Abstract
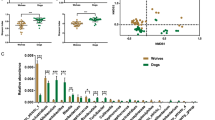
For different breeds of dogs with acute diarrhea, the gut microbiota and metabolome profiles are unclear. This prospective observational study analyzed the gut microbiomes of poodles with acute diarrhea and Labrador retrievers with acute diarrhea based on 16S amplicon sequencing, with respective healthy dogs as controls. Fecal non-target metabolomics and metagenomics were performed on poodles with acute diarrhea. This study found that the diversity and structure of the microbial community differed significantly between the two breeds in cohorts of healthy dogs. Two breeds of dogs with acute diarrhea demonstrated different changes in microbial communities and metabolic functions. The metabolism of starch and sucrose was significantly decreased in dogs with acute diarrhea, which may be attributed to the reduced activity of dextran dextrinase. Non-targeted metabolomics identified 21 abnormal metabolic pathways exhibited by dogs with acute diarrhea, including starch, amino acid, bile acid metabolism, etc., and were closely related to specific intestinal flora. This study provided new insights into breed specificity and the development of dietary treatment strategy in canine gastrointestinal disease.





Similar content being viewed by others
Data availability
The datasets generated in this study are all submitted to publicly available repositories. The 16 s and metagenomic high-throughput sequencing data generated in this study have been submitted to NCBI’s Sequence Read Archive repository with accession ID PRJNA917802.The metabolome data generated in this study have been submitted to MetaboLights database with the unique identifier MTBLS6823.
References
Begley M, Gahan CG, Hill C (2005) The interaction between bacteria and bile. FEMS Microbiol Rev 29(4):625–651. https://doi.org/10.1016/j.femsre.2004.09.003
Feng Q, Liang S, Jia H, Stadlmayr A, Tang L, Lan Z et al (2015) Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun 6:6528. https://doi.org/10.1038/ncomms7528
Ferreira-Halder CV, Faria AVS, Andrade SS (2017) Action and function of Faecalibacterium prausnitzii in health and disease. Best Pract Res Clin Gastroenterol 31(6):643–648. https://doi.org/10.1016/j.bpg.2017.09.011
Galler AI, Suchodolski JS, Steiner JM, Sung C-H, Hittmair KM, Richter B et al (2022) Microbial dysbiosis and fecal metabolomic perturbations in Yorkshire Terriers with chronic enteropathy. Sci Rep 12(1):12977. https://doi.org/10.1038/s41598-022-17244-6
Guard BC, Barr JW, Reddivari L, Klemashevich C, Jayaraman A, Steiner JM et al (2015) Characterization of microbial dysbiosis and metabolomic changes in dogs with acute diarrhea. PLoS ONE 10(5):e0127259. https://doi.org/10.1371/journal.pone.0127259
Guo Q, Goldenberg JZ, Humphrey C, El Dib R, Johnston BC (2019) Probiotics for the prevention of pediatric antibiotic-associated diarrhea. Cochrane Database Syst Rev 4(4):Cd004827. https://doi.org/10.1002/14651858.CD004827.pub5
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G et al (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21(3):494–504. https://doi.org/10.1101/gr.112730.110
Honneffer JB, Steiner JM, Lidbury JA, Suchodolski JS (2017) Variation of the microbiota and metabolome along the canine gastrointestinal tract. Metabolomics 13(3):26. https://doi.org/10.1007/s11306-017-1165-3
Huerta-Cepas J, Szklarczyk D, Heller D, Hernández-Plaza A, Forslund SK, Cook H et al (2019) eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucl Acids Res. 47(D1):D309–D314. https://doi.org/10.1093/nar/gky1085
Johnson JS, Spakowicz DJ, Hong BY, Petersen LM, Demkowicz P, Chen L et al (2019) Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat Commun 10(1):5029. https://doi.org/10.1038/s41467-019-13036-1
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucl Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Karlsson FH, Fåk F, Nookaew I, Tremaroli V, Fagerberg B, Petranovic D et al (2012) Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat Commun 3:1245. https://doi.org/10.1038/ncomms2266
Li J, Li Y, Zhou Y, Wang C, Wu B, Wan J (2018) Actinomyces and alimentary tract diseases: a review of its biological functions and pathology. Biomed Res Int 2018:3820215. https://doi.org/10.1155/2018/3820215
Li M, Shao D, Zhou J, Gu J, Qin J, Chen W, et al (2020) Signatures within esophageal microbiota with progression of esophageal squamous cell carcinoma. Chin J Cancer Res 32(6):755–767. https://doi.org/10.21147/j.issn.1000-9604.2020.06.09
