Abstract
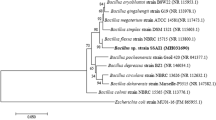
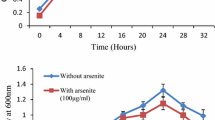
A novel arsenite resistant bacterial strain SSBW5 was isolated from the battery waste site of Corlim, Goa, India. This strain interestingly exhibited rapid arsenite oxidation with an accumulation of 5 mM arsenate within 24 h and a minimum inhibitory concentration (MIC) of 18 mM. The strain SSBW5 was identified as Paenarthrobacter nicotinovorans using 16S rDNA sequence analysis. Fourier-transformed infrared (FTIR) spectroscopy of arsenite-exposed cells revealed the interaction of arsenite with several important functional groups present on the cell surface, possibly involved in the resistance mechanism. Interestingly, the whole genome sequence analysis also clearly elucidated the presence of genes, such as GlpF, aioAB and aioE encoding transporter, arsenite oxidase and oxidoreductase enzyme, respectively, conferring their role in arsenite resistance. Furthermore, this strain also revealed the presence of several other genes conferring resistance to various metals, drugs, antibiotics and disinfectants. Further suggesting the probable direct or indirect involvement of these genes in the detoxification of arsenite thereby increasing its tolerance limit. In addition, clumping of bacterial cells was observed through microscopic analysis which could also be a strategy to reduce arsenite toxicity thus indicating the existence of multiple resistance mechanisms in strain SSBW5. In the present communication, we are reporting for the first time the potential of P. nicotinovorans strain SSBW5 to be used in the bioremediation of arsenite via arsenite oxidation along with other toxic metals and metalloids.





Similar content being viewed by others
Data availability
The data sets generated during and/or analyzed during the current study are available in the GenBank repository (MN640912 and SZVS00000000). All the data sets generated or analyzed during this study are included in this published article and its supplementary information files.
References
Aguilar NC, Faria MC, Pedron T, Batista BL, Mesquita JP, Bomfeti CA, Rodrigues JL (2020) Isolation and characterization of bacteria from a brazilian gold mining area with a capacity of arsenic bioaccumulation. Chemosphere 240:124871
Andres J, Bertin PN (2016) The microbial genomics of arsenic. FEMS Microbiol Rev 40(2):299–322
Antenozio ML, Giannelli G, Marabottini R, Brunetti P, Allevato E, Marzi D, Capobianco G, Bonifazi G, Serranti S, Visioli G, Stazi SR (2021) Phytoextraction efficiency of Pteris vittata grown on a naturally As-rich soil and characterization of As-resistant rhizosphere bacteria. Sci Rep 11(1):6794
Banerjee S, Datta S, Chattyopadhyay D, Sarkar P (2011) Arsenic accumulating and transforming bacteria isolated from contaminated soil for potential use in bioremediation. J Environ Sci Health A 46(14):1736–1747
Banerjee A, Sarkar S, Gorai S, Kabiraj A, Bandopadhyay R (2021) High arsenic tolerance in Brevundimonas aurantiaca PFAB1 from an arsenic-rich Indian hot spring. Electron J Biotechnol 53:1–7
Ben Fekih I, Zhang C, Li YP, Zhao Y, Alwathnani HA, Saquib Q, Rensing C, Cervantes C (2018) Distribution of arsenic resistance genes in prokaryotes. Front Microbiol 9:2473
Cai X, Zhang Z, Yin N, Du H, Li Z, Cui Y (2016) Comparison of arsenate reduction and release by three As (V)-reducing bacteria isolated from arsenic-contaminated soil of Inner Mongolia, China. Chemosphere 161:200–207
Ceballos-Escalera A, Pous N, Chiluiza-Ramos P, Korth B, Harnisch F, Bañeras L, Balaguer MD, Puig S (2021) Electro-bioremediation of nitrate and arsenite polluted groundwater. Water Res 190:116748
Chang JS (2015) Biotransformation of arsenite and bacterial aox activity in drinking water produced from surface water of floating houses: arsenic contamination in Cambodia. Environ Pollut 206:315–323
Crognale S, Amalfitano S, Casentini B, Fazi S, Petruccioli M, Rossetti S (2017) Arsenic-related microorganisms in groundwater: a review on distribution, metabolic activities and potential use in arsenic removal processes. Rev Environ Sci Biotechnol 16:647–665
