Abstract
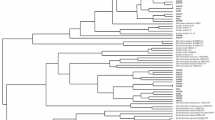
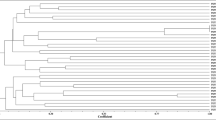
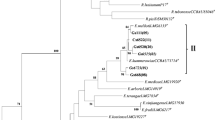
High concentrations of heavy metals in mine soil disturb the interactions between legumes and microorganisms leading to select strains adapted to these specific conditions. In this work, we analyzed the diversity of fifty strains isolated from Trifolium sp. nodules growing on Pb–Zn mine soil, in the Northeastern of Algeria and highlighted their potential symbiotic traits. The phylogeny of the 16S rRNA gene sequences revealed a high bacterial diversity with a predominance of non-rhizobial endophytes. The identified isolates belong to the thirteen following genera Cupriavidus, Pseudomonas, Bacillus, Acinetobacter, Enterobacter, Roseomonas, Paracoccus, Frondihabitans, Microbacterium, Kocuria, Providencia, Micrococcus and Staphylococcus. Regarding rhizobial strains, only isolates affiliated to Rhizobium genus were obtained. The symbiotic gene nodC and the nitrogen fixation gene nifH present showed that Rhizobium isolates belonged to the symbiovar trifolii. In addition to bacterial, one yeast strain was isolated and identified as Rhodotorula mucilaginosa by sequencing the internal transcribed spacer (ITS) region.



Similar content being viewed by others
Availability of data and materials
The data sets used and/or analysed during the current study are available from the corresponding author (Sarah Rahal) on reasonable request.
References
Abdelguerfi A, Abdelguerfi-Laouar M (2004) Genetic resources of forage and/or pastoral interest: diversity, collection and valorization at the Mediterranean level. Cah Options Mediterr 62:29–41. http://om.ciheam.org/om/pdf/c62/04600124
Amari T, Ghnaya T, Abdelly C (2017) Nickel, cadmium and lead phytotoxicity and potential of halophytic plants in heavy metal extraction. South Afric J Bot 111:99–110. https://doi.org/10.1016/j.sajb.2017.03.011
Aserse AA, Räsänen LA, Aseffa F, Hailemariam A, Lindström K (2013) Diversity of sporadic symbionts and non-symbiotic endophytic bacteria isolated from nodules of woody, shrub, and food legumes in Ethiopia. Appl Microbiol Biotechnol 97:10117–10134. https://doi.org/10.1007/s00253-013-5248-4
Borowik A, Wyszkowska J, Kucharski J, Baćmaga M, Tomkiel M (2017) Response of microorganisms and enzymes to soil contamination with a mixture of terbuthylazine, mesotrione, and S-metolachlor. Environ Sci Pollut Res 24:1910–1925. https://doi.org/10.1007/s11356-016-7919-z
Borymski S, Cycon M, Beckmann M, Mur LAJ, Piotrowska-Seget Z (2018) Plant species and heavy metals affect biodiversity of microbial communities associated with metal-tolerant plants in metalliferous soils. Front Microbiol 9:1425. https://doi.org/10.3389/fmicb.2018.01425
Cardoso P, Alves A, Silveira P, Sá C, Fidalgo C, Freitas R, Figueira E (2018) Bacteria from nodules of wild legume species: phylogenetic diversity, plant growth promotion abilities and osmotolerance. Sci Total Environ 645:1094–1102. https://doi.org/10.1016/j.scitotenv.2018.06.399
De Meyer SE, De Beuf K, Vekeman B, Willems A (2015) A large diversity of non-rhizobial endophytes found in legume root nodules in Flanders (Belgium). Soil Biol Biochem 83:1–11. https://doi.org/10.1016/j.soilbio.2015.01.002
Efrose RC, Rosu CM, Stedel C, Stefan A, Sirbu C, Gorgan LD, Labrou NE, Flemetakis E (2018) Molecular diversity and phylogeny of indigenous Rhizobium leguminosarum strains associated with Trifolium repens plants in Romania. Antonie Van Leeuwenhoek 111:135–153. https://doi.org/10.1007/s10482-017-0934-3
