Abstract
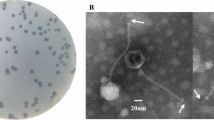
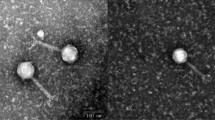
A novel lytic Enterococcus faecalis phage, EFC1, was isolated from the sewage of a farm in Handan, China, and its genome was analyzed and described. The phage could infect 87.5% of the chicken-derived Enterococcus faecalis preserved in our laboratory. The genome of phage EFC1 consists of a circular double-stranded DNA with a length of 56,099 bp and a G + C content of 39.96%, containing 89 predicted protein-coding genes as well as 2 tRNAs, which are involved in phage intron, structure, transcription, packaging, DNA replication, modification, cell lysis, and other functions, indicating the genetic and functional characteristics of this phage. Genome comparison analysis revealed that phage EFC1 can be regarded as new genus Saphexavirus phage in the Siphoviridae family.



Similar content being viewed by others
References
Bebeacua C, Lorenzo Fajardo JC, Blangy S et al (2013) X-ray structure of a superinfection exclusion lipoprotein from phage TP-J34 and identification of the tape measure protein as its target. Mol Microbiol 89:152–165. https://doi.org/10.1111/mmi.12267
Blanco AE, Barz M, Icken W et al (2016) Twenty years of amyloid arthropathy research in chickens. Worlds Poult Sci J 72:495–508. https://doi.org/10.1017/S0043933916000453
Boetzer M, Pirovano W (2012) Toward almost closed genomes with GapFiller. Genome Biol 13:R56. https://doi.org/10.1186/gb-2012-13-6-r56
Boetzer M, Henkel CV, Jansen HJ et al (2011) Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27:578–579. https://doi.org/10.1093/bioinformatics/btq683
Chen S, Zhou Y, Chen Y et al (2018a) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Chen Y, Sun E, Yang L et al (2018b) Therapeutic application of bacteriophage PHB02 and its putative depolymerase against Pasteurella multocida capsular Type A in mice. Front Microbiol 9:1678. https://doi.org/10.3389/fmicb.2018.01678
Cheng M, Liang J, Zhang Y et al (2017) The Bacteriophage EF-P29 efficiently protects against lethal vancomycin-resistant Enterococcus faecalis and alleviates gut microbiota imbalance in a murine bacteremia model. Front Microbiol 8:837–837. https://doi.org/10.3389/fmicb.2017.00837
Chibani CM, Farr A, Klama S et al (2019) Classifying the unclassified: a phage classification method. Viruses. https://doi.org/10.3390/v11020195
Cisek AA, Dąbrowska I, Gregorczyk KP et al (2017) Phage therapy in bacterial infections treatment: one hundred years after the discovery of bacteriophages. Curr Microbiol 74:277–283. https://doi.org/10.1007/s00284-016-1166-x
Furuta M, Nasu T, Umeki K et al (2017) Characterization and application of lytic bacteriophages against campylobacter jejuni isolated from poultry in Japan. Biocontrol Sci 22:213–221. https://doi.org/10.4265/bio.22.213
Gibb EA, Edgell DR (2007) Multiple controls regulate the expression of mobE, an HNH homing endonuclease gene embedded within a ribonucleotide reductase gene of phage Aeh1. J Bacteriol 189:4648–4661. https://doi.org/10.1128/JB.00321-07
Hashmi JA, Al-Harbi KM, Ramzan K et al (2016) A novel splice-site mutation in the ASPM gene underlies autosomal recessive primary microcephaly. Ann Saudi Med 36:391–396. https://doi.org/10.5144/0256-4947.2016.391
Hyatt D, Chen GL, Locascio PF et al (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119. https://doi.org/10.1186/1471-2105-11-119
Kim J, Shen R, Olcott MC et al (2005) Adenylate kinase of Escherichia coli, a component of the phage T4 dNTP synthetase complex. J Biol Chem 280:28221–28229. https://doi.org/10.1074/jbc.M502201200
Kim MH, Moon DC, Kim S-J et al (2021) Nationwide surveillance on antimicrobial resistance profiles of Enterococcus faecium and Enterococcus faecalis isolated from healthy food animals in South Korea, 2010 to 2019. Microorganisms 9:925. https://doi.org/10.3390/microorganisms9050925