Debiec-Andrzejewska K, Krucon T, Piatkowska K, Drewniak L (2020) Enhancing the plants growth and arsenic uptake from soil using arsenite-oxidizing bacteria. Environ Pollut 264:114692
Haghi M, Diznabi SH, Karaboz I, Omeroglu EE (2023) Arsenic pollution and arsenic-resistant bacteria of drying Urmia Salt Lake. Front Environ Sci 11:1195643
Han YH, Jia MR, Liu X, Zhu Y, Cao Y, Chen DL, Chen Y, Ma LQ (2017) Bacteria from the rhizosphere and tissues of As-hyperaccumulator Pteris vittata and their role in arsenic transformation. Chemosphere 186:599–606
Hartl FU (1996) Molecular chaperones in cellular protein folding. Nature 381(6583):571–580
Imran M, Das KR, Naik MM (2019) Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: an emerging health threat. Chemosphere 215:846–857
Jebelli MA, Maleki A, Amoozegar MA, Kalantar E, Gharibi F, Darvish N, Tashayoe H (2018) Isolation and identification of the native population bacteria for bioremediation of high levels of arsenic from water resources. J Environ Manag 212:39–45
Ji Y, Wang L, Jiang M, Lu J, Ferronato C, Chovelon JM (2017) The role of nitrite in sulfate radical-based degradation of phenolic compounds: an unexpected nitration process relevant to groundwater remediation by in-situ chemical oxidation (ISCO). Water Res 123:249–257
Kabiraj A, Biswas R, Halder U, Bandopadhyay R (2022) Bacterial arsenic metabolism and its role in arsenic bioremediation. Curr Microbiol 79(5):1–5
Kashyap DR, Botero LM, Lehr C, Hassett DJ, McDermott TR (2006) A Na+: H+ antiporter and a molybdate transporter are essential for arsenite oxidation in Agrobacterium tumefaciens. J Bacteriol 188(4):1577–1584
Khairul I, Wang QQ, Jiang YH, Wang C, Naranmandura H (2017) Metabolism, toxicity and anticancer activities of arsenic compounds. Oncotarget 8(14):23905
Khanna K, Kohli SK, Kumar P, Ohri P, Bhardwaj R, Alam P, Ahmad P (2022) Arsenic as hazardous pollutant: perspectives on engineering remediation tools. Sci Total Environ 838:155870
Khoei AJ, Joogh NJ, Darvishi P, Rezaei K (2018) Application of physical and biological methods to remove heavy metal, arsenic and pesticides, malathion and diazinon from water. Turk J Fish Aquat Sci 19(1):21–28
Koechler S, Cleiss-Arnold J, Proux C, Sismeiro O, Dillies MA, Goulhen-Chollet F, Hommais F, Lièvremont D, Arsène-Ploetze F, Coppée JY, Bertin PN (2010) Multiple controls affect arsenite oxidase gene expression in Herminiimonas arsenicoxydans. BMC Microbiol 10(1):53
Kognou AL, Chio C, Khatiwada JR, Shrestha S, Chen X, Zhu Y, Ngono Ngane RA, Agbor Agbor G, Jiang ZH, Xu CC, Qin W (2023) Characterization of potential virulence, resistance to antibiotics and heavy metals, and biofilm-forming capabilities of soil lignocellulolytic bacteria. Microb Physiol 33(1):36–48
Koutros S, Lenz P, Hewitt SM, Kida M, Jones M, Schned AR, Baris D, Pfeiffer R, Schwenn M, Johnson A, Karagas MR (2018) RE: Elevated bladder cancer in northern new England: the role of drinking water and arsenic. JNCI Nat Cancer Inst 110(11):1273–1274
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Laha A, Bhattacharyya S, Sengupta S, Bhattacharyya K, GuhaRoy S (2021) Investigation of arsenic-resistant, arsenite-oxidizing bacteria for plant growth promoting traits isolated from arsenic contaminated soils. Arch Microbiol 7:4677–4692
Lemire JA, Harrison JJ, Turner RJ (2013) Antimicrobial activity of metals: mechanisms, molecular targets and applications. Nat Rev Microbiol 11(6):371–384
Lescure T, Joulian C, Charles C, Saanda TB, Charron M, Breeze D, Bauda P, Battaglia-Brunet F (2020) Simple or complex organic substrates inhibit arsenite oxidation and aioA gene expression in two β-Proteobacteria strains. Res Microbiol 171(1):13–20
Lett MC, Paknikar KM, Lievremont D (2001) A simple and rapid method for arsenite and arsenate speciation. Process Metall 11:541–546
Li X, Gu AZ, Zhang Y, Xie B, Li D, Chen J (2019) Sub-lethal concentrations of heavy metals induce antibiotic resistance via mutagenesis. J Hazard Mater 369:9–16
Liu S, Zhang F, Chen J, Sun G (2011) Arsenic removal from contaminated soil via biovolatilization by genetically engineered bacteria under laboratory conditions. J Environ Sci 23(9):1544–1550