Efstathiadou E, Savvas D, Tampakaki AP (2020) Genetic diversity and phylogeny of indigenous rhizobia nodulating faba bean (Vicia faba L.) in Greece. Syst Appl Microbiol 43:126149. https://doi.org/10.1016/j.syapm.2020.126149
Etesami H, Beattie GA (2017) Plant-microbe interactions in adaptation of agricultural crops to abiotic stress conditions. Probiotics and Plant Health, Springer, Singapore, p 163–200
Fan M, Liu Z, Nan L, Wang E, Chen W, Lin Y, Wei G (2018) Isolation, characterization, and selection of heavy metal-resistant and plant growth-promoting endophytic bacteria from root nodules of Robinia pseudoacacia in a Pb/Zn mining area. Microbiol Res 217:51–59. https://doi.org/10.1016/j.micres.2018.09.002
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.2307/2408678
Garza-Gonzalez MT, Perez DB, Rodriguez AV, Garcia- Gutierrez DI, Zarate X, Cardenas MEC, Medina-Ruiz P (2016) Correction: Metal-induced production of a novel bioadsorbent exopolysaccharide in a native Rhodotorula mucilaginosa from the Mexican northeastern region. PLoS ONE 11:e0150522. https://doi.org/10.1371/journal.pone.0150522
Giller KE, Witter E, McGrath SP (1998) Toxicity of heavy metals to microorganisms and microbial processes in agricultural soils: a review. Soil Biol Biochem 30:1389–1414. https://doi.org/10.1016/S0038-0717(97)00270-8
Gopalakrishnan S, Sathya A, Vijayabharathi R, Varshney RK, Gowda CLL, Krishnamurthy L (2015) Plant growth promoting rhizobia: challenges and opportunities. Biotech 5:355–377. https://doi.org/10.1007/s13205-014-0241-x
Graham PH (2008) Ecology of the root-nodule bacteria of legumes. In: Dilworth MJ, James EK, Sprent JI, Newton WE (eds) Nitrogenfixing leguminous symbioses. Springer, Dordrecht, pp 23–43
Guo D, Fan Z, Lu S, Ma Y, Nie X, Tong F, Peng X (2019) Changes in rhizosphere bacterial communities during remediation of heavy metal-accumulating plants around the Xikuangshan mine in southern China. Nature 9:1947. https://doi.org/10.1038/s41598-018-38360-2
Hao X, Taghavi S, Xie P, Orbach MJ, Alwathnani HA, Rensing C, Wei G (2014) Phytoremediation of heavy and transitionmetals aided by Legume-Rhizobia Symbiosis. Int J Phytorem 16:179–202. https://doi.org/10.1080/15226514.2013.773273
Hardy RWF, Holsten RD, Jackson EK, Burns RC (1968) The acetylene-ethylene assay for N2 fixation: laboratory and field evaluation. Plant Physiol 43:1185–1207. https://doi.org/10.1104/pp.43.8.1185
Kumar S, Stecher G, Tamura K (2016) MEGA7: Mol. evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Laguerre G, Nour SM, Macheret V, Sanjuan J, Drouin P, Amarger N (2001) Classification of rhizobia based on nodC and nifH gene analysis reveals a close phylogenetic relationship among Phaseolus vulgaris symbionts. Microbiology 147:981–993. https://doi.org/10.1099/00221287-147-4-981
Leff JW, Bardgett RD, Wilkinson A, Jackson BG, Pritchard WJ, de Long JR, Oakley S, Mason KE, Ostle NJ, Johnson D, Baggs EM, Fierer N (2018) Predicting the structure of soil communities from plant community taxonomy, phylogeny, and traits. ISME J 12:1794–1805. https://doi.org/10.1038/s41396-018-0089-x
Leite J, Fischer D, Rouws LFM, Fernandes-Júnior PI, Hofmann A, Kublik S, Schloter M, Xavier GR, Radl V (2017) Cowpea nodules harbor non-rhizobial bacterial communities that are shaped by soil type rather than plant genotype. Front Plant Sci 7:2064. https://doi.org/10.3389/fpls.2016.02064