Korf IHE, Kittler S, Bierbrodt A et al (2020) In vitro evaluation of a phage cocktail controlling infections with Escherichia coli. Viruses 12:1470. https://doi.org/10.3390/v12121470
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Le S, He X, Tan Y et al (2013) Mapping the tail fiber as the receptor binding protein responsible for differential host specificity of Pseudomonas aeruginosa bacteriophages PaP1 and JG004. PLoS ONE 8:e68562–e68562. https://doi.org/10.1371/journal.pone.0068562
Li M, Lin H, Jing Y et al (2020) Broad-host-range Salmonella bacteriophage STP4-a and its potential application evaluation in poultry industry. Poult Sci 99:3643–3654. https://doi.org/10.1016/j.psj.2020.03.051
Olsen RH, Frantzen C, Christensen H et al (2012) An investigation on first-week mortality in layers. Avian Dis 56:51–57. https://doi.org/10.1637/9777-051011-Reg.1
Pell LG, Kanelis V, Donaldson LW et al (2009) The phage lambda major tail protein structure reveals a common evolution for long-tailed phages and the type VI bacterial secretion system. Proc Natl Acad Sci USA 106:4160–4165. https://doi.org/10.1073/pnas.0900044106
Penadés JR, Chen J, Quiles-Puchalt N et al (2015) Bacteriophage-mediated spread of bacterial virulence genes. Curr Opin Microbiol 23:171–178. https://doi.org/10.1016/j.mib.2014.11.019
Rabinovitch I, Yanku M, Yeheskel A et al (2010) Staphylococcus aureus NrdH redoxin is a reductant of the class Ib ribonucleotide reductase. J Bacteriol 192:4963–4972. https://doi.org/10.1128/JB.00539-10
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686-689. https://doi.org/10.1093/nar/gki366
Stothard P, Wishart DS (2005) Circular genome visualization and exploration using CGView. Bioinformatics 21:537–539. https://doi.org/10.1093/bioinformatics/bti054
Tavares P, Zinn-Justin S, Orlova EV (2012) Genome gating in tailed bacteriophage capsids. Adv Exp Med Biol 726:585–600. https://doi.org/10.1007/978-1-4614-0980-9_25
Van Boeckel TP, Brower C, Gilbert M et al (2015) Global trends in antimicrobial use in food animals. Proc Natl Acad Sci USA 112:5649–5654. https://doi.org/10.1073/pnas.1503141112
Yang D, Chen Y, Sun E et al (2020) Characterization of a lytic bacteriophage vB_EfaS_PHB08 harboring endolysin Lys08 against Enterococcus faecalis biofilms. Microorganisms 8:1332. https://doi.org/10.3390/microorganisms8091332
Yasui R, Yoshida C, Yamaguchi T et al (2017) Characterization of an anti-varicella-zoster virus compound that targets the portal protein encoded by ORF54. Microbiol Immunol 61:398–402. https://doi.org/10.1111/1348-0421.12507
Zerbino DR, McEwen GK, Margulies EH et al (2009) Pebble and rock band: heuristic resolution of repeats and scaffolding in the velvet short-read de novo assembler. PLoS ONE 4:e8407. https://doi.org/10.1371/journal.pone.0008407
Acknowledgements
This study received the support of the Department of science and technology of Hebei Province, China and the school of life sciences and food engineering, Hebei University of Engineering, China. Thanks to the Electron Microscopy Experimental Center of Hebei Medical University for providing equipment and guidance training. The library preparation, sequencing and assembly, and annotation guidance are all supported and guided by GENEWIZ Technology Co., Ltd.
Funding
This study was funded by Special project for tackling key technical problems of science and Technology Department of Hebei Province (19226609D).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author has no conflict of interests to disclose.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Nucleotide sequence accession number
The GenBank accession number for phage EFC1 is MW677132.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Q., Liu, N. Complete genome analysis of bacteriophage EFC1 infecting Enterococcus faecalis from chicken. Arch Microbiol 204, 413 (2022). https://doi.org/10.1007/s00203-022-02838-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-02838-5