Lloyd NA, Nazaret S, Barkay T (2018) Whole genome sequences to assess the link between antibiotic and metal resistance in three coastal marine bacteria isolated from the mummichog gastrointestinal tract. Mar Pollut Bull 135:514–520
Lupo A, Coyne S, Berendonk TU (2012) Origin and evolution of antibiotic resistance: the common mechanisms of emergence and spread in water bodies. Front Microbiol 3:18
Ma B, Song W, Zhang X, Chen M, Li J, Yang X, Zhang L (2023) Potential application of novel cadmium-tolerant bacteria in bioremediation of Cd-contaminated soil. Ecotoxicol Environ Saf 255:114766
Mahtani S, Mavinkurve S (1979) Microbial purification of longifolene-a sesquiterpene. J Ferment Technol 57:529–533
Mandal BK, Suzuki KT (2002) Arsenic round the world: a review. Talanta 58(1):201–235
Mandal D, Sonar R, Saha I, Ahmed S, Basu A (2021) Isolation and identification of arsenic resistant bacteria: a tool for bioremediation of arsenic toxicity. Int J Environ Sci Technol 19:9883–9900
Meng YL, Liu Z, Rosen BP (2004) As(III) and Sb(III) uptake by GlpF and efflux by ArsB in Escherichia coli. J Biol Chem 279(18):18334–18341
Miranda-Carrazco A, Vigueras-Cortés JM, Villa-Tanaca L, Hernández-Rodríguez C (2018) Cyanotrophic and arsenic oxidizing activities of Pseudomonas mendocina P6115 isolated from mine tailings containing high cyanide concentration. Arch Microbiol 200(7):1037–1048
Mujawar SY, Shamim K, Vaigankar DC, Dubey SK (2019) Arsenite biotransformation and bioaccumulation by Klebsiella pneumoniae strain SSSW7 possessing arsenite oxidase (aioA) gene. Biometals 32(1):65–76
Mujawar SY, Vaigankar DC, Dubey SK (2021) Biological characterization of Bacillus flexus strain SSAI1 transforming highly toxic arsenite to less toxic arsenate mediated by periplasmic arsenite oxidase enzyme encoded by aioAB genes. Biometals 34(4):895–907
Mukhopadhyay R, Rosen BP, Phung LT, Silver S (2002) Microbial arsenic: from geocycles to genes and enzymes. FEMS Microbiol Rev 26(3):311–325
Mukhopadhyay R, Bhattacharjee H, Rosen BP (2014) Aquaglyceroporins: generalized metalloid channels. Biochim Biophys Acta (BBA) 1840(5):1583–1591
Naumann D, Helm D, Labischinski H, Giesbrecht P (1991) The characterization of microorganisms by Fourier-transform infrared spectroscopy. In: Nelson WH (ed) Modern techniques for rapid microbiological analysis. VCH, New York, pp 43–96
Páez-Espino D, Tamames J, de Lorenzo V, Cánovas D (2009) Microbial responses to environmental arsenic. Biometals 22(1):117–130
Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DJ (2014) BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42(D1):D737–D743
Palma-Lara I, Martínez-Castillo M, Quintana-Pérez JC, Arellano-Mendoza MG, Tamay-Cach F, Valenzuela-Limón OL, García-Montalvo EA, Hernández-Zavala A (2020) Arsenic exposure: a public health problem leading to several cancers. Regul Toxicol Pharmacol 110:104539
Rahman S, Kim KH, Saha SK, Swaraz AM, Paul DK (2014) Review of remediation techniques for arsenic (As) contamination: a novel approach utilizing bio-organisms. J Environ Manag 134:175–185
Rathod J, Dhanani AS, Jean JS, Jiang WT (2019) The whole genome insight on condition-specific redox activity and arsenopyrite interaction promoting As-mobilization by strain Lysinibacillus sp. B2A1. J Hazard Mater 364:671–681
Raturi G, Chaudhary A, Rana V, Mandlik R, Sharma Y, Barvkar V, Salvi P, Tripathi DK, Kaur J, Deshmukh R, Dhar H (2023) Microbial remediation and plant-microbe interaction under arsenic pollution. Sci Total Environ 864:160972
Rosen BP (2002) Biochemistry of arsenic detoxification. FEBS Lett 529(1):86–92
Satyapal GK, Mishra SK, Srivastava A, Ranjan RK, Prakash K, Haque R, Kumar N (2018) Possible bioremediation of arsenic toxicity by isolating indigenous bacteria from the middle Gangetic plain of Bihar, India. Biotechnol Rep 17:117–125
Sher S, Hussain SZ, Cheema MT, Hussain A, Rehman A (2022) Efficient removal potential of Microbacterium sp. strain 1S1 against arsenite isolated from polluted environment. J King Saud Univ Sci 34(5):102066
