Abstract
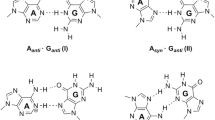
DNA G-Quadruplex is highly sensitive to oxidation as their structures include \(\pi\)-stacked guanine quartets allowing fast hole transfer between the nucleobases. These transfers can be described using vertical energy gap and electronic coupling between the different diabatic states at play in a guanine pair. Using classical molecular dynamics simulation and the constrained DFT/MM implementation in deMon2k, we determine these quantities for all the interacting guanine pairs of six G-quadruplex structures including one to four quartets and corresponding to different DNA folding. We then described an uni-directional transfer within a quartet, with high electronic coupling and vertical energy gap values, which contrasts with the hole transfer between \(\pi\)-stacked guanine, bi-directional and corresponding to smaller charge transfer parameters. The influence of the geometrical parameters on the electronic coupling is explored, while the external or internal position of the guanine may impact its oxidation probability according to the vertical energy gaps.





Similar content being viewed by others
References
Marchand A, Gabelica V (2016) Folding and misfolding pathways of G-quadruplex DNA. Nucl Acids Res 44(22):10999–11012
Moyzis RK, Buckingham JM, Cram LS, Dani M, Deaven LL, Jones MD, Meyne J, Ratliff RL, Wu JR (1988) A highly conserved repetitive dna sequence, (ttaggg)n, present at the telomeres of human chromosomes. Proc Natl Acad Sci 85(18):6622–6626
Bryan TM (2020) G-quadruplexes at telomeres: friend or foe? Molecules 25(16):3686
Armanios M, Blackburn EH (2012) The telomere syndromes. Nat Rev Genet 13(10):693–704
Artandi SE, DePinho RA (2010) Telomeres and telomerase in cancer. Carcinogenesis 31(1):9–18. https://doi.org/10.1093/carcin/bgp268
Balasubramanian S, Hurley LH, Neidle S (2011) Targeting G-quadruplexes in gene promoters: a novel anticancer strategy? Nat Rev Drug Discov 10(4):261–275
Neidle S (2017) Quadruplex nucleic acids as targets for anticancer therapeutics. Nat Rev Chem 1(5):1–10
Harris LM, Merrick CJ (2015) G-quadruplexes in pathogens: a common route to virulence control? PLOS Pathogens 11(2):1–15. https://doi.org/10.1371/journal.ppat.1004562
Cadet J, Davies KJA, Medeiros MH, Di Mascio P, Wagner JR (2017) Formation and repair of oxidatively generated damage in cellular DNA. Free Radical Biol Med 107:13–34
Fleming AM, Burrows CJ (2013) G-quadruplex folds of the human telomere sequence alter the site reactivity and reaction pathway of guanine oxidation compared to duplex DNA. Chem Res Toxicol 26(4):593–607
Balanikas E, Banyasz A, Baldacchino G, Markovitsi D (2019) Populations and dynamics of guanine radicals in DNA strands-direct versus indirect generation. Molecules 24(13):2347
Banyasz A, Balanikas E, Martinez-Fernandez L, Baldacchino G, Douki T, Improta R, Markovitsi D (2019) Radicals generated in tetramolecular guanine quadruplexes by photoionization: spectral and dynamical features. J Phys Chem B 123(23):4950–4957
Zhang X, Jie J, Song D, Su H (2020) Deprotonation of guanine radical cation G\(\cdot +\) mediated by the protonated water cluster. J Phys Chem A 124(29):6076–6083. https://doi.org/10.1021/acs.jpca.0c03748
Perrier S, Hau J, Gasparutto D, Cadet J, Favier A, Ravanat JL (2006) Characterization of Lysine–Guanine cross-links upon one-electron oxidation of a guanine-containing oligonucleotide in the presence of a Trilysine peptide. J Am Chem Soc 128(17):5703–5710. https://doi.org/10.1021/ja057656i
Martinez-Fernandez L, Changenet P, Banyasz A, Gustavsson T, Markovitsi D, Improta R (2019) Comprehensive study of guanine excited state relaxation and photoreactivity in G-quadruplexes. J Phys Chem Lett 10(21):6873–6877
Banyasz A, Martínez-Fernández L, Balty C, Perron M, Douki T, Improta R, Markovitsi D (2017) Absorption of low-energy UV radiation by human telomere G-quadruplexes generates long-lived guanine radical cations. J Am Chem Soc 139(30):10561–10568. https://doi.org/10.1021/jacs.7b05931
