Abstract
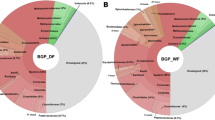
The search for sustainable development has increased interest in the improvement of technologies that use renewable energy sources. One of the alternatives in the production of renewable energy comes from the use of waste including urban solids, animal excrement from livestock, and biomass residues from agro-industrial plants. These materials may be used in the production of biogas, making its production highly sustainable and environmentally friendly. The present study aimed to evaluate the cultivated and uncultivated microbial community from a substrate (starter) used as an adapter for biogas production in anaerobic digestion processes. 16S rDNA metabarcoding revealed the domain of bacteria belonging to the phyla Firmicutes, Bacteroidota, Chloroflexi and Synergistota. The methanogenic group was represented by the phyla Halobacterota and Euryarchaeota. Through 16S rRNA sequencing of isolates recovered from the starter culture, the genera Rhodococcus (Actinobacteria phylum), Vagococcus, Lysinibacillus, Niallia, Priestia, Robertmurraya, Proteiniclasticum (Firmicutes phylum), and Luteimonas (Proteobacteria phylum) were identified, genera that were not observed in the metabarcoding data. The volatile solids, volatile organic acids, and total inorganic carbon reached 659.10 g kg−1, 717.70 g kg−1, 70,005.0 g kg−1, respectively. The cultured groups are involved in the metabolism of sugars and other compounds derived from lignocellulosic material, as well as in anaerobic methane production processes. The results demonstrate that culture-dependent approaches, such as isolation and sequencing, and culture-independent studies, such as the Metabarcoding approach, are complementary methodologies that, when integrated provide robust and comprehensive information about the microbial communities involved in processes of the production of biogas in anaerobic digestion processes.




Similar content being viewed by others
References
Aamir S, Sutar S, Sk S, Baghela A (2015) A rapid and efficient method of fungal genomic DNA extraction, suitable for PCR based molecular methods. Plant Pathol Quarant 5:74–81. https://doi.org/10.5943/ppq/5/2/6
Agustini CB (2014). Isolamento microbiano na biodegradação de resíduos de curtumes.
Ali S, Hua B, Jeanne J et al (2019) Bioresource Technology Effect of different initial low pH conditions on biogas production, composition, and shift in the aceticlastic methanogenic population. Bioresour Technol 289:121579. https://doi.org/10.1016/j.biortech.2019.121579
Ali Shah F, Mahmood Q, Maroof Shah M, Pervez A, Ahmad Asad S (2014) Microbial ecology of anaerobic digesters: the key players of anaerobiosis. Sci World J
Andrews S (2017) FastQC: a quality control tool for high throughput sequence data
Anguita-Maeso M, Olivares-García C, Haro C et al (2020) Culture-dependent and culture-independent characterization of the olive xylem microbiota: effect of sap extraction methods. Front Plant Sci. https://doi.org/10.3389/fpls.2019.01708
Anthony WE, Carr RR, Delorenzo DM et al (2019) Development of Rhodococcus opacus as a chassis for lignin valorization and bioproduction of high-value compounds. Biotechnol Biofuels 12:1–14. https://doi.org/10.1186/s13068-019-1535-3
APHA. American Publish Health Association (2017) Standard methods for the examination of water and wastewater, 23rd edn. American Water Works Association, Washington
Belgini DRB, Dias RS, Siqueira VM et al (2014) Culturable bacterial diversity from a feed water of a reverse osmosis system, evaluation of biofilm formation and biocontrol using phages. World J Microbiol Biotechnol 30:2689–2700. https://doi.org/10.1007/s11274-014-1693-1