Shi K, Wang Q, Wang G (2020) Microbial oxidation of arsenite: regulation, chemotaxis, phosphate metabolism and energy generation. Front Microbiol 11:569282
Silver S, Phung LT (1996) Bacterial heavy metal resistance: new surprises. Ann Rev Microbiol 50(1):753–789
Silver S, Phung LT (2005) Genes and enzymes involved in bacterial oxidation and reduction of inorganic arsenic. Appl Environ Microbiol 71(2):599–608
Singh N, Gupta S, Marwa N, Pandey V, Verma PC, Rathaur S, Singh N (2016) Arsenic mediated modifications in Bacillus aryabhattai and their biotechnological applications for arsenic bioremediation. Chemosphere 164:524–534
Sodhi KK, Kumar M, Agrawal PK, Singh DK (2019) Perspectives on arsenic toxicity, carcinogenicity and its systemic remediation strategies. Environ Technol Innov 16:100462
Studholme DJ, Jackson RA, Leak DJ (1999) Phylogenetic analysis of transformable strains of thermophilic Bacillus species. FEMS Microbiol Lett 172(1):85–90
Suhadolnik ML, Salgado AP, Scholte LL, Bleicher L, Costa PS, Reis MP, Dias MF, Ávila MP, Barbosa FA, Chartone-Souza E, Nascimento A (2017) Novel arsenic-transforming bacteria and the diversity of their arsenic-related genes and enzymes arising from arsenic-polluted freshwater sediment. Sci Rep 7(1):1–17
Sultana M, Vogler S, Zargar K, Schmidt AC, Saltikov C, Seifert J, Schlömann M (2012) New clusters of arsenite oxidase and unusual bacterial groups in enrichments from arsenic-contaminated soil. Arch Microbiol 194(7):623–635
Sun LN, Guo B, Lyu WG, Tang XJ (2020) Genomic and physiological characterization of an antimony and arsenite-oxidizing bacterium Roseomonas rhizosphaerae. Environ Res 191:110136
Teixeira P, Tacão M, Alves A, Henriques I (2016) Antibiotic and metal resistance in a ST395 Pseudomonas aeruginosa environmental isolate: a genomics approach. Mar Pollut Bull 110(1):75–81
Wang Q, Han Y, Shi K, Fan X, Wang L, Li M, Wang G (2017) An oxidoreductase AioE is responsible for bacterial arsenite oxidation and resistance. Sci Rep 7(1):1–10
WHO (2017) Guidelines for drinking-water quality: fourth edition incorporating the first addendum, 4th edn. World Health Organization, Geneva
Wu Y, Feng S, Li B, Mi X (2010) The characteristics of Escherichia coli adsorption of arsenic(III) from aqueous solution. World J Microbiol Biotechnol 26(2):249–256
Yang Z, Wu Z, Liao Y, Liao Q, Yang W, Chai L (2017) Combination of microbial oxidation and biogenic schwertmannite immobilization: a potential remediation for highly arsenic-contaminated soil. Chemosphere 181:1–8
Yuan C, Li P, Qing C, Kou Z, Wang H (2022) Different regulatory strategies of arsenite oxidation by two isolated Thermus tengchongensis strains from hot springs. Front Microbiol 13:817891
Zhao M, Zheng G, Kang X, Zhang X, Guo J, Zhang M, Zhang J, Chen Y, Xue L (2023) Arsenic pollution remediation mechanism and preliminary application of arsenic-oxidizing bacteria isolated from industrial wastewater. Environ Pollut 324:121384
Acknowledgements
Mujawar SY is grateful to University Grants Commission, New Delhi, for financial support as Senior Research Fellow (SRF). The authors are also thankful to Dr. B. R. Srinivasan and Mr. Rahul Kerkar from Department of Chemistry, Goa University for FTIR analysis.
Funding
This work was supported by University Grants Commission, New Delhi as Senior Research Fellow (SRF) [Award Letter No. F1-17.1/2015-16/MANF-2015-17-Goa-50641].
Author information
Authors and Affiliations
Contributions
Experimental design, data collection and analysis was performed by SYM. The first draft of the manuscript was written by SYM and all the authors provided their comments and suggestions on the previous versions of the manuscript. SKD mentored the experiments, corrected and thoroughly edited the manuscript. All the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mujawar, S.Y., Shamim, K., Vaigankar, D.C. et al. Rapid arsenite oxidation by Paenarthrobacter nicotinovorans strain SSBW5: unravelling the role of GlpF, aioAB and aioE genes. Arch Microbiol 205, 333 (2023). https://doi.org/10.1007/s00203-023-03673-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03673-y