Abstract
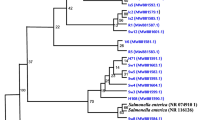
Escherichia coli has been recognized as a pathogen of concern in the antimicrobial resistance (AMR) perspective. Globally initiatives were taken to control AMR. Bacteriophages are recognized as promising alternative to antibiotics. Harnessing broad-spectrum bacteriophages for augmenting phage repositories is being prioritized across continents for future health care needs. In this context, a study was conducted to isolate coliphages against a collection of AMR E. coli isolated from diverse aquatic niche. Thirty pooled water samples (5 each from rivers, aquaculture ponds, lake, sewage treatment plant, domestic waste and canals) were analysed, and fifty-four lytic coliphages were isolated against the wide range of E. coli host strains. Broad host-spectrum phages were isolated predominantly from sewage water samples. Enriched phages were quantified, and the concentrations ranged from 106 to 107 PFU/mL. Ten phages, viz. ФEC-S-18, ФEC-S-21, ФEC-S-22, ФEC-S-23, ФEC-S-24, ФEC-S-25, ФEC-S-28, ФEC-S-30, ФEC-S-39 and ФEC-S-49, exhibited lytic activity against more than ten AMR strains of E. coli. PCR analysis of the 54 phages using the major capsid protein (MCP) specific primers coupled with gene sequence analysis identified two phages related to Berlinvirus and 35 phages to Kayfunavirus of Autographiviridae. However, the remaining 17 phages did not show amplification using the MCP primers. The study has demonstrated that aquatic environment harboured phages with broad host spectrum that can potentially be used as agents for biological control of E. coli for infection control and food safety.



Similar content being viewed by others
References
Adams MH (1959) Enumeration of bacteriophage particles. In: The bacteriophages. Interscience Publishers, London, pp 27–34
Adriaenssens EM, Sullivan MB, Knezevic P, van Zyl LJ, Sarkar BL, Dutilh BE, Krupovic M (2020) Taxonomy of prokaryotic viruses: 2018–2019 update from the ICTV Bacterial and Archaeal Viruses Subcommittee. Arch Virol 165(5):1–8
Aertsen A, Faster D, Michiels CW (2005) Induction of Shiga toxin-converting prophage in Escherichia coli by high hydrostatic pressure. Appl Environ Microbiol 71(3):1155–1162. https://doi.org/10.1128/AEM.71.3.1155-1162.2005
Aggarwal P, Uppal B, Ghosh R, Prakash SK, Rajeshwari K (2013) Highly-resistant E. coli as a common cause of paediatric diarrhoea in India. J Health Popul Nutr 31(3):409–412
Akhwale JK, Rohde M, Rohde C, Bunk B, Spröer C, Boga HI, Klenk HP, Wittmann J (2019) Isolation, characterization and analysis of bacteriophages from the haloalkaline lake Elmenteita, Kenya. PLoS ONE 14(4):e0215734. https://doi.org/10.1371/journal.pone.0215734
Almeida A, Cunha Â, Gomes N, Alves E, Costa L, Faustino MA (2009) Phage therapy and photodynamic therapy: low environmental impact approaches to inactivate microorganisms in fish farming plants. Mar Drugs 7(3):268–313
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Bai M, Cheng YH, Sun XQ, Wang ZY, Wang YX, Cui XL, Xiao W (2019) Nine novel phages from a plateau Lake in Southwest China: insights into Aeromonas phage diversity. Viruses 11(7):615
Basha KA, Ranjit NK, Vineetha D, Reshmi K, Rao BM, Lalitha KV, TomsJoseph C (2019) Prevalence, molecular characterization, genetic heterogeneity and antimicrobial resistance of Listeria monocytogenes associated with fish and fishery environment in Kerala, India. Lett Appl Microbiol 69:286–293
Benala M, Vaiyapuri M, Visnuvinayagam S, George JC, Raveendran K, George I, Mothadaka MP, Badireddy MR (2021) A revisited two-step microtiter plate assay: optimization of in vitro multiplicity of infection (MOI) for Coliphage and Vibriophage. J Virol Methods 294:114177. https://doi.org/10.1016/j.jviromet.2021.114177
