Abstract
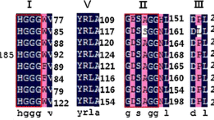
A novel lipolytic gene, estq7, was identified from a fosmid metagenomic library. The recombinant enzyme EstQ7 consists of 370 amino acids with an anticipated molecular mass of 42 kDa. Multiple sequence alignments showed that EstQ7 contained a pentapeptide motif GHSMG, and a putative catalytic triad Ser174–Asp306–His344. Interestingly, EstQ7 was found to have very little similarity to the characterized lipolytic enzymes. Phylogenetic analysis revealed that EstQ7 may be a member of a novel family of lipolytic enzymes. Biochemical characterization of the recombinant enzyme revealed that it constitutes a slightly alkalophilic, moderate thermophilic and highly active carboxylesterase against short-chain fatty acid esters with optimum temperature 50 ℃ and pH 8.2. The Km and kcat values toward p-nitrophenyl acetate were determined to be 0.17 mM and 1910s−1, respectively. Moreover, EstQ7 was demonstrated to have acyltransferase activity by GC–MS analysis. Structural modeling of the three-dimensional structure of this new enzyme showed that it exhibits a typical α/β hydrolase fold, and the catalytic triad residues are spatially close. Molecular docking revealed the interactions between the enzyme and the ligand. The high levels of lipolytic activity of EstQ7, combined with its moderate thermophilic property and acyltransferase activity, render this novel enzyme a promising candidate biocatalyst for food, pharmaceutical and biotechnological applications.








Similar content being viewed by others
References
Alex D, Shainu A, Pandey A, Sukumaran RK (2014) Esterase active in polar organic solvents from the yeast Pseudozyma sp. NII 08165. Enzyme Res 2014:494682. https://doi.org/10.1155/2014/494682
Arpigny JL, Jaeger KE (1999) Bacterial lipolytic enzymes: classification and properties. Biochem J 343:177–183. https://doi.org/10.1042/0264-6021:3430177
Bachar A et al (2010) Soil microbial abundance and diversity along a low precipitation gradient. Microb Ecol 60:453–461. https://doi.org/10.1007/s00248-010-9727-1
Biver S, Vandenbol M (2013) Characterization of three new carboxylic ester hydrolases isolated by functional screening of a forest soil metagenomic library. J Ind Microbiol Biot 40:191–200. https://doi.org/10.1007/s10295-012-1217-7
Bommarius AS (2015) Biocatalysis: a status report. Annu Rev Chem Biomol Eng 6:319–345. https://doi.org/10.1146/annurev-chembioeng-061114-123415
Bornscheuer UT (2002) Microbial carboxyl esterases: classification, properties and application in biocatalysis. Fems Microbiol Rev 26:73–81. https://doi.org/10.1111/j.1574-6976.2002.tb00599.x
Brady SF (2007) Construction of soil environmental DNA cosmid libraries and screening for clones that produce biologically active small molecules. Nat Protoc 2:1297–1305. https://doi.org/10.1038/nprot.2007.195
Buller AR, Townsend CA (2013) Intrinsic evolutionary constraints on protease structure, enzyme acylation, and the identity of the catalytic triad. Proc Natl Acad Sci Usa 110:E653–E661. https://doi.org/10.1073/pnas.1221050110
Byun JS et al (2007) Crystal structure of hyperthermophilic esterase EstE1 and the relationship between its dimerization and thermostability properties. BMC Struct Biol 7:47. https://doi.org/10.1186/1472-6807-7-47
Choi JE, Kwon MA, Na HY, Hahm DH, Song JK (2013) Isolation and characterization of a metagenome-derived thermoalkaliphilic esterase with high stability over a broad pH range. Extremophiles 17:1013–1021. https://doi.org/10.1007/s00792-013-0583-z
Chu X, He H, Guo C, Sun B (2008) Identification of two novel esterases from a marine metagenomic library derived from South China Sea. Appl Microbiol Biotechnol 80:615–625. https://doi.org/10.1007/s00253-008-1566-3
Daniel W, Janaina J, Rachel R, Jaigeeth D, Kui W, Zdzislaw W (2017) Biochemical and structural analyses of two cryptic esterases in bacteroides intestinalis and their synergistic activities with cognate xylanases. J Mol Biol 429:2509–2527. https://doi.org/10.1016/j.jmb.2017.06.017
Ferrer M, Martinez-Martinez M, Bargiela R, Streit WR, Golyshina OV, Golyshin PN (2016) Estimating the success of enzyme bioprospecting through metagenomics: current status and future trends. Microb Biotechnol 9:22–34. https://doi.org/10.1111/1751-7915.12309
Garg R, Srivastava R, Brahma V, Verma L, Karthikeyan S, Sahni G (2016) Biochemical and structural characterization of a novel halotolerant cellulase from soil metagenome. Sci Rep 6:39634. https://doi.org/10.1038/srep39634
Gerits LR, Pareyt B, Decamps K, Delcour JA (2014) Lipases and their functionality in the production of wheat-based food systems. Compr Rev Food Sci F 13:978–989. https://doi.org/10.1111/1541-4337.12085
Gong BL et al (2017) Improvement of enzyme activity and soluble expression of an alkaline protease isolated from oil-polluted mud flat metagenome by random mutagenesis. Enzyme Microb Technol 106:97–105. https://doi.org/10.1016/j.enzmictec.2017.06.015
Gurung N, Ray S, Bose S, Rai V (2013) A broader view: Microbial enzymes and their relevance in industries, medicine, and beyond. Biomed Res Int Doi: Artn. https://doi.org/10.1155/2013/329121
Hitch TCA, Clavel T (2019) A proposed update for the classification and description of bacterial lipolytic enzymes. PeerJ. https://doi.org/10.7717/peerj.7249
