled metabolites, orthogonal partial least-squares discriminant analysis identified 19 metabolites that could distinguish between DN with macroalbuminuria and diabetic patients without albuminuria. These identified metabolites included creatinine, aspartic acid, γ-butyrobetaine, citrulline, symmetric dimethylarginine (SDMA), kynurenine, azelaic acid, and galactaric acid. Significant correlations between all these metabolites and urinary albumin-to-creatinine ratios (p < 0.009, Spearman’s rank test) were observed. When five metabolites (including γ-butyrobetaine, SDMA, azelaic acid and two unknowns) were selected from 19 metabolites and applied for multiple logistic regression model, AUC value for diagnosing DN was 0.927 using the whole dataset, and 0.880 in a cross-validation test. In addition, when four known metabolites (aspartic acid, SDMA, azelaic acid and galactaric acid) were applied, the resulting AUC was still high at 0.844 with the whole dataset and 0.792 with cross-validation. Combination of serum metabolomics with multivariate analyses enabled accurate discrimination of DN patients. The results suggest that capillary electrophoresis-mass spectrometry based metabolome analysis could be used for DN diagnosis.


Similar content being viewed by others
References
Gross JL, de Azevedo MJ, Silveiro SP, Canani LH, Caramori ML, Zelmanovitz T (2005) Diabetic nephropathy: diagnosis, prevention, and treatment. Diabetes Care 28(1):164–176
Kim HJ, Cho EH, Yoo JH, Kim PK, Shin JS, Kim MR, Kim CW (2007) Proteome analysis of serum from type 2 diabetics with nephropathy. J Proteome Res 6(2):735–743
Molitch ME, DeFronzo RA, Franz MJ, Keane WF, Mogensen CE, Parving HH, Steffes MW (2004) Nephropathy in diabetes. Diabetes Care 27(Suppl 1):S79–S83
Tabaei BP, Al-Kassab AS, Ilag LL, Zawacki CM, Herman WH (2001) Does microalbuminuria predict diabetic nephropathy? Diabetes Care 24(9):1560–1566
Dihazi H, Muller GA, Lindner S, Meyer M, Asif AR, Oellerich M, Strutz F (2007) Characterization of diabetic nephropathy by urinary proteomic analysis: identification of a processed ubiquitin form as a differentially excreted protein in diabetic nephropathy patients. Clin Chem 53(9):1636–1645
Mischak H, Kaiser T, Walden M, Hillmann M, Wittke S, Herrmann A, Knueppel S, Haller H, Fliser D (2004) Proteomic analysis for the assessment of diabetic renal damage in humans. Clin Sci (Lond) 107(5):485–495
Tiziani S, Lopes V, Gunther UL (2009) Early stage diagnosis of oral cancer using 1H NMR-based metabolomics. Neoplasia 11(3):269–276
Kind T, Tolstikov V, Fiehn O, Weiss RH (2007) A comprehensive urinary metabolomic approach for identifying kidney cancer. Anal Biochem 363(2):185–195
Sieber M, Wagner S, Rached E, Amberg A, Mally A, Dekant W (2009) Metabonomic study of ochratoxin a toxicity in rats after repeated administration: phenotypic anchoring enhances the ability for biomarker discovery. Chem Res Toxicol 22(7):1221–1231
Hirayama A, Kami K, Sugimoto M, Sugawara M, Toki N, Onozuka H, Kinoshita T, Saito N, Ochiai A, Tomita M, Esumi H, Soga T (2009) Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res 69(11):4918–4925
Soga T, Baran R, Suematsu M, Ueno Y, Ikeda S, Sakurakawa T, Kakazu Y, Ishikawa T, Robert M, Nishioka T, Tomita M (2006) Differential metabolomics reveals ophthalmic acid as an oxidative stress biomarker indicating hepatic glutathione consumption. J Biol Chem 281(24):16768–16776
Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M (2010) Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics 6(1):78–95
Xia JF, Liang QL, Hu P, Wang YM, Li P, Luo GA (2009) Correlations of six related purine metabolites and diabetic nephropathy in Chinese type 2 diabetic patients. Clin Biochem 42(3):215–220
Jiang Z, Liang Q, Luo G, Hu P, Li P, Wang Y (2009) HPLC-electrospray tandem mass spectrometry for simultaneous quantitation of eight plasma aminothiols: application to studies of diabetic nephropathy. Talanta 77(4):1279–1284