Biedendieck R, Knuuti T, Moore SJ, Jahn D (2021) The “beauty in the beast”—the multiple uses of Priestia megaterium in biotechnology. Appl Microbiol Biotechnol 105:5719–5737. https://doi.org/10.1007/s00253-021-11424-6
Agustini CB (2014) Isolamento microbiano na biodegradação deresíduos de curtumes
Borges AC (2014) Isolamento microbiano na biodegradação de resíduos de cortume Dissertação. Universidade Federal do Rio Grande do Sul, Escola de Engenharia
Brandt C, Bongcam-Rudloff E, Müller B (2020) Abundance tracking by long-read nanopore sequencing of complex microbial communities in samples from 20 different biogas/wastewater plants. Appl Sci 10:1–14. https://doi.org/10.3390/app10217518
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) correspondence QIIME allows analysis of high- throughput community sequencing data Intensity normalization improves color calling in SOLiD sequencing. Nat Publ Gr 7:335–336. https://doi.org/10.1038/nmeth0510-335
Cerqueira MBR, Dias AN, Caldas SS et al (2011) Validação de método para determinação de ácidos orgânicos voláteis em efluentes de reatores anaeróbios empregando cromatografia líquida. Quim Nova 34:156–159. https://doi.org/10.1590/s0100-40422011000100029
Chaikitkaew S, Seengenyoung J, Mamimin C et al (2021) Simultaneous biogas upgrading and acetic acid production by homoacetogens consortium enriched from peatland soil. Bioresour Technol Reports 15:100701. https://doi.org/10.1016/j.biteb.2021.100701
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Stat 11:265–270. https://doi.org/10.2307/4615964
Chen S, Cheng H, Liu J et al (2017) Unexpected competitiveness of Methanosaeta populations at elevated acetate concentrations in methanogenic treatment of animal wastewater. Appl Microbiol Biotechnol 101:1729–1738. https://doi.org/10.1007/s00253-016-7967-9
Chernicharo CAL. Reatores anaeróbios. 2.ed. Belo Horizonte: DESA, UFMG, 2007. 380p. (Princípios do Tratamento Biológico de Águas Residuárias, v.5)
Covey KR, Megonigal JP (2019) Methane production and emissions in trees and forests. New Phytol 222(1):35–51. https://doi.org/10.1111/nph.15624
Dong L, Cao G, Guo X et al (2019) Efficient biogas production from cattle manure in a plug flow reactor: a large scale long term study. Bioresour Technol 278:450–455. https://doi.org/10.1016/j.biortech.2019.01.100
Ferry JG, Smith PH, Wolfe RS (1974) Methanospirillum, a new genus of methanogenic bacteria, and characterization of Methanospirillum hungatii sp.nov. Int J Syst Bacteriol 24:465–469. https://doi.org/10.1099/00207713-24-4-465
Gielen D, Boshell F, Saygin D et al (2019) The role of renewable energy in the global energy transformation. Energy Strateg Rev 24:38–50. https://doi.org/10.1016/j.esr.2019.01.006
Gulhane M, Pandit P, Khardenavis A et al (2017) Study of microbial community plasticity for anaerobic digestion of vegetable waste in Anaerobic Baffled Reactor. Renew Energy 101:59–66. https://doi.org/10.1016/j.renene.2016.08.021
Gupta RS, Patel S, Saini N, Chen S (2020) Robust demarcation of 17 distinct Bacillus species clades, proposed as novel Bacillaceae genera, by phylogenomics and comparative genomic analyses: Description of Robertmurraya kyonggiensis sp. nov. and proposal for an emended genus Bacillus limiting it only to the members of the Subtilis and Cereus clades of species. Int J Syst Evol Microbiol 70:5753–5798. https://doi.org/10.1099/ijsem.0.004475
Hahnke S, Striesow J, Elvert M et al (2014) Clostridium bornimense sp. nov., isolated from a mesophilic, two-phase, laboratory-scale biogas reactor. Int J Syst Evol Microbiol 64:2792–2797. https://doi.org/10.1099/ijs.0.059691-0
Han R, Liu L, Meng Y et al (2021) Archaeal and bacterial community structures of rural household biogas digesters with different raw materials in Qinghai Plateau. Biotechnol Lett 43:1337–1348. https://doi.org/10.1007/s10529-021-03105-1
