Abstract
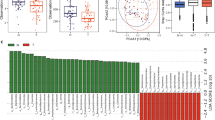
Human microbiome studies have shown diversity to exist among different ethnic populations. However, studies pertaining to the microbial composition of CRC among the Indian population have not been well explored. We aimed to decipher the microbial signature in tumor tissues from North Indian CRC patients. Next-generation sequencing of tumor and adjacent tissue-derived bacterial 16S rRNA V3-V4 hypervariable regions was performed to investigate the abundance of specific microbes. The expression profile analysis deciphered a decreased diversity among the tumor-associated microbial communities. At the phyla level, Proteobacteria was differentially expressed in CRC tissues than adjacent normal. Further, DeSeq2 normalization identified 4 out of 79 distinct species (p < 0.005) only in CRC, Bacteroides massiliensis, Alistipes onderdonkii, Bifidobacterium pseudocatenulatum, and Corynebacterium appendicis. Thus, the findings suggest that microbial signatures can be used as putative biomarkers in diagnosis, prognosis and treatment management of CRC.





Similar content being viewed by others
References
Amin MB, Greene FL, Edge SB, et a. (2017) The Eighth Edition AJCC Cancer Staging Manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin 67(2):93–99. https://doi.org/10.3322/caac.21388
Arnold M, Sierra MS, Laversanne M, Soerjomataram I, Jemal A, Bray F (2017) Global patterns and trends in colorectal cancer incidence and mortality. Gut 66(4):683–691. https://doi.org/10.1136/gutjnl-2015-310912
Arnold M, Abnet CC, Neale RE, et al (2020) Global burden of 5 major types of gastrointestinal cancer. Gastroenterology 159(1):335–349. https://doi.org/10.1053/j.gastro.2020.02.068
Chen W, Liu F, Ling Z, Tong X, Xiang C (2012) Human intestinal lumen and mucosa-associated microbiota in patients with colorectal cancer. PLoS ONE 7(6):e39743. https://doi.org/10.1371/journal.pone.0039743
Das B, Ghosh TS, Kedia S et al (2018) Analysis of the gut microbiome of rural and urban healthy Indians living in sea level and high altitude areas. Sci Rep 8(1):10104. https://doi.org/10.1038/s41598-018-28550-3
Dhakan DB, Maji A, Sharma AK, et al (2019) The unique composition of Indian gut microbiome, gene catalogue, and associated fecal metabolome deciphered using multi-omics approaches. Gigascience. https://doi.org/10.1093/gigascience/giz004
Dubey AK, Uppadhyaya N, Nilawe P et al (2018) LogMPIE, pan-India profiling of the human gut microbiome using 16S rRNA sequencing. Sci Data 5:180232. https://doi.org/10.1038/sdata.2018.232
Eckburg PB, Bik EM, Bernstein CN et al (2005) Diversity of the human intestinal microbial flora. Science 308(5728):1635–1638. https://doi.org/10.1126/science.1110591
Fidler MM, Soerjomataram I, Bray F (2016) A global view on cancer incidence and national levels of the human development index. Int J Cancer 139(11):2436–2446. https://doi.org/10.1002/ijc.30382
Goetz DH, Holmes MA, Borregaard N, Bluhm ME, Raymond KN, Strong RK (2002) The neutrophil lipocalin NGAL is a bacteriostatic agent that interferes with siderophore-mediated iron acquisition. Mol Cell 10(5):1033–1043. https://doi.org/10.1016/s1097-2765(02)00708-6
Gupta A, Dhakan DB, Maji A, et al (2019) Association of Flavonifractor plautii, a flavonoid-degrading bacterium, with the gut microbiome of colorectal cancer patients in India. mSystems. https://doi.org/10.1128/mSystems.00438-19
He T, Cheng X, Xing C (2021) The gut microbial diversity of colon cancer patients and the clinical significance. Bioengineered 12(1):7046–7060. https://doi.org/10.1080/21655979.2021.1972077