Hyun P, Jeong J, Kyungmoon P, Yong K, Young Y (2013) Prediction of the solvent affecting site and the computational design of stable Candida antarctica lipase B in a hydrophilic organic solvent. J Biotechnol 163:346–352. https://doi.org/10.1016/j.jbiotec.2012.11.00610.1016/j
Jaeger KE, Eggert T (2002) Lipases for biotechnology. Curr Opin. Biotech 13:390–397. https://doi.org/10.1016/S0958-1669(02)00341-5
Jayanath G, Sowmya P, Bhavya K, Solly S, Bright S, Rosamma P (2018) A novel solvent tolerant esterase of GDSGG motif subfamily from solar saltern through metagenomic approach: Recombinant expression and characterization. Int J Biol Macromol 119:393–401. https://doi.org/10.1016/j.ijbiomac.2018.06.057
Katz M, Hover BM, Brady SF (2016) Culture-independent discovery of natural products from soil metagenomes. J Ind Microbiol Biotechnol 43:129–141. https://doi.org/10.1007/s10295-015-1706-6
Kawata T, Ogino H (2010) Amino acid residues involved in organic solvent-stability of the LST-03 lipase. Biochem Biophys Res Commun 400:384–388. https://doi.org/10.1016/j.bbrc.2010.08.080
Li PY et al (2015) Interdomain hydrophobic interactions modulate the thermostability of microbial esterases from the hormone-sensitive lipase family. J Biol Chem 290:11188–11198. https://doi.org/10.1074/jbc.M115.646182
Li X et al (2018) Identification of a novel feruloyl esterase by functional screening of a soil metagenomic library. Appl Biochem Biotech 187:424–437. https://doi.org/10.1007/s12010-018-2832-1
Maester TC, Pereira MR, Malaman AMG, Borges JP, Pereira PAM, Lemos EGM (2020) Exploring metagenomic enzymes: a novel esterase useful for short-chain ester synthesis. Catal. https://doi.org/10.3390/catal10101100
Mestrom L, Claessen JGR, Hanefeld U (2019) Enzyme-catalyzed synthesis of esters in water. ChemCatChem 11:2004–2010. https://doi.org/10.1002/cctc.201801991
Milshteyn A, Schneider JS, Brady SF (2014) Mining the metabiome: Identifying novel natural products from microbial communities. Chem Biol 21:1211–1223. https://doi.org/10.1016/j.chembiol.2014.08.006
Morris GM et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Muller H et al (2021) Discovery and design of family VIII carboxylesterases as highly efficient acyltransferases. Angew Chem Int Ed Engl 60:2013–2017. https://doi.org/10.1002/anie.202014169
Noby N, Saeed H, Embaby AM, Pavlidis IV, Hussein A (2018) Cloning, expression and characterization of cold active esterase (EstN7) from Bacillus cohnii strain N1: a novel member of family IV. Int J Biol Macromol 120:1247–1255. https://doi.org/10.1016/j.ijbiomac.2018.07.169
Oh KH et al (2012) Characterization of a novel esterase isolated from intertidal flat metagenome and its tertiary alcohols synthesis. J Mol Catal B 80:67–73. https://doi.org/10.1016/j.molcatb.2012.04.015
Park J-M, Kang C-H, Won S-M, Oh K-H, Yoon J-H (2020) Characterization of a novel moderately thermophilic solvent-tolerant esterase isolated from a compost metagenome library. Front Microbiol. https://doi.org/10.3389/fmicb.2019.03069
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786. https://doi.org/10.1038/nmeth.1701
Ramnath L, Sithole B, Govinden R (2017) Classification of lipolytic enzymes and their biotechnological applications in the pulping industry. Can J Microbiol 63:179–192. https://doi.org/10.1139/cjm-2016-0447
Rappe MS, Giovannoni SJ (2003) The uncultured microbial majority. Annu Rev Microbiol 57:369–394. https://doi.org/10.1146/annurev.micro.57.030502.090759
Roland L, James U, David E (1992) Assessment of protein models with three-dimensional profiles. Nature. https://doi.org/10.1038/356083a0
Sameh H, Soror V, Verma R, Shafaq R, John C (2007) A cold-active esterase of Streptomyces coelicolor A3(2): from genome sequence to enzyme activity. J Ind Microbiol Biot 34:525–531. https://doi.org/10.1007/s10295-007-0224-6
Sood S, Sharma A, Sharma N, Kanwar SS (2018) Carboxylesterases: Sources, characterization and broader applications. Insights Enzyme Res. https://doi.org/10.21767/2573-4466.100002
Waterhouse A et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296–W303. https://doi.org/10.1093/nar/gky427
Yang JY, Yan RX, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12:7–8. https://doi.org/10.1038/nmeth.3213
Yang C et al (2016) Discovery of new cellulases from the metagenome by a metagenomics-guided strategy. Biotechnol Biofuels 9:138. https://doi.org/10.1186/s13068-016-0557-3
Zarafeta D et al (2016) EstDZ3: A new esterolytic enzyme exhibiting remarkable thermostability. Front Microbiol. https://doi.org/10.3389/fmicb.2016.01779
Acknowledgements
This work was supported by the Key Research and Development Program of Jiangsu Province (BE2017374-2) and special funds of agro-product quality safety risk assessment of Ministry of Agriculture of the People’s Republic of China (GJFP201701505; GJFP20191101).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yan, Z., Ding, L., Zou, D. et al. Identification and characterization of a novel carboxylesterase EstQ7 from a soil metagenomic library. Arch Microbiol 203, 4113–4125 (2021). https://doi.org/10.1007/s00203-021-02398-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02398-0