Bintsis T (2018) Foodborne pathogens. AIMS Microbiol 3(3):529–563. https://doi.org/10.3934/microbiol.2017.3.529
Born Y, Knecht LE, Eigenmann M, Bolliger M, Klumpp J, Fieseler L (2019) A major-capsid-protein-based multiplex PCR assay for rapid identification of selected virulent bacteriophage types. Arch Virol 164(3):819–830
Breitbart M (2011) Marine viruses: truth or dare. Ann Rev Mar Sci 4:425–448
Carlson K (2005) Working with bacteriophages: common techniques and methodological approach. In: Kutter E, Sulakvelidze A (eds) Bacteriophages: biology and applications. CRC Press, Boca Raton, pp 437–494
Clokie MR, Millard AD, Letarov AV, Heaphy S (2011) Phages in nature. Bacteriophage 1(1):31–45
Dini C, De Urraza PJ (2010) Isolation and selection of coliphages as potential biocontrol agents of enterohemorrhagic and Shiga toxin-producing E. coli (EHEC and STEC) in cattle. J Appl Microbiol 109(3):873–887
Domingo-Calap P, Beamud B, Vienne J, González-Candelas F, Sanjuán R (2020) Isolation of four lytic phages infecting Klebsiella pneumoniae K22 clinical isolates from Spain. Int J Mol Sci 21(2):425
Efrony R, Loya Y, Bacharach E, Rosenberg E (2007) Phage therapy of coral disease. Coral Reefs 26(1):7–13
Fiedoruk K, Zakrzewska M, Daniluk T, Piktel E, Chmielewska S, Bucki R (2020) Two lineages of Pseudomonas aeruginosa filamentous phages: structural uniformity over integration preferences. Genome Biol Evol 12(10):1765–1781
Guidolin A, Morelli G, Kamke M, Manning PA (1984) Vibrio cholerae bacteriophage CP-T1: characterization of bacteriophage DNA and restriction analysis. J Virol 51(1):163–169
Hibma AM, Jassim SA, Griffiths MW (1997) Infection and removal of L-forms of Listeria monocytogenes with bred bacteriophage. Int J Food Microbiol 34(3):197–207
Jørgensen JB, Djurhuus AM, Carstens AB, Kot W, Neve H, Morris CE, Hansen LH (2020) Presentation of three novel tailed phages targeting multiple strains of Pseudomonas syringae. PHAGE 1(4):245–250
Karthika R, George I, Murugadas V, Jeswin J, Anna Sherin PS, Sanjeev D, Visnuvinayagam S, Manikanta B, Prasad MM, Madhusudana Rao B (2021) Characterization of tailed phages against E. coli using DNA restriction digestion analysis. Fish Technol 58:53–58
Kim J, Kim GH, Lee NG, Lee JS, Yoon SS (2018) Whole-genome sequencing and genomic analysis of a virulent bacteriophage infecting Bacillus cereus. Intervirology 61(6):272–280
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549
Lee DS, Lee SJ, Choe HS (2018) Community-acquired urinary tract infection by Escherichia coli in the era of antibiotic resistance. BioMed Res Int 2018:7656752
Loc-Carrillo C, Abedon ST (2011) Pros and cons of phage therapy. Bacteriophage 1(2):111–114
Malik AN, McLean PM, Roberts A, Barnett PS, Demaine AG, Banga JP, McGregor AM (1990) A simple high yield method for the preparation of lambda gt10 DNA suitable for subcloning, amplification and direct sequencing. Nucleic Acids Res 18(13):4031
Moghadam MT, Khoshbayan A, Chegini Z, Farahani I, Shariati A (2020) Bacteriophages, a new therapeutic solution for inhibiting multidrug-resistant bacteria causing wound infection: lesson from animal models and clinical trials. Drug Des Devel Ther 14:1867–1883
Muniesa M, Jofre J (1998) Abundance in sewage of bacteriophages that infect Escherichia coli O157: H7 and that carry the shiga toxin 2 gene. Appl Environ Microbiol 64(7):2443–2448
Murugadas V, Joseph TC, Reshmi K, Lalitha KV (2016a) Prevalence of methicillin resistant Staphylococcus aureus in selected seafood markets and aquaculture farms in Kerala, south-west coast of India. Indian J Fish 63(4):150–153