Helmink BA, Khan MAW, Hermann A, Gopalakrishnan V, Wargo JA (2019) The microbiome, cancer, and cancer therapy. Nat Med 25(3):377–388. https://doi.org/10.1038/s41591-019-0377-7
Hughes ER, Winter MG, Duerkop BA et al (2017) Microbial respiration and formate oxidation as metabolic signatures of inflammation-associated dysbiosis. Cell Host Microbe 21(2):208–219. https://doi.org/10.1016/j.chom.2017.01.005
Khochamit N, Siripornadulsil S, Sukon P, Siripornadulsil W (2015) Antibacterial activity and genotypic-phenotypic characteristics of bacteriocin-producing Bacillus subtilis KKU213: potential as a probiotic strain. Microbiol Res 170:36–50. https://doi.org/10.1016/j.micres.2014.09.004
Kim K, Castro EJT, Shim H, Advincula JVG, Kim YW (2018) Differences regarding the molecular features and gut microbiota between right and left colon cancer. Ann Coloproctol 34(6):280–285. https://doi.org/10.3393/ac.2018.12.17
Mashima I, Nakazawa F (2014) The influence of oral Veillonella species on biofilms formed by Streptococcus species. Anaerobe 28:54–61. https://doi.org/10.1016/j.anaerobe.2014.05.003
Matson V, Fessler J, Bao R et al (2018) The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359(6371):104–108. https://doi.org/10.1126/science.aao3290
Milani C, Hevia A, Foroni E et al (2013) Assessing the fecal microbiota: an optimized ion torrent 16S rRNA gene-based analysis protocol. PLoS ONE 8(7):e68739. https://doi.org/10.1371/journal.pone.0068739
Moschen AR, Gerner RR, Wang J et al (2016) Lipocalin 2 protects from inflammation and tumorigenesis associated with gut microbiota alterations. Cell Host Microbe 19(4):455–469. https://doi.org/10.1016/j.chom.2016.03.007
Nagal S, Okimura K, Kaizawa N, Ohki K, Kanatomo S (1996) Study on surfactin, a cyclic depsipeptide. II. Synthesis of surfactin B2 produced by Bacillus natto KMD 2311. Chem Pharm Bull (Tokyo) 44(1):5–10. https://doi.org/10.1248/cpb.44.5
Ozawa K, Yagu-Uchi K, Yamanaka K, Yamashita Y, Ueba K, Miwatani T (1979) Antagonistic effects of Bacillus natto and Streptococcus faecalis on growth of Candida albicans. Microbiol Immunol 23(12):1147–1156. https://doi.org/10.1111/j.1348-0421.1979.tb00547.x
Parker BJ, Wearsch PA, Veloo ACM, Rodriguez-Palacios A (2020) The genus Alistipes: gut bacteria with emerging implications to inflammation, cancer, and mental health. Front Immunol 11:906. https://doi.org/10.3389/fimmu.2020.00906
Raskov H, Burcharth J, Pommergaard HC (2017) Linking gut microbiota to colorectal cancer. J Cancer 8(17):3378–3395. https://doi.org/10.7150/jca.20497
Ringel Y, Maharshak N, Ringel-Kulka T, Wolber EA, Sartor RB, Carroll IM (2015) High throughput sequencing reveals distinct microbial populations within the mucosal and luminal niches in healthy individuals. Gut Microbes 6(3):173–181. https://doi.org/10.1080/19490976.2015.1044711
Rivadeneyra MA, Martin-Algarra A, Sanchez-Roman M, Sanchez-Navas A, Martin-Ramos JD (2010) Amorphous Ca–phosphate precursors for Ca–carbonate biominerals mediated by Chromohalobacter marismortui. ISME J 4(7):922–932. https://doi.org/10.1038/ismej.2010.17
Rizzatti G, Lopetuso LR, Gibiino G, Binda C, Gasbarrini A (2017) Proteobacteria: a common factor in human diseases. Biomed Res Int 2017:9351507. https://doi.org/10.1155/2017/9351507
Ryan MC, Stucky M, Wakefield C, et al (2019) Interactive clustered heat map builder: an easy web-based tool for creating sophisticated clustered heat maps. F1000Res. https://doi.org/10.12688/f1000research.20590.2
Shah MS, DeSantis T, Yamal JM et al (2018) Re-purposing 16S rRNA gene sequence data from within case paired tumor biopsy and tumor-adjacent biopsy or fecal samples to identify microbial markers for colorectal cancer. PLoS ONE 13(11):e0207002. https://doi.org/10.1371/journal.pone.0207002