Merdas B (2006) Contribution to the geological study of the mineralizations of the Hammam N'bail region (North East Algeria). Dissertation, USTHB, Alger
Nemeth G (1959) Nature 183:1460–1461
Olenska E, Małek W (2015) Genetic differentiation of Trifolium repens microsymbionts deriving from Zn–Pb waste-heap and control area in Poland. J Basic Microbiol 55:462–470. https://doi.org/10.1002/jobm.201400604
Olenska E, Imperato V, Małek W, Włostowski T, Wójcik M, Swiecicka I, Vangronsveld J, Thijs S (2020) Trifolium repens-associated bacteria as a potential tool to facilitate phytostabilization of zinc and lead polluted waste heaps. Plants 9:1002. https://doi.org/10.3390/plants9081002
Pajuelo E, Rodríguez-Llorente ID, Lafuente A, Caviedes MÁ (2011) Legume–Rhizobium Symbioses as a Tool for Bioremediation of Heavy Metal Polluted Soils. In: Khan MS, Zaidi A, Goel R, Musarrat J (eds) Biomanagement of metal-contaminated soils. Environmental pollution, vol 20. Springer, Dordrecht, pp 95–123. https://doi.org/10.1007/978-94-007-1914-9_4
Paul K, Chinmay S, Nag M, Mandal D, Naiya H, Sen D, Mitra S, Kumar M, Bose D, Mukherjee G, Naskar N, Lahiri S, Ghosh U, Tripathi S, Sarkar MP, Banerjee M, Kleinert A, Valentine AJ, Tripathy S, Sinharoy S, Seal A (2019) Tripartite interaction among the basidiomycete Rhodotorula mucilaginosa, N2-fixing endobacteria, and rice improves nitrogen nutrition in plants. Plant Cell 32:486–507. https://doi.org/10.1105/tpc.19.00385
Prasad MNV, Freitas HMD (2003) Metal hyperaccumulation in plants—biodiversity prospecting for phytoremediation technology. Electron J Biotechnol 93:285–321. https://doi.org/10.2225/vol6-issue3-fulltext-6
Rehman K, Fatima F, Waheed I, Akash MSH (2017) Prevalence of exposure of heavy metals and their impact on health consequences. J Cell Biochem 119:157–184. https://doi.org/10.1002/jcb.26234
Román-Ponce B, Ramos-Garza J, Vásquez-Murrieta MS, RiveraOrduña FN, Chen WF, Yan J, Wang ET (2016) Cultivable endophytic bacteria from heavy metal(loid)-tolerant plants. Arch Microbiol 198:941–956. https://doi.org/10.1007/s00203-016-1252-2
Rösch C, Mergel A, Bothe H (2002) Biodiversity of denitrifying and dinitrogen-fixing bacteria in an acid forest soil. Appl Environ Microbiol 68:3818–3829. https://doi.org/10.1128/aem.68.8.3818-3829.2002
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sbabou L, Idir Y, Bruneel O, Le Quéré A, Aurag J (2016) Characterization of root-nodule bacteria isolated from Hedysarum spinosissimum L, growing in mining sites of northeastern region of Morocco. SOJ Microbiol Infect Dis 4:1–8
Sen D, Paul K, Saha C, Mukherjee G, Nag M, Ghosh S, Das A, Seal A, Tripathy S (2019) A unique life-strategy of an endophytic yeast Rhodotorula mucilaginosa JGTA-S1—a comparative genomics viewpoint. DNA Res 26:131–146. https://doi.org/10.1093/dnares/dsy044
Shetta ND, Al-Shaharani TS, Abdel-Aal M (2011) Identification and characterization of Rhizobium associated with woody legume trees grown under Saudi Arabia condition. Am J Agric Environ Sci 10:410–418
Shi Z, Zhang Z, Yuan M, Wang S, Yang M, Yao Q et al (2020) Characterization of a high cadmium accumulating soil bacterium, Cupriavidus sp. WS2. Chemosphere 247:125834. https://doi.org/10.1016/j.chemosphere.2020.125834
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, pp 611–651
