Abstract
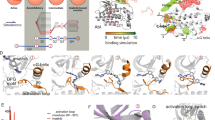
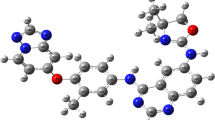
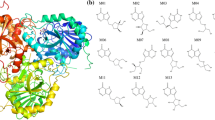
Nilotinib is a novel anticancer drug, which specifically binds to the Abl kinase and blocks its signaling activity. In order to model the nilotinib/protein interactions, we have developed a molecular mechanics force field for nilotinib, consistent with the CHARMM force field for proteins and nucleic acids. Atomic charges were derived by utilizing a supermolecule ab initio approach. We considered the ab initio energies and geometries of a probe water molecule that interacts with nilotinib fragments at six different positions. We investigated both neutral and protonated states of nilotinib. The final rms deviation between the ab initio and the force field energies, averaged over both forms, was equal 0.2 kcal/mol. The model reproduces the ab initio geometry and flexibility of nilotinib. To apply the force field to nilotinib/Abl simulations, it is also necessary to determine the most likely protein and nilotinib protonation state when it binds to Abl. This task was carried out using molecular dynamics free energy simulations. The simulations indicate that nilotinib can interact with Abl in protonated and deprotonated forms, with the protonated form more favoured for the interaction. In the course of our calculations, we established that the His361, a titratable amino acid residue that mediates the interaction, prefers to be neutral. These insights and models should be of interest for drug design.






Similar content being viewed by others
References
Gambacorti-Passerini CB, Gunby RH, Piazza R, Galietta A, Rostagno R, Scapozza L (2003) Lancet Oncol 4:75–85
Levitski A (1996) Curr Opin Cell Biol 8:239–244
Chen J, Zhang X, Fernandez A (2007) Curr Drug Targets 7:1443–1454
Aleksandrov A, Simonson T (2010) J Biol Chem 285:13807–13815
Nagar B, Hantschel O, Young M, Scheffzek K, Veach D, Bornmann W, Clarkson B, Superti-Furga G, Kuriyan J (2003) Cell 112:859–871
Deininger M, Buchdunger E, Druker BJ (2005) Blood 105:2640–2653
Vajpai N, Strauss A, Fendrich G, Cowan-Jacob S, Manley P, Grzesiek S, Jahnke W (2008) J Biol Chem 283:18292–18302
Weisberg E, Manley P, Mestan J, Cowan-Jacob S, Ray A, Griffin JD (2006) Br J Cancer 94:1765–1769
DeRemer DL, Ustun C, Natarajan K (2008) Clin Ther 30:1956–1975
Giles F, Rosti G, Beris P, Clark R, le Coutre P, Mahon F, Steegmann J, Valent P, Saglio G (2010) Exp Rev Hematol 3:665–673
Weisberg E et al (2005) Cancer Cell 7:129–141
Mackerell A et al (1998) J Phys Chem B 102:3586–3616
Mackerell A, Wiorkiewicz-Kuczera J, Karplus M (1995) J Am Chem Soc 117:11946–11975
Jorgensen W, Chandrasekar J, Madura J, Impey R, Klein M (1983) J Chem Phys 79:926–935
Trylska J, Antosiewicz J, Geller M, Hodge C, Klabe R, Head M, Gilson M (1999) Prot Sci 8:180–195
Aleksandrov A, Proft J, Hinrichs W, Simonson T (2007) ChemBioChem 8:675–685
Donnini S, Villa A, Groenhof G, Mark AE, Wierenga RK, Juffer A (2009) Proteins 76:138–150
Sham Y, Chu Z, Warshel A (1997) J Phys Chem B 101:4458–4472
Simonson T, Carlsson J, Case DA (2004) J Am Chem Soc 126:4167–4180
Archontis G, Simonson T (2005) Biophys J 88:3888–3904
Aleksandrov A, Simonson T (2006) J Comp Chem 27:1517–1533
Aleksandrov A, Simonson T (2009) J Comp Chem 30:243–255
Aleksandrov A, Simonson T (2010) J Comp Chem 31: 1550–1560
Foloppe N, MacKerell A (2000) J Comp Chem 21:86–104
Beglov D, Roux B (1994) J Chem Phys 100:9050–9063
Simonson T (2000) J Phys Chem B 104:6509–6513
Stote R, States D, Karplus M (1991) J Chem Phys 88:2419–2433
Brooks B, Bruccoleri R, Olafson B, States D, Swaminathan S, Karplus M (1983) J Comp Chem 4:187–217
Szakacs Z, Beni S, Varga Z, Orfi L, Keri G, Noszal B (2005) J Med Chem 48:249–255
Vaughan JD, Vaughan VL, Daly SS, Smith WA (1980) J Org Chem 45:3108–3111
Bruice T, Schmir G (1958) J Am Chem Soc 80:148–156
Simonson T (2001) Free energy calculations. In: Becker O, Mackerell A Jr, Roux B, Watanabe M (eds) Computational biochemistry & biophysics, Chap. 9. Marcel Dekker, New York
Thompson D, Plateau P, Simonson T (2006) ChemBioChem 7:337–344
Simonson T, Archontis G, Karplus M (2002) Acc Chem Res 35:430–437
Hodel A, Simonson T, Fox RO, Brünger AT (1993) J Phys Chem 97:3409–3417
Reinhardt W, Miller M, Amon L (2001) Acc Chem Res 34:607–614
Shirts MR, Pitera JW, Swope WC, Pande VS (2003) J Chem Phys 119:5740–5761
Hermans J (1991) J Phys Chem 95:9029–9032
Wood RH (1991) J Phys Chem 95:4838–4842
Hummer G (2001) J Chem Phys 114:7330–7337
Thompson D, Simonson T (2006) J Biol Chem 281:23792–23803
Aleksandrov A, Simonson T (2008) Biochemistry 47:13594–13603
Eberini I, Baptista A, Gianazza E, Fraternali F, Beringhelli T (2004) Proteins 54:744–758
Simonson T, Brünger AT (1994) J Phys Chem 98:4683–4694
Mackerell A Jr (2001) In: Becker O, Mackerell A Jr, Roux B, Watanabe M (eds) Computational biochemistry & biophysics, Chap. 1. Marcel Dekker, New York
Simonson T (2001) Curr Opin Struct Biol 11:243–252
Richarz R, Wuthrich K (1975) Biopolymers 17:2133–2141
Asaki T, Sugiyama Y, Hamamoto T, Higashioka M, Umehara M, Naito H, Niwa T (2006) Bioorg Med Chem Lett 16:1421–1425
Puttini M et al (2008) Haematologica 93:653–661
Acknowledgments
We thank Thomas Simonson for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Valeyev, N.V., Aleksandrov, A. An atomistic model for simulations of nilotinib and nilotinib/kinase binding. Theor Chem Acc 129, 747–756 (2011). https://doi.org/10.1007/s00214-011-0931-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00214-011-0931-y