Hania WB, Bouanane-Darenfed A, Cayol JL et al (2016) Reclassification of Anaerobaculum mobile, Anaerobaculum thermoterrenum, Anaerobaculum hydrogeniformans as Acetomicrobium mobile comb. nov., Acetomicrobium thermoterrenum comb. nov. and Acetomicrobium hydrogeniformans comb. nov., respectively, and emendation of the genus Acetomicrobium. Int J Syst Evol Microbiol 66:1506–1509. https://doi.org/10.1099/ijsem.0.000910
He C, Zhang B, Jiang Y et al (2021) Microbial electrolysis cell produced biogas as sustainable electron donor for microbial chromate reduction. Chem Eng J 403:126429. https://doi.org/10.1016/j.cej.2020.126429
Heitkamp K, Latorre-Pérez A, Nefigmann S et al (2021) Monitoring of seven industrial anaerobic digesters supplied with biochar. Biotechnol Biofuels 14:1–14. https://doi.org/10.1186/s13068-021-02034-5
Kakuk B, Wirth R, Maróti G et al (2021) Early response of methanogenic archaea to H2 as evaluated by metagenomics and metatranscriptomics. Microb Cell Fact. https://doi.org/10.1186/s12934-021-01618-y
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Knoblauch C, Beer C, Liebner S et al (2018) Methane production as key to the greenhouse gas budget of thawing permafrost. Nat Clim Chang. https://doi.org/10.1038/s41558-018-0095-z
Kunz A, Steinmetz RLR, do Amaral AC (2019) Fundamentos da digestão anaeróbia, purificação do biogás, uso e tratamento do digestato- Concórdia. Embrapa Suínos e Aves, Sbera, p 209
Litti YV, Kovalev DA, Kovalev AA, Merkel AY, Vishnyakova AV, Russkova YI, Nozhevnikova AN (2021) Auto-selection of microorganisms of sewage sludge used as an inoculum for fermentative hydrogen production from different substrates. Int J Hydrogen Energy 46(58):29834–29845. https://doi.org/10.1016/j.ijhydene.2021.06.174
Lohani SP, Havukainen J (2018) Anaerobic digestion: factors affecting anaerobic digestion process. Energy Environ Sustain. https://doi.org/10.1007/978-981-10-7413-4_18
Manimegalai R, Gopinath LR, Christy PM, Divya D (2014) Isolation and identification of acetogenic and methanogenic bacteria from anoxic black sediments and their role in biogas production. Int J Plant Anim Environ Sci 4(3):156–164
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J 17(1):10–12
Mcmurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. https://doi.org/10.1371/journal.pone.0061217
Mukhuba M, Roopnarain A, Moeletsi ME, Adeleke R (2020) Metagenomic insights into the microbial community and biogas production pattern during anaerobic digestion of cow dung and mixed food waste. J Chem Technol Biotechnol 95:151–162. https://doi.org/10.1002/jctb.6217
Murunga SI, Duncan OM, Ayub NG et al (2016) Isolation and characterization of methanogenic bacteria from brewery wastewater in Kenya. African J Biotechnol 15:2687–2697. https://doi.org/10.5897/ajb2016.15551
Muturi SM, Muthui LW, Njogu PM et al (2021) Metagenomics survey unravels diversity of biogas microbiomes with potential to enhance productivity in Kenya. PLoS ONE 16:e0244755. https://doi.org/10.1371/journal.pone.0244755
Nordgård ASR, Bergland WH, Vadstein O et al (2017) Anaerobic digestion of pig manure supernatant at high ammonia concentrations characterized by high abundances of Methanosaeta and non-euryarchaeotal archaea. Sci Rep 7:1–14. https://doi.org/10.1038/s41598-017-14527-1
Orhorhoro EK, Ebunilo PO, Sadjere GE (2018) Effect of organic loading rate (OLR) on biogas yield using a single and three-stages continuous anaerobic digestion reactors. Int J Eng Res Africa 39:147–155. https://doi.org/10.4028/www.scientific.net/JERA.39.147