Siegel RL, Miller KD, Goding Sauer A et al (2020) Colorectal cancer statistics. CA Cancer J Clin 70(3):145–164. https://doi.org/10.3322/caac.21601
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Tomkovich S, Yang Y, Winglee K et al (2017) Locoregional effects of microbiota in a preclinical model of colon carcinogenesis. Cancer Res 77(10):2620–2632. https://doi.org/10.1158/0008-5472.CAN-16-3472
Ventura M, O’Flaherty S, Claesson MJ et al (2009) Genome-scale analyses of health-promoting bacteria: probiogenomics. Nat Rev Microbiol 7(1):61–71. https://doi.org/10.1038/nrmicro2047
Wang Q, Ye J, Fang D et al (2020) Multi-omic profiling reveals associations between the gut mucosal microbiome, the metabolome, and host DNA methylation associated gene expression in patients with colorectal cancer. BMC Microbiol 20(Suppl 1):83. https://doi.org/10.1186/s12866-020-01762-2
Wong SH, Zhao L, Zhang X, et al (2017) Gavage of fecal samples from patients with colorectal cancer promotes intestinal carcinogenesis in germ-free and conventional mice. Gastroenterology 153(6):1621–1633. https://doi.org/10.1053/j.gastro.2017.08.022
Wu M, Wu Y, Li J, Bao Y, Guo Y, Yang W (2018) The dynamic changes of gut microbiota in Muc2 deficient mice. Int J Mol Sci. https://doi.org/10.3390/ijms19092809
Zelante T, Iannitti RG, Cunha C et al (2013) Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39(2):372–385. https://doi.org/10.1016/j.immuni.2013.08.003
Zhou P, Liu J, Merritt J, Qi F (2015) A YadA-like autotransporter, Hag1 in Veillonella atypica is a multivalent hemagglutinin involved in adherence to oral streptococci, Porphyromonas gingivalis, and human oral buccal cells. Mol Oral Microbiol 30(4):269–279. https://doi.org/10.1111/omi.12091
Zmora N, Zilberman-Schapira G, Suez J, et al. (2018) Personalized gut mucosal colonization resistance to empiric probiotics is associated with unique host and microbiome features. Cell 174(6):1388–1405. https://doi.org/10.1016/j.cell.2018.08.041
Acknowledgements
The authors would like to thank Parul Chugh (SGRH) for statistical help and the co-workers at Department of Research, SGRH.
Funding
This study was supported by the grant provided by Research and Development program (RDP Grant No.: 4.9.36-011), Sir Ganga Ram Hospital (SGRH), India.
Author information
Authors and Affiliations
Contributions
All authors contributed towards writing the manuscript and have given approval to the final version of the manuscript. RH performed collection of human samples/isolation/NGS assays and framing the manuscript. SB guided, framed the manuscript, inferenced the data and edited the manuscript. RR performed the metagenomics data analysis and prepared the figures. SR enrolled the patients as per inclusion criteria, provided the clinical samples and demographics of the patients. DB contributed in manuscript proof reading. PN & NKS assisted in experimental standardization and NGS data acquisition and cleaning. SC conceptualized, guided, analyzed the data, framed and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This study was reviewed and approved by Sir Ganga Ram Hospital Ethics Committee (Ethics approval code: EC/06/16/1000). The patients/participants provided their informed consent to participate in this study.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hasan, R., Bose, S., Roy, R. et al. Tumor tissue-specific bacterial biomarker panel for colorectal cancer: Bacteroides massiliensis, Alistipes species, Alistipes onderdonkii, Bifidobacterium pseudocatenulatum, Corynebacterium appendicis. Arch Microbiol 204, 348 (2022). https://doi.org/10.1007/s00203-022-02954-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-02954-2