Liu YX, Qin Y, Chen T, Lu M, Qian X, Guo X et al (2021) A practical guide to amplicon and metagenomic analysis of microbiome data. Protein Cell 12(5):315–330. https://doi.org/10.1007/s13238-020-00724-8
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Minamoto Y, Minamoto T, Isaiah A, Sattasathuchana P, Buono A, Rangachari VR et al (2019) Fecal short-chain fatty acid concentrations and dysbiosis in dogs with chronic enteropathy. J Vet Intern Med 33(4):1608–1618. https://doi.org/10.1111/jvim.15520
Pilla R, Suchodolski JS (2019) The role of the canine gut microbiome and metabolome in health and gastrointestinal disease. Front Vet Sci 6:498. https://doi.org/10.3389/fvets.2019.00498
Pilla R, Suchodolski JS (2021) The Gut microbiome of dogs and cats, and the influence of diet. Vet Clin North Am Small Anim Pract 51(3):605–621. https://doi.org/10.1016/j.cvsm.2021.01.002
Pryde SE, Duncan SH, Hold GL, Stewart CS, Flint HJ (2002) The microbiology of butyrate formation in the human colon. FEMS Microbiol Lett 217(2):133–139. https://doi.org/10.1111/j.1574-6968.2002.tb11467.x
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285):59–65. https://doi.org/10.1038/nature08821
Ramos MT, Otto CM (2022) Canine mobility maintenance and promotion of a healthy lifestyle. Vet Clin North Am Small Anim Pract 52(4):907–924. https://doi.org/10.1016/j.cvsm.2022.03.001
Shah MM, Odoyo E, Ichinose Y (2019) Epidemiology and pathogenesis of Providencia alcalifaciens Infections. Am J Trop Med Hyg 101(2):290–293. https://doi.org/10.4269/ajtmh.18-0376
Singh A (2020) Tools for metabolomics. Nat Methods 17(1):24. https://doi.org/10.1038/s41592-019-0710-6
Suchodolski JS (2011) Companion animals symposium: microbes and gastrointestinal health of dogs and cats. J Anim Sci 89(5):1520–1530. https://doi.org/10.2527/jas.2010-3377
Suchodolski JS (2022) Analysis of the gut microbiome in dogs and cats. Vet Clin Pathol. 50(Suppl 1):6–17. https://doi.org/10.1111/vcp.13031
Suchodolski JS, Markel ME, Garcia-Mazcorro JF, Unterer S, Heilmann RM, Dowd SE et al (2012) The fecal microbiome in dogs with acute diarrhea and idiopathic inflammatory bowel disease. PLoS ONE 7(12):e51907. https://doi.org/10.1371/journal.pone.0051907
Tajima S, Waki M, Ohata A, Koda K, Maruyama Y (2015) Xanthogranulomatous gastritis associated with actinomycosis: report of a case presenting as a large submucosal mass. Int J Clin Exp Pathol 8(1):1013–1018
Treiber ML, Taft DH, Korf I, Mills DA, Lemay DG (2020) Pre- and post-sequencing recommendations for functional annotation of human fecal metagenomes. BMC Bioinform 21(1):74. https://doi.org/10.1186/s12859-020-3416-y
Vetter ND, Palmer DRJ (2021) Substrate substitution in kanosamine biosynthesis using phosphonates and phosphite rescue. Biochemistry 60(24):1926–1932. https://doi.org/10.1021/acs.biochem.1c00283
Weingarden AR, Chen C, Bobr A, Yao D, Lu Y, Nelson VM et al (2014) Microbiota transplantation restores normal fecal bile acid composition in recurrent Clostridium difficile infection. Am J Physiol Gastrointest Liver Physiol 306(4):G310-319. https://doi.org/10.1152/ajpgi.00282.2013
Wen B, Mei Z, Zeng C, Liu S (2017) metaX: a flexible and comprehensive software for processing metabolomics data. BMC Bioinform 18(1):183. https://doi.org/10.1186/s12859-017-1579-y
Yang ZW, Men Y, Zhang J, Liu ZH, Luo JY, Wang YH et al (2021) Evaluation of sample preservation approaches for better insect microbiome research according to next-generation and third-generation sequencing. Microb Ecol 82(4):971–980. https://doi.org/10.1007/s00248-021-01727-6
Zhou Q, Verne ML, Fields JZ, Lefante JJ, Basra S, Salameh H et al (2019) Randomised placebo-controlled trial of dietary glutamine supplements for postinfectious irritable bowel syndrome. Gut 68(6):996–1002. https://doi.org/10.1136/gutjnl-2017-315136
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
HSB and ZZW designed and supervised the study. TL, SJW, and LYS collected samples, recorded information, and generated data. HSB performed data analysis. ZZW conducted a data review. HSB wrote the manuscript with contributions from ZZW.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bai, H., Liu, T., Wang, S. et al. Variations in gut microbiome and metabolites of dogs with acute diarrhea in poodles and Labrador retrievers. Arch Microbiol 205, 97 (2023). https://doi.org/10.1007/s00203-023-03439-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03439-6