Abstract
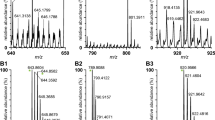
Desorption electrospray ionization mass spectrometry (DESI-MS) was investigated as a method to detect and identify peptides from tryptic digests of cytochrome c and myoglobin separated on ProteoChrom® HPTLC Silica gel 60 F254s plates and ProteoChrom® HPTLC Cellulose sheets. Full-scan mass spectra and data-dependent tandem mass spectra were acquired in separate plate scans and used to identify peptide ions. Peptide distributions along the development lane were mapped for each separated protein digest. Signal levels ranged over several orders of magnitude. In general, highest signal levels were obtained for the peptides with the highest R f values on a plate, while peptides with very low R f values were often not detected. Sequence coverages for cytochrome c were 58% for the digest separated on the silica gel plate and 72% for the separation on the cellulose sheet; myoglobin sequence coverages were 62% and 68% on silica gel and cellulose, respectively. Weak correlations between peptide hydrophilicity and R f values on the silica gel and cellulose plates were found, with the more hydrophilic peptides having lower R f values.





Similar content being viewed by others
References
Takáts Z, Wiseman JM, Gologan B, Cooks RG (2004) Mass spectrometry sampling under ambient conditions with desorption electrospray ionization. Science 306:471–473
Takáts Z, Wiseman JM, Cooks RG (2005) Ambient mass spectrometry using desorption electrospray ionization (DESI): instrumentation, mechanisms and applications in forensics, chemistry, and biology. J Mass Spectrom 40:1261–1275
Cooks RG, Ouyang Z, Takáts Z, Wiseman JM (2006) Ambient mass spectrometry. Science 311:1566–1570
Myung S, Wiseman JM, Valentine SJ, Takáts Z, Cooks RG, Clemmer DE (2006) Coupling desorption electrospray ionization with ion mobility/mass spectromtery for analysis of protein structure: evidence for desorption of folded and denatured states. J Phys Chem 110:5045–5051
Bereman MS, Nyadong L, Fernandez FM, Muddiman DC (2006) Direct high-resolution peptide and protein analysis by desorption electrspray ionization fourier transform ion cyclotron resonance mass spectrometry. Rapid Commun Mass Spectrom 20:3409–3411
Shin Y-S, Drolet B, Mayer R, Dolence K, Basile F (2007) Desorption electrospray ionization mass spectrometry of proteins. Anal Chem 79:3514–3518
Kaur-Atwal G, Weston DJ, Green PS, Crosland S, Bonner PLR, Creaser CS (2007) Analysis of tryptic peptides using desorption electrospray ionization combined with ion mobility spectrometry/mass spectromtetry. Rapid Commun Mass Spectrom 21:1131–1138
Cotte-Rodriguez I, Mulligan CC, Cooks RG (2007) Non-proximate detection of small and large molecules by desorption electrospray ionization and desorption atmospheric pressure chemical ionzation mass spectrometry: instrumentation and applications in forensics, chemistry, and biology. Anal Chem 79:7069–7077
Van Berkel GJ, Ford MJ, Deibel MA (2005) Thin-layer chromatography and mass spectrometry coupled using desorption electrospray ionization. Anal Chem 77:1207–1215
Van Berkel GJ, Kertesz V (2006) Automated sampling and imaging of analytes separated on thin-layer chromatography plates using desorption electrospray ionization mass spectrometry. Anal Chem 78:4938–4944
Kauppila TJ, Talaty N, Salo PK, Kotiaho T, Kostiainen R, Cooks RG (2006) New surfaces for desorption electrospray ionization mass spectrometry: porous silicon and ultra-thin layer chromatography plates. Rapid Commun Mass Spectrom 20:2143–2150
Van Berkel GJ, Tomkins BA, Kertesz V (2007) Thin-layer chromatography/desorption electrospray ionization mass spectrometry: investigation of goldenseal alkaloids. Anal Chem 79:2778–2789
Frölich T, Arnold GJ (2006) Proteome research based on modern liquid chromatography–tandem mass spectrometry: separation, identification and quantification. J Neural Transm 113:973–994
Tabb DL, Narasimhan C, Strader BL, Hettich RL (2005) DBDigger: reorganized proteomic database identification that improves flexibility and speed. Anal Chem 77:2464–2474
Narasimhan C, Tabb DL, VerBerkmoes NC, Thompson MR, Hettich RL, Uberbacher EC (2005) MASPIC: intensity-based tandem mass spectrometry scoring scheme that improves peptide identification at high confidence. Anal Chem 77:7581–7593
Wysocki VH, Resing KA, Zhang Q, Cheng G (2005) Mass spectrometry of peptides and proteins. Methods 35:211–222
Laboratory of Mass Spectrometry and Gaseous Ion Chemistry, The Rockefeller University. http://prowl.rockefeller.edu. Accessed 10 Jan 2008
Hydrophilicity calculator. Innovagen, Lund. http://www.innovagen.se/custom-peptide-synthesis/peptide-property-calculator. Accessed 10 Jan 2008
Hopp TP, Woods KR (1981) Prediction of protein antigenic determinants from amino acid sequences. Proc Nat Acad Sci USA 78:3824–3828
Cech NB, Enke CG (2000) Relating electrospray ionization response to nonpolar character of small peptides. Anal Chem 72:2717–2723
Acknowledgements
S.P.P. acknowledges an Oak Ridge National Laboratory (ORNL) appointment through the ORNL Postdoctoral Research Associates Program. Dr. Julian Philips (Thermo Fisher Scientific) is thanked for the loan of the LCQ DECA mass spectrometer. The MicroIonSpray head used to fabricate the DESI emitter was provided through a cooperative research and development agreement (CRADA) with MDS Sciex (ORNL02-0662). This research was supported by the Battelle Memorial Institute Technology Maturation Fund. The surface scanning platform and associated control software used in this study was developed with support from ORNL Technology Transfer and Economic Development (TTED) Royalty Funds. ORNL is managed and operated by UT-Battelle, LLC, for the United States Department of Energy under contract DE-AC05-00OR22725.
Author information
Authors and Affiliations
Corresponding author
Additional information
This manuscript has been authored by a contractor of the US Government under contract DE-AC05-00OR22725. Accordingly, the US Government retains a paid-up, nonexclusive, irrevocable, worldwide license to publish or reproduce the published form of this contribution, prepare derivative works, distribute copies to the public, and perform publicly and display publicly, or allow others to do so, for US Government purposes.
Rights and permissions
About this article
Cite this article
Pasilis, S.P., Kertesz, V., Van Berkel, G.J. et al. Using HPTLC/DESI-MS for peptide identification in 1D separations of tryptic protein digests. Anal Bioanal Chem 391, 317–324 (2008). https://doi.org/10.1007/s00216-008-1874-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-008-1874-6