Delaney S, Barton JK (2003) Charge transport in DNA duplex/quadruplex conjugates. Biochemistry 42(48):14159–14165. https://doi.org/10.1021/bi0351965
Champdoré Md, Napoli LD, Montesarchio D, Piccialli G, Caminal C, Mulazzani QG, Navacchia ML, Chatgilialoglu C (2004) Excess electron transfer in G-quadruplex. Chem Commun 15:1756–1757
Gervasio FL, Laio A, Iannuzzi M, Parrinello M (2004) Influence of DNA structure on the reactivity of the guanine radical cation. Chem Eur J 10(19):4846–4852. https://doi.org/10.1002/chem.200400171
Martínez-Fernández L, Banyasz A, Markovitsi D, Improta R (2018) Topology controls the electronic absorption and delocalization of electron holes in guanine quadruplexes. Chem Eur J 24(57):15185–15189
Marcus RA, Sutin N (1985) Electron transfers in chemistry and biology. Biochim Biophys Acta (BBA) Rev Bioenerge 811(3):265–322
Lech CJ, Phan AT, Michel-Beyerle ME, Voityuk AA (2013) Electron–hole transfer in G-quadruplexes with different tetrad stacking geometries: a combined QM and MD study. J Phys Chem B 117(34):9851–9856. https://doi.org/10.1021/jp404788t
Woiczikowski PB, Kubař T, Gutiérrez R, Cuniberti G, Elstner M (2010) Structural stability versus conformational sampling in biomolecular systems: Why is the charge transfer efficiency in G4-DNA better than in double-stranded DNA? J Chem Phys 133(3):035103. https://doi.org/10.1063/1.3460132
Lim KW, Amrane S, Bouaziz S, Xu W, Mu Y, Patel DJ, Luu KN, Phan AT (2009) Structure of the human telomere in K+ solution: a stable basket-type G-quadruplex with only two G-tetrad layers. J Am Chem Soc 131(12):4301–4309
Wang Y, Patel DJ (1993) Solution structure of the human telomeric repeat d[AG3(T2AG3)3] G-tetraplex. Structure 1(4):263–282
Ambrus A, Chen D, Dai J, Jones RA, Yang D (2005) Solution structure of the biologically relevant G-quadruplex element in the human c-MYC promoter. Implications for G-quadruplex stabilization. Biochemistry 44(6):2048–2058. https://doi.org/10.1021/bi048242p
Luu KN, Phan AT, Kuryavyi V, Lacroix L, Patel DJ (2006) Structure of the human telomere in K+ solution: an intramolecular (3 + 1) G-quadruplex scaffold. J Am Chem Soc 128(30):9963–9970. https://doi.org/10.1021/ja062791w
Winnerdy FR, Bakalar B, Maity A, Vandana JJ, Mechulam Y, Schmitt E, Phan AT (2019) NMR solution and X-ray crystal structures of a DNA molecule containing both right- and left-handed parallel-stranded G-quadruplexes. Nucl Acids Res 47(15):8272–8281. https://doi.org/10.1093/nar/gkz349
Bakalar B, Heddi B, Schmitt E, Mechulam Y, Phan AT (2019) A minimal sequence for left-handed G-quadruplex formation. Angew Chem Int Ed 58(8):2331–2335. https://doi.org/10.1002/anie.201812628
Wu Q, Van Voorhis T (2005) Direct optimization method to study constrained systems within density-functional theory. Phys Rev A 72(2):024502
Wu Q, Van Voorhis T (2006) Direct calculation of electron transfer parameters through constrained density functional theory. J Phys Chem A 110(29):9212–9218. https://doi.org/10.1021/jp061848y
Salahub DR, Noskov SY, Lev B, Zhang R, Ngo V, Goursot A, Calaminici P, Köster AM, Alvarez-Ibarra A, Mejía-Rodríguez D, Řezáč J, Cailliez F, De la Lande A (2015) QM/MM calculations with deMon2k. Molecules 20(3):4780–4812
Humphrey W, Dalke A, Schulten K (1996) VMD—visual molecular dynamics. J Mol Graph 14:33–38
Case DA, Belfon K, Ben-Shalom IY, Brozell SR, Cerutti DS, Cheatham TE, Cruzeiro VWD, Darden TA, Duke RE, Giambasu G, Gilson MK, Gohlke H, Goetz AW, Harris R, Izadi S, Ismailov SA, Kasavajhala K, Kovalenko A, Krasny R, Kurtzman T, Lee T, LeGrand S, Li P, Lin C, Liu J, Luchko T, Luo R, Man V, Merz KM, Miao Y, Mikhailovskii O, Monard G, Nguyen H, Onufriev A, Pan F, Pantano S, Qi R, Roe DR, Roitberg A, Sagui C, Schott-Verdugo S, Shen J, Simmerling C, Skrynnikov NR, Smith J, Swails JM, Walker RC, Wang J, Wilson L, Wolf RM, Wu X, Xiong Y, Xue Y, York DM, Kollman PA (2020) AMBER 2020. University of California, San Francisco