Thompson JD, Higgins DG, Gibson TJ (1994) ClustalW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Wang L, Lin H, Dong Y, He Y, Liu C (2018) Isolation of vanadium resistance endophytic bacterium PRE01 from Pteris vittata in stone coal smelting district and characterization for potential use in phytoremediation. J Hazard Mater 341:1–9. https://doi.org/10.1016/j.jhazmat.2017.07.036
Wdowiak-Wrobel S, Małek W (2000) Numerical analysis of Astragalus cicer microsymbionts. Curr Microbiol 41:142–148. https://doi.org/10.1007/s002840010108
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR protocols: a guide to methods and applications, Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds). Academic Press, San Diego
Wong MH (2003) Ecological restoration of mine degraded soils, with emphasis on metal contaminated soils. Chemosphere 50:775–780. https://doi.org/10.1016/S0045-6535(02)00232-1
Xiong T, Leveque T, Shahid M, Foucault Y, Mombo S, Dumat C (2014) Lead and cadmium phytoavailability and human bioaccessibility for vegetables exposed to soil or atmospheric pollution by process ultrafine particles. J Environ Qual 43:1593–1600. https://doi.org/10.2134/jeq2013.11.0469
Yang F, Wu J, Zhang D, Chen Q, Zhang Q, Cheng XL (2018) Soil bacterial community composition and diversity in relation to edaphic properties and plant traits in grasslands of southern China. Appl Soil Ecol 128:43–53. https://doi.org/10.1016/j.apsoil.2018.04.001
Young JPW, Moeskjær S, Afonin A, Rahi P, Maluk M, James EK, Cavassim MIA, Rashid MH, Aserse AA, Perry BJ, Wang ET, Velázquez E, Andronov EE, Tampakaki A, Félix JDF, González RR, Youseif SH, Lepetit M, Boivin S, Jorrin B, Kenicer GJ, Peix A, Hynes MF, Ramírez-Bahena MH, Gulati A, Tian C (2021) Defining the Rhizobium leguminosarum species complex. Genes 12:111. https://doi.org/10.3390/genes12010111
Youseif SH, El-Megeed FHA, Mohamed AH, Ageez A, Veliz E, Martínez-Romero E (2020) Diverse Rhizobium strains isolated from root nodules of Trifolium alexandrinum in Egypt and symbiovars. Syst Appl Microbiol 44:126156. https://doi.org/10.1016/j.syapm.2020.126156
Zhang JJ, Jing XY, de Lajudie P, Ma C, He PX, Singh RP, Chen WF, Wang ET (2016) Association of white clover (Trifolium repens L.) with rhizobia of sv. trifolii belonging to three genomic species in alkaline soils in North and East China. Plant Soil 407:417–427. https://doi.org/10.1007/s11104-016-2899-9.13
Acknowledgements
We are grateful to Benjamin Gourion for hosting Sarah Rahal in Laboratory of Plant-Microbe Interactions (LIPM), for the help in the experiments and his extremely useful advice throughout this research. The authors would like to thank also Laurent Sauviac, Claire Benezech and Bryan Ruiz for their help and advice.
Funding
This work was supported by the Ministry of Higher Education and Scientific Research of Algeria. The funding bodies had no role in the design of the study and collection, analysis, and interpretation of data, or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
SR performed the experiments. SR and DC wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors state no competing interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rahal, S., Chekireb, D. Diversity of rhizobia and non-rhizobia endophytes isolated from root nodules of Trifolium sp. growing in lead and zinc mine site Guelma, Algeria. Arch Microbiol 203, 3839–3849 (2021). https://doi.org/10.1007/s00203-021-02362-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02362-y