Abstract
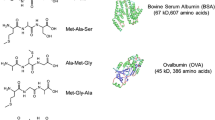
Hydrophobic-interaction chromatography coupled on-line with chemical-vapor-generation atomic-fluorescence spectrometry (HIC–CVGAFS), optimized recently for the analysis of thiol-containing proteins under denaturing conditions, has been used to study the chemical reduction of denatured proteins. Four proteins chosen as models (human serum albumin (HSA), bovine serum albumin (BSA), α-lactalbumin (α-Lac) from bovine milk, and lysozyme from chicken egg (Lys)) were denatured with urea and reduced with dithiothreitol (DTT), with selenol as catalyst. The method is based on derivatization of the –SH groups of proteins with p-hydroxymercurybenzoate (PHMB), followed by HIC separation and post-column on-line reaction of the derivatized reduced, denatured proteins with bromine generated in situ. HgII, derived from rapid conversion of uncomplexed and protein-complexed PHMB, is selectively detected by AFS in an Ar/H2 miniaturized flame after sodium borohydride (NaBH4) reduction to Hg°. The yield of the reduction was studied as a function of reductant concentration, reduction time (tred), and urea concentration. Results showed that the optimum values for DTT and selenol concentrations and for tred were between 1 and 100 mmol L−1 and between 1 and 20 min, respectively, depending on the protein studied. The percentage disulfide bond reduction increases as the urea concentration used for protein denaturation increases, giving a single-step sigmoid increment for single-domain, low-MW proteins (α-Lac and Lys), and a two-step sigmoid increment for multi-domain, high MW proteins (HSA and BSA). The shapes of plots of percentage reduced disulfide against urea concentration are characteristic of each protein and are correlated with the location of S–S in the protein. Under the adopted conditions complete protein denaturation is the conditio sine qua non for obtaining 100% S–S reduction. The detection limit for denatured, reduced proteins examined under the optimized conditions was found to be in the range 1–5×10−12 mol L−1 (10–30 pg), depending on the protein considered.





Similar content being viewed by others
References
Wedemeyer WJ, Welker E, Narayan M, Scheraga HA (2000) Biochemistry 39:4207–4216
Creighton TE (1986) Methods Enzymol 131:83–106
Gekko K, Kimoto A, Kamiyama T (2003) Biochemistry 42:13746–13753
Edwin F, Jagannadham MV (2000) Biochim Biophys Acta 1479:69–82
Tachibana H, Ohta K, Sawano H, Koumoto Y, Segawa S (1994) Biochemistry 33:15008–15016
Chang Y, Schindler P, Chatrenet B (1995) J Biol Chem 270:11992–11997
Darby JNJ, Morin PE, Talbo G, Creighton TE (1995) J Mol Biol 249:463–477
Takeda K, Ogawa K, Ohara M, Hamada S, Moriyama Y (1995) J Protein Chem 14:679–684
Haniu M, Hui J, Young Y, Le J, Katta V, Lee R, Shimamoto G, Rohde MF (1996) Biochemistry 35:16799–16805
Onda M, Tatsumi E, Takahashi N, Hirose M (1997) J Biochem 122:83–89
Xu X, Scheraga HA (1998) Biochemistry 37:7561–7571
Hober S, Uhlen M, Nilsson B (1997) Biochemistry 36:4616–4622
Kumar N, Kella D, Kinsella JE (1985) J Biochem Biophys Met 11:251–263
Cayot P, Rosiers H, Roullier L, Haertlé T, Tainturier G (2002) Food Chem 77:309–315
Kuwajima K, Ikeguchi M, Sugawara T, Hiraoka Y, Sugai S (1990) Biochemistry 29:8240–8249