Persinoti GF, Paixão DAA, Bugg TDH, Squina FM (2018) Genome sequence of Lysinibacillus sphaericus, a lignin-degrading bacterium isolated from municipal solid waste soil. Genome Announc 6:1–2. https://doi.org/10.1128/genomeA.00353-18
Petriglieri F, Nierychlo M, Nielsen PH, McIlroy SJ (2018) In situ visualisation of the abundant Chloroflexi populations in full-scale anaerobic digesters and the fate of immigrating species. PLoS ONE 13:1–14. https://doi.org/10.1371/journal.pone.0206255
Poirier S, Déjean S, Midoux C et al (2020) Integrating independent microbial studies to build predictive models of anaerobic digestion inhibition by ammonia and phenol. Bioresour Technol 316:123952. https://doi.org/10.1016/j.biortech.2020.123952
Pu C, Liu L, Yao M et al (2018) Responses and successions of sulfonamides, tetracyclines and fluoroquinolones resistance genes and bacterial community during the short-term storage of biogas residue and organic manure under the incubator and natural conditions. Environ Pollut 242:749–759. https://doi.org/10.1016/j.envpol.2018.07.063
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Rabah A, Baki A, Hassan L et al (2010) Production of biogas using abattoir waste at different retention time. Sci Wold J 5:23–26
Rashid GMM, Durán-Peña MJ, Rahmanpour R et al (2017) Delignification and enhanced gas release from soil containing lignocellulose by treatment with bacterial lignin degraders. J Appl Microbiol 123:159–171. https://doi.org/10.1111/jam.13470
Ren NQ, Chua H, Chan SY et al (2007) Assessing optimal fermentation type for bio-hydrogen production in continuous-flow acidogenic reactors. Bioresour Technol 98:1774–1780. https://doi.org/10.1016/j.biortech.2006.07.026
Roh SW, Kim KH, Do NY et al (2008) Luteimonas aestuarii sp. nov., isolated from tidal flat sediment. J Microbiol 46:525–529. https://doi.org/10.1007/s12275-008-0189-9
Saitou N, Nei M (1987) ESCALA CIWA-AR Escala CIWA-Ar (Clinical Institute Withdrawal Assesment for Alcohol) Evaluación del Síndrome de Abstinencia Alcohólica. Mol Biol Evol 4:406–425
Soares CMT, Feiden A, Tavares SG (2017) Fatores que influenciam o processo de digestão anaeróbia na produção de biogás Factors that influence the anaerobic digestion process in biogas production. Rev Nativ 5:522–528
Sun H, Bjerketorp J, Levenfors JJ, Schnürer A (2020) Isolation of antibiotic-resistant bacteria in biogas digestate and their susceptibility to antibiotics. Environ Pollut 266:115265
The Scientific World Journal (2017) Retracted: microbial ecology of anaerobic digesters: the key players of anaerobiosis. Sci World J 2017:3852369. https://doi.org/10.1155/2017/3852369
Vale M, Mateus MM, Galhano dos Santos R et al (2019) Replacement of petroleum-derived diols by sustainable biopolyols in one component polyurethane foams. J Clean Prod 212:1036–1043. https://doi.org/10.1016/j.jclepro.2018.12.088
Valijanian E, Meisam T, Mortaza A, Alawi S, Yusuf C (2018) Biogas production systems. In: Tabatabaei M, Ghanavati H (eds) Biogas: fundamentals, process and operation. Springer, Cham, pp 95–116
Vrieze J, Verstraete W (2016) Perspectives for microbial community composition in anaerobic digestion: from abundance and activity to connectivity. Environ Microbiol 18:2797–2809. https://doi.org/10.1111/1462-2920.13437
Wang L, Cui YS, Kwon CS, Lee ST, Lee JS, Im WT (2011) Vagococcus acidifermentans sp. nov., isolated from an acidogenic fermentation reactor. Int J Syst Evol Microbiol 61(5):1123–1126. https://doi.org/10.1099/ijs.0.022087-0
Watcharasukarn M, Kaparaju P, Steyer JP, Krogfelt KA, Angelidaki I (2009) Screening Escherichia coli, Enterococcus faecalis, and Clostridium perfringens as indicator organisms in evaluating pathogen-reducing capacity in biogas plants. Microb Ecol 58(2):221–230. https://doi.org/10.1007/s00248-009-9497-9