Murugadas V, Joseph TC, Lalitha KV (2016b) Antibiotic resistance pattern of Staphylococcus aureus isolated from seafood. Fish Tech 53:232–237
Murugadas V, Jospeh TC, Lalitha KV (2016c) Distribution of pathotypes of Escherichia coli in seafood from retail markets of Kerala, India. Indian J Fish 63(1):152–155
Murugadas V, Joseph TC, Rao BM, Lalitha KV, Prasad MM (2019) Methicillin-resistant Staphylococcus aureus in seafood: prevalence, laboratory detection, clonal nature, and control in seafood chain. J Food Sci 84:3341–3351
O’flaherty S, Coffey A, Meaney WJ, Fitzgerald GF, Ross RP (2005) Inhibition of bacteriophage K proliferation on Staphylococcus aureus in raw bovine milk. Lett Appl Microbiol 41(3):274–279
Oliveira J, Castilho F, Cunha A, Pereira MJ (2012) Bacteriophage therapy as a bacterial control strategy in aquaculture. Aquac Int 20(5):879–910
Plaza N, Castillo D, Pérez-Reytor D, Higuera G, García K, Bastías R (2018) Bacteriophages in the control of pathogenic vibrios. Electron J Biotechnol 31:24–33
Rao BM, Lalitha KV (2015) Bacteriophages for aquaculture: are they beneficial or inimical. Aquaculture 437:146–154
Rao BM, Surendran PK (2000) Occurrence of coliphages in fish and aquaculture farms. Fish Technol 37(2):146–149
Rao BM, Surendran PK (2003) Coliphage test: a quick and easy method to detect faecal pollution in water and Fish. In: Surendran PK, Mathew PT, Thampuran N, Nambiar VN, Joseph J, Boopendranath MR, Lakshmanan PT, Nair PGV (eds) Seafood safety. Society of Fisheries Technologists (India), Cochin, pp 556–561
Rao BM, Surendran PK (2013) Detection of ctx gene positive non-O1/non-O139 V. cholerae in shrimp aquaculture environments. J Food Sci Technol 50(3):496–504
Rao BM, Murthy LN, Jesmi D, Prasad MM (2014) Resistance of E. coli and Salmonella isolated from marine and freshwater fishes towards Carbapenems. Fish Tech 51:207–212
Romero-Calle D, Guimarães Benevides R, Góes-Neto A, Billington C (2019) Bacteriophages as alternatives to antibiotics in clinical care. Antibiotics 8(3):138
Sillankorva S, Neubauer P, Azeredo J (2008) Isolation and characterization of a T7-like lytic phage for Pseudomonas fluorescens. BMC Biotechnol 8:80
Sillankorva SM, Oliveira H, Azeredo J (2012) Bacteriophages and their role in food safety. Int J Microbiol. 863945. https://doi.org/10.1155/2012/863945
Soni KA, Nannapaneni R (2010) Removal of Listeria monocytogenes biofilms with bacteriophage P100. J Food Prot 73(8):1519–1524
Soni KA, Nannapaneni R, Hagens S (2010) Reduction of Listeria monocytogenes on the surface of fresh channel catfish fillets by bacteriophage Listex P100. Foodborne Pathog Dis 7(4):427–434. https://doi.org/10.1089/fpd.2009.0432
Spencer R (1955) A marine bacteriophage. Nature 175(4459):690–691
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10(3):512–526
Tang SS, Biswas SK, Tan WS, Saha AK, Leo BF (2019) Efficacy and potential of phage therapy against multidrug resistant Shigella spp. PeerJ 7:e6225
Thanki AM, Brown N, Millard AD, Clokie MR (2019) Genomic characterization of jumbo Salmonella phages that effectively target United Kingdom pig-associated Salmonella serotypes. Front Microbiol 10:1491
Thurgood TL, Sharma R, Call JJ, Chronis JD, Dawson DD, Finnegan ZK et al (2020) Genome sequences of 12 phages that infect Klebsiella pneumoniae. Microbiol Resour Announc 9(16):e00024-20. https://doi.org/10.1128/MRA.00024-20
Vaiyapuri M, Pailla S, Rao Badireddy M, Pillai D, Chandragiri Nagarajarao R, Prasad Mothadaka M (2021a) Antimicrobial resistance in Vibrios of shrimp aquaculture: incidence, identification schemes, drivers and mitigation measures. Aquac Res 52:2923–2941. https://doi.org/10.1111/are.15142