Zhang J, Yan L, Chen W, Lin L, Song X, Yan X, Hang W, Huang B (2009) Metabonomics research of diabetic nephropathy and type 2 diabetes mellitus based on UPLC-oaTOF-MS system. Anal Chim Acta 650(1):16–22
Soga T, Ohashi Y, Ueno Y, Naraoka H, Tomita M, Nishioka T (2003) Quantitative metabolome analysis using capillary electrophoresis mass spectrometry. J Proteome Res 2(5):488–494
Soga T, Igarashi K, Ito C, Mizobuchi K, Zimmermann HP, Tomita M (2009) Metabolomic profiling of anionic metabolites by capillary electrophoresis mass spectrometry. Anal Chem 81(15):6165–6174
Witten I, Frank E (2000) Data mining: practical machine learning algorithms with Java implementations. Morgan Kaufmann Publishers, CA
Xia JF, Liang QL, Liang XP, Wang YM, Hu P, Li P, Luo GA (2009) Ultraviolet and tandem mass spectrometry for simultaneous quantification of 21 pivotal metabolites in plasma from patients with diabetic nephropathy. J Chromatogr B Analyt Technol Biomed Life Sci 877(20–21):1930–1936
Perrone RD, Madias NE, Levey AS (1992) Serum creatinine as an index of renal function: new insights into old concepts. Clin Chem 38(10):1933–1953
Chuang CK, Lin SP, Chen HH, Chen YC, Wang TJ, Shieh WH, Wu CJ (2006) Plasma free amino acids and their metabolites in Taiwanese patients on hemodialysis and continuous ambulatory peritoneal dialysis. Clin Chim Acta 364(1–2):209–216
Fleck C, Janz A, Schweitzer F, Karge E, Schwertfeger M, Stein G (2001) Serum concentrations of asymmetric (ADMA) and symmetric (SDMA) dimethylarginine in renal failure patients. Kidney Int 59:S14–S18
Pahlich S, Zakaryan RP, Gehring H (2006) Protein arginine methylation: cellular functions and methods of analysis. Biochim Biophys Acta 1764(12):1890–1903
Saito K, Fujigaki S, Heyes MP, Shibata K, Takemura M, Fujii H, Wada H, Noma A, Seishima M (2000) Mechanism of increases in L-kynurenine and quinolinic acid in renal insufficiency. American Journal of Physiology-Renal Physiology 279(3):F565
Toyohara T, Akiyama Y, Suzuki T, Takeuchi Y, Mishima E, Tanemoto M, Momose A, Toki N, Sato H, Nakayama M, Hozawa A, Tsuji I, Ito S, Soga T, Abe T (2010) Metabolomic profiling of uremic solutes in CKD patients. Hypertens Res 33(9):944–952
Akamatsu H, Komura J, Asada Y, Miyachi Y, Niwa Y (1991) Inhibitory effect of azelaic acid on neutrophil functions: a possible cause for its efficacy in treating pathogenetically unrelated diseases. Arch Dermatol Res 283(3):162–166
Watanabe S, Yamada M, Ohtsu I, Makino K (2007) α-Ketoglutaric semialdehyde dehydrogenase isozymes involved in metabolic pathways of D-glucarate, D-galactarate, and hydroxy-L-proline. J Biol Chem 282(9):6685
Acknowledgments
We thank Dr. Astuko Watarai, Japan Labour Health and Welfare Organization Chubu Rosai Hospital, for assistance with sample collection. We also thank Maki Sugawara and Hiroko Ueda, Institute for Advanced Biosciences, Keio University, and Jiro Nakamura, Department of Endocrinology and Diabetes, Nagoya University Graduate School of Medicine, for technical support and fruitful discussions. This work was supported by a Health and Labour Sciences Research Grant “Research on Biological Markers for New Drug Development”, Grants from the Ministry of Health, Labour and Welfare of Japan “Research on Rare and Intractable Disease”, KAKENHI Grants-in-Aid for Scientific Research on Priority Areas “Systems Genomes” and “Lifesurveyor” from the Ministry of Education, Culture, Sports, Science and Technology of Japan, and research funds from the Yamagata prefectural government and the City of Tsuruoka.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hirayama, A., Nakashima, E., Sugimoto, M. et al. Metabolic profiling reveals new serum biomarkers for differentiating diabetic nephropathy. Anal Bioanal Chem 404, 3101–3109 (2012). https://doi.org/10.1007/s00216-012-6412-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-012-6412-x