Wei Y, Wang F, Gao J, Huang Y, Ren W, Sheng H (2021) Culture-dependent and culture-independent characterization of bacterial community diversity in different types of sandy lands: the case of Minqin County, China. BMC Microbiol 21(1):1–15. https://doi.org/10.1186/s12866-021-02150-0
Westerholm M, Schnürer A (2019) Microbial responses to different operating practices for biogas production systems. Anaerob Dig. https://doi.org/10.5772/intechopen.82815
Wongfaed N, Kongjan P, Prasertsan P, Sompong O (2020) Effect of oil and derivative in palm oil mill effluent on the process imbalance of biogas production. J Clean Prod 247:119110. https://doi.org/10.1016/j.jclepro.2019.119110
Xie Z, Meng X, Ding H, Cao Q, Chen Y, Liu X, Li D (2021) The synergistic effect of rumen cellulolytic bacteria and activated carbon on thermophilic digestion of cornstalk. Biores Technol 338:125566. https://doi.org/10.1016/j.biortech.2021.125566
Xu Q, Luo TY, Wu RL, Wei W, Sun J, Dai X, Ni BJ (2021) Rhamnolipid pretreatment enhances methane production from two-phase anaerobic digestion of waste activated sludge. Water Res 194:116909. https://doi.org/10.1016/j.watres.2021.116909
Yan BH, Selvam A, Wong JW (2020) Bio-hydrogen and methane production from two-phase anaerobic digestion of food waste under the scheme of acidogenic off-gas reuse. Biores Technol 297:122400. https://doi.org/10.1016/j.biortech.2019.122400
Yıldırım E, Ince O, Aydin S, Ince B (2017) Improvement of biogas potential of anaerobic digesters using rumen fungi. Renewable Energy 109:346–353. https://doi.org/10.1016/j.renene.2017.03.021
Yoon YM, Kim M, Kim ET, Park SJ (2021) Draft genome sequence of Bacillus sp. strain B1–b2 isolated from meconium. Microbiol Soc Korea 57(1):55–57
Zamorano-López N, Borrás L, Seco A, Aguado D (2020) Unveiling microbial structures during raw microalgae digestion and co-digestion with primary sludge to produce biogas using semi-continuous AnMBR systems. Sci Total Environ 699:134365. https://doi.org/10.1016/j.scitotenv.2019.134365
Zhang J, Zhang R, He Q, Ji B, Wang H, Yang K (2020) Adaptation to salinity: response of biogas production and microbial communities in anaerobic digestion of kitchen waste to salinity stress. J Biosci Bioeng 130(2):173–178. https://doi.org/10.1016/j.jbiosc.2019.11.011
Zhao HY, Jie LI, Liu JJ, Lü YC, Wang XF, Cui ZJ (2013) Microbial community dynamics during biogas slurry and cow manure compost. J Integr Agric 12(6):1087–1097. https://doi.org/10.1016/S2095-3119(13)60488-8
Zhou J, Yang J, Yu Q, Yong X, Xie X, Zhang L, Jia H (2017) Different organic loading rates on the biogas production during the anaerobic digestion of rice straw: a pilot study. Biores Technol 244:865–871. https://doi.org/10.1016/j.biortech.2017.07.146
Acknowledgements
We thank the International Center for Renewable Energies—Biogás supported by Itaipu Binacional (supported by Itaipu Technological Park Foundation), EDITAL PRPPG Nº 105/2020—Latin America and the Caribbean Priority Institutional Program and EDITAL PRPPG n° 80/2019—Researcher Integration Assistance Program (PAIP).
Funding
Financial support granted by International Center for Renewable Energies – Biogás, supported by Itaipu Binacional; Latin American and the Caribbean Priority Institutional Program (PRPPG 105/2020), Researcher Integration Assistance Program (PAIP)(Edital PRPPG nº80/2019).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ottoni, J.R., Bernal, S.P.F., Marteres, T.J. et al. Cultured and uncultured microbial community associated with biogas production in anaerobic digestion processes. Arch Microbiol 204, 340 (2022). https://doi.org/10.1007/s00203-022-02819-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-02819-8