Vaiyapuri M, Raveendran K, George I, Gundubilli D, Sivam V, Krishnan SG et al (2021b) Comparison of single and multi-host enrichment approach for harnessing lytic phages against antimicrobial-resistant E. coli: repurposing the enrichment step. Biologia. 76, 1041–1052. https://doi.org/10.2478/s11756-020-00652-2
Vaiyapuri M, Sebastian AS, George I, Variem SS, Vasudevan RN, George JC, Badireddy MR, Sivam V, Peeralil S, Sanjeev D, Thandapani M, Moses SA, Nagarajarao RC, Mothadaka MP (2021c) Predominance of genetically diverse ESBL Escherichia coli identified in resistance mapping of Vembanad Lake, the largest fresh-cum-brackishwater lake of India. Environ Sci Pollut Res Int 28(46):66206–66222. https://doi.org/10.1007/s11356-021-15110-y
Van Twest R, Kropinski AM (2009) Bacteriophage enrichment from water and soil. In: Clokie MR, Kropinski AM (eds) Bacteriophages, vol 501. Humana Press, Totowa, pp 15–21
Visnuvinayagam S, Joseph TC, Murugadas V, Chakrabarti R, Lalitha KV (2015) Status on methicillin resistant and multiple drug resistant Staphylococcus aureus in fishes of Cochin and Mumbai coast, India. J Environ Biol 36(3):571
Weinbauer MG, Arrieta JM, Griebler C, Herndlb GJ (2009) Enhanced viral production and infection of bacterioplankton during an iron-induced phytoplankton bloom in the Southern Ocean. Limnol Oceanogr 54(3):774–784
Whichard JM, Sriranganathan N, Pierson FW (2003) Suppression of Salmonella growth by wild-type and large-plaque variants of bacteriophage Felix O1 in liquid culture and on chicken frankfurters. J Food Prot 66(2):220–225
Yamamoto KR, Alberts BM, Benzinger R, Lawhorne L, Treiber G (1970) Rapid bacteriophage sedimentation in the presence of polyethylene glycol and its application to large-scale virus purification. Virol J 40(3):734–744
Yang C, Wang H, Ma H, Bao R, Liu H, Yang L, Song H (2018) Characterization and genomic analysis of SFPH2, a novel T7virus infecting Shigella. Front Microbiol 9:3027
Yuan X, Zhang S, Wang J, Li C, Li N, Yu S et al (2021) Isolation and characterization of a novel Escherichia coli Kayfunavirus phage DY1. Virus Res 293:198274
Acknowledgements
The authors are highly indebted to Dr. C. N. Ravishankar, Director, ICAR-CIFT, Cochin, India, for providing necessary support to complete the research work. The authors are also thankful to the Department of Biotechnology (DBT), Govt. of India, for providing financial support for the project. The project is funded by Department of Biotechnology, Government of India (Grant no: BT/PR26949/AAQ/3/888/2017).
Funding
This work was funded by Department of Biotechnology, Ministry of Science and Technology (Grant no: BT/PR26949/AAQ/3/888/2017).
Author information
Authors and Affiliations
Contributions
IG and KR involved in isolation of phages, characterization of phages, analysis of data, gene sequencing analysis, phylogenetic tree construction and drafting of manuscript; MV, MPM and MRB conceptualized the work and analysis of data and reviewed the manuscript; MB involved in standardization of phage isolation and enumeration process; AS and DS carried out PCR identification of the phages; and IG, KR, SK and VS collected the samples from various aquatic niche and determined the host range for all the AMR E. coli.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest among authors in publishing this research work.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
George, I., Raveendran, K., Vaiyapuri, M. et al. Gene sequencing analysis of tailed phages identified diverse (Kayfunavirus and Berlinvirus) coliphages in aquatic niche against AMR Escherichia coli. Arch Microbiol 204, 429 (2022). https://doi.org/10.1007/s00203-022-03055-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-03055-w