Cayot P, Lorient D (1997) Structure–function relationships of whey proteins (Chap 8). In: Damodaran S, Paraf A (eds) Food proteins and their applications, pp 225–256
Wolf WJ (1993) J Agric Food Chem 41:168–176
Mangino ME, Harper WJ (1996) Factors affecting application of native and modified proteins in food product. In: Magdassi S (ed) Surface activity of proteins: chemical and physicochemical modifications. Marcel Dekker, New York, pp 285–322
Honeychurch MJ (1997) Bioelectrochem Bioeng 44:13–21
Raspi G, Lo Moro A, Spinetti M, Tesi G (1997) Anal Commun 34:307–309
Means GE, Feeney RE (1971) Chemical modification of proteins. Holden Day, San Francisco
Fontana A, Toniolo C (1974) Detection and determination of thiols. In: Patai S (ed) The chemistry of the thiol group, Wiley, New York, pp 272–319
Jocelyn P (1987) Methods Enzymol 143:246–256
DeCollo TV, Lees WJ (2001) J Org Chem 66:4244–4249
Ivanov AR, Nazimov IV, Baratova LA (2000) J Chromatogr A 870:433–442
Burmeister Getz E, Xiao M, Chakrabarty T, Cooke R, Selvin PR (1999) Anal Biochem 273:73–80
Singh R, Whitesides GM (1991) J Org Chem 56:2332–2337
Singh R, Whitesides GM (1991) J Org Chem 56:6931–6933
Singh R, Kats L (1995) Anal Biochem 232:86–91
Bramanti E, Lucchesini S, D’Ulivo A, Lampugnani L, Zamboni R, Spinetti M, Raspi G (2001) J Anal Atom Spectrom 16:166–171
Bramanti E, Lomonte C, Onor M, Zamboni R, Raspi G, D’Ulivo A (2004) Talanta 63:383–389
Ando Y, Steiner M (1973) Biochim Biophys Acta 311:38–44
Pace CN, Shirley BA, Thompson JA (1989) Protein structure: a practical approach, Creighton TE (ed) IRL Press, Oxford, pp 311–330
Wallevik K (1973) J Biol Chem 245:2650–2655
Razumovsky L, Damodaran S (1999) Langmuir 15:1392–1399
D’Ulivo A, Bramanti E, Lampugnani L, Zamboni R (2001) Spectrochim Acta B 56:1893–1907
Malmsten M (1998) J Colloid Interface Sci 207:186–199
Gu Z, Su Z, Janson J-C (2001) J Chromatogr A 918:311–318
Creigthon TE (1993) Proteins structures and molecular properties. Freeman VH and Company, New York
Khan MY (1986) Biochem J 236:307–310
Peters T Jr (1985) Adv Protein Chem 37:161–247
Ahmad N, Qasim MA (1995) Eur J Biochem 227:563–565
Sugai S, Ikeguchi M (1994) Adv Biophys 30:37–84
Pardon E, Haezebrouck P, De Baetselier A, Hooke SD, Fancourt KT, Desmet J, Dobson CM, Van Dael H, Joniau M (1995) J Biol Chem 270:10514–10524
Chang JY, Li L (2002) FEBS Letters 511:73–78
Gilbert HF (1994) Mechanism of protein folding. In: Pain RH (ed), Oxford University Press, Cambridge
Farruggia B, Picò GA (1999) Int J Biol Macromol 26:317–323
Ahmad F, Bigelow CC (1982) J Biol Chem 257:12935–12938
Green RF, Pace CN (1974) J Biol Chem 249:5388–5393
Acknowledgments
This work was financially supported by CNR and Ambiente s.c.r.L. (Carrara, MS, Italy; DOCUP 2000–2006, Regione Toscana). The authors would like to thank ThermoQuest, for providing part of the instrumentation, and Mrs M. Cempini, C. Lanza and M.C.M. Mascherpa for their technical support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bramanti, E., Lomonte, C., Onor, M. et al. Study of the disulfide reduction of denatured proteins by liquid chromatography coupled with on-line cold-vapor-generation atomic-fluorescence spectrometry (LC–CVGAFS). Anal Bioanal Chem 380, 310–318 (2004). https://doi.org/10.1007/s00216-004-2746-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-004-2746-3