Ivani I, Dans PD, Noy A, Pérez A, Faustino I, Hospital A, Walther J, Andrio P, Goñi R, Balaceanu A, Portella G, Battistini F, Gelpí JL, González C, Vendruscolo M, Laughton CA, Harris SA, Case DA, Orozco M (2016) Parmbsc1: a refined force field for DNA simulations. Nat Methods 13(1):55–58
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103(19):8577–8593
Roe DR, Cheatham TE (2013) Ptraj and cpptraj: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput 9(7):3084–3095
Köster AM, Calaminici P, Casida ME, Flores-Moreno R, Greudtner G, Goursot A, Heine T, Ipatov A, Janetzko F, del Campo JM, Patchkovskii S, Reveles JU, Vela A, Zuniga B, Salahub D (2006) demon2K. deMon developers. Cinvestav, Mexico City
de la Lande A, Salahub DR (2010) Derivation of interpretative models for long range electron transfer from constrained density functional theory. J Mol Struct THEOCHEM 943(1):115–120
Adamo C, Barone V (1999) Toward reliable density functional methods without adjustable parameters: the PBE0 model. J Chem Phys 110(13):6158–6170. https://doi.org/10.1063/1.478522
Kendall RA, Dunning TH Jr, Harrison RJ (1992) Electron affinities of the first-row atoms revisited. Systematic basis sets and wave functions. J Chem Phys 96(9):6796–6806. https://doi.org/10.1063/1.462569
Grimme S (2006) Semiempirical GGA-type density functional constructed with a long-range dispersion correction. J Comput Chem 27(15):1787–1799. https://doi.org/10.1002/jcc.20495
Gillet N, Berstis L, Wu X, Gajdos F, Heck A, de la Lande A, Blumberger J, Elstner M (2016) Electronic coupling calculations for bridge-mediated charge transfer using constrained density functional theory (CDFT) and effective hamiltonian approaches at the density functional theory (DFT) and fragment-orbital density functional tight binding (FODFTB) level. J Chem Theory Comput 12(10):4793–4805. https://doi.org/10.1021/acs.jctc.6b00564
Calaminici P, Janetzko F, Köster AM, Mejia-Olvera R, Zuniga-Gutierrez B (2007) Density functional theory optimized basis sets for gradient corrected functionals: 3d transition metal systems. J Chem Phys 126(4):044108–10. https://doi.org/10.1063/1.2431643
Bultinck P, Ayers PW, Fias S, Tiels K, Van Alsenoy C (2007) Uniqueness and basis set dependence of iterative Hirshfeld charges. Chem Phys Lett 444(1–3):205–208
Wu Q, Van Voorhis T (2006) Extracting electron transfer coupling elements from constrained density functional theory. J Chem Phys 125(16):164105. https://doi.org/10.1063/1.2360263
Blumberger J, Lamoureux G (2008) Reorganization free energies and quantum corrections for a model electron self-exchange reaction: comparison of polarizable and non-polarizable solvent models. Mol Phys 106(12–13):1597–1611. https://doi.org/10.1080/00268970802220112
de la Lande A, Alvarez-Ibarra A, Hasnaoui K, Cailliez F, Wu X, Mineva T, Cuny J, Calaminici P, Löpez-Sosa L, Geudtner G, Navizet I, Garcia Iriepa C, Salahub DR, Köster AM (2019) Molecular simulations with in-demon2k qm/mm, a tutorial-review. Molecules 24(9):1653
Acknowledgements
R.R. is grateful for the IISER-ENS de Lyon fellowship which has financed her visit at the ENS de Lyon. The authors thank the SYSPROD Project and AXELERA Pôle de Compétitivité for financial support (PSMN Data Center). This work was performed using HPC resources from GENCI-IDRIS (Grant 2020-A0070800609). N. G. thanks A. de la Lande for providing the deMon2k version with the last cDFT and QM/MM modules and his kind explanations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published as part of the special collection of articles “20th deMon Developers Workshop”.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ravindranath, R., Mondal, P. & Gillet, N. Radical cation transfer in a guanine pair: an insight to the G-quadruplex structure role using constrained DFT/MM. Theor Chem Acc 140, 89 (2021). https://doi.org/10.1007/s00214-021-02787-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00214-021-02787-0