Abstract
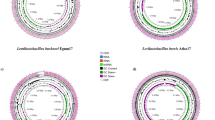
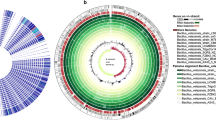
Bacillus is an excellent organic matter degrader, and it has exhibited various abilities required for lignocellulose degradation. Several B. velezensis strains encode lignocellulosases, however their ability to efficiently transform biomass has not been appreciated. In the present study, through the comparative genomic analysis of the whole genome sequences of 21 B. velezensis strains, CAZyome related to lignocellulose degradation was identified and their similarities and differences were compared. Subsequently, the secretome of B. velezensis LC1 by liquid chromatography-tandem mass spectrometry (LC–MS/MS) were identified and confirmed that a considerable number of proteins were involved in lignocellulose degradation. Moreover, after 6-day treatment, the degradation efficiency of the B. velezensis LC1 toward cellulose, hemicellulose and lignin were 59.90%, 75.44% and 23.41%, respectively, the hydrolysate was subjected to ethanol fermentation with Saccharomyces cerevisiae and Escherichia coli KO11, yielded 10.44 g/L ethanol after 96 h. These results indicate that B. velezensis LC1 has the ability to effectively degrade bamboo lignocellulose and has the potential to be used in bioethanol production.





Similar content being viewed by others
Availability of data and material
The sequence reads from this article have been deposited at the NCBI Sequence Read Archive under the accession PRJNA574012. The assembly data set supporting the results of this article has been deposited at GenBank under the accession CP044349. The version described in this paper is CP044349.
Abbreviations
- B. velezensis :
-
Bacillus velezensis
- GH:
-
Glycoside Hydrolase
- GT:
-
Glycosyltransferase
- CE:
-
Carbohydrate esterase
- CBM:
-
Carbohydrate binding domain
- PL:
-
Polysaccharide lyase;
- AA:
-
Auxiliary Activity
- CAZyme:
-
Carbohydrate-active enzyme
- ABTS:
-
[2,2′-Azino-bis (3-ethylbenzothiazoline-6-sulfonic acid)]
- BP:
-
Bamboo powder
- YPD:
-
Yeast Extract Peptone Dextrose Medium
- DNS:
-
3,5-Dinitrosalicylic acid
- CDS:
-
Sequence coding for amino acids in protein
- NCBI:
-
National Centre for Biotechnology Information
References
Adav SS, Li AA, Manavalan A, Punt P, Sze SK (2010) Quantitative iTRAQ secretome analysis of Aspergillus niger reveals novel hydrolytic enzymes. J Proteome Res 9(8):3932–3940
Belbahri L, Chenari Bouket A, Rekik I, Alenezi FN, Vallat A, Luptakova L, Petrovova E, Oszako T, Cherrad S, Vacher S, Rateb ME (2017) Comparative Genomics of Bacillus amyloliquefaciens Strains Reveals a Core Genome with Traits for Habitat Adaptation and a Secondary Metabolites Rich Accessory Genome. Front Microbiol 8:1438
Capolupo L, Faraco V (2016) Green methods of lignocellulose pretreatment for biorefinery development. Appl Microbiol Biotechnol 100(22):9451–9467
Chen L (2017) Complete genome sequence of Bacillus velezensis LM2303, a biocontrol strain isolated from the dung of wild yak inhabited Qinghai-Tibet plateau. J Biotechnol 251:124–127
Chen TY, Wen JL, Wang B, Wang HM, Liu CF, Sun RC (2017) Assessment of integrated process based on autohydrolysis and robust delignification process for enzymatic saccharification of bamboo. Bioresour Technol 244(Pt 1):717–725
Chen XH, Koumoutsi A, Scholz R, Schneider K, Vater J, Süssmuth R, Piel J, Borriss R (2009) Genome analysis of Bacillus amyloliquefaciens FZB42 reveals its potential for biocontrol of plant pathogens. J Biotechnol 140(1–2):27–37
Crowell DN, Reznikoff WS, Raetz CR (1987) Nucleotide sequence of the Escherichia coli gene for lipid A disaccharide synthase. J Bacteriol 169(12):5727–5734
Dam P, Kataeva I, Yang SJ, Zhou F, Yin Y, Chou W, Poole FL 2nd, Westpheling J, Hettich R, Giannone R, Lewis DL, Kelly R, Gilbert HJ, Henrissat B, Xu Y, Adams MW (2011) Insights into plant biomass conversion from the genome of the anaerobic thermophilic bacterium Caldicellulosiruptor bescii DSM 6725. Nucleic Acids Res 39(8):3240–3254
Goldman AR, Beer LA, Tang HY, Hembach P, Zayas-Bazan D, Speicher DW (2019) Proteome analysis using Gel-LC-MS/MS. Curr Protoc Protein Sci 96(1):e93
Gong G, Lee SM, Woo HM, Park TH, Um Y (2017) Influences of media compositions on characteristics of isolated bacteria exhibiting lignocellulolytic activities from various environmental sites. Appl Biochem Biotechnol 183(3):931–942
He P, Hao K, Blom J, Rückert C, Vater J, Mao Z, Wu Y, Hou M, He P, He Y, Borriss R (2012) Genome sequence of the plant growth promoting strain Bacillus amyloliquefaciens subsp plantarum B9601–Y2 and expression of mersacidin and other secondary metabolites. J Biotechnol 164(2):281–291
Huang XF, Santhanam N, Badri DV, Hunter WJ, Manter DK, Decker SR, Vivanco JM, Reardon KF (2013) Isolation and characterization of lignin-degrading bacteria from rainforest soils. Biotechnol Bioeng 110(6):1616–1626
Jin Q, Jiang Q, Zhao L, Su C, Li S, Si F, Li S, Zhou C, Mu Y, Xiao M (2017) Complete genome sequence of Bacillus velezensis S3–1, a potential biological pesticide with plant pathogen inhibiting and plant promoting capabilities. J Biotechnol 259:199–203
Kim SY, Lee SY, Weon HY, Sang MK, Song J (2017a) Complete genome sequence of Bacillus velezensis M75, a biocontrol agent against fungal plant pathogens, isolated from cotton waste. J Biotechnol 241:112–115
Kim SY, Song H, Sang MK, Weon HY, Song J (2017b) The complete genome sequence of Bacillus velezensis strain GH1-13 reveals agriculturally beneficial properties and a unique plasmid. J Biotechnol 259:221–227
Khelil O, Choubane S, Cheba BA (2016) Polyphenols content of spent coffee grounds subjected to physico-chemical pretreatments influences lignocellulolytic enzymes production by Bacillus sp. R2. Bioresour Technol 211:769–773
Li Y, Lei L, Zheng L, Xiao X, Tang H, Luo C (2020) Genome sequencing of gut symbiotic Bacillus velezensis LC1 for bioethanol production from bamboo shoots. Biotechnol Biofuels 13:34
Liu G, Kong Y, Fan Y, Geng C, Peng D, Sun M (2017) Whole-genome sequencing of Bacillus velezensis LS69, a strain with a broad inhibitory spectrum against pathogenic bacteria. J Biotechnol 249:20–24
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42(Database issue):D490-495
Luo C, Li Y, Liao H, Yang Y (2018) De novo transcriptome assembly of the bamboo snout beetle Cyrtotrachelus buqueti reveals ability to degrade lignocellulose of bamboo feedstock. Biotechnol Biofuels 11:292
Mai-Gisondi G, Maaheimo H, Chong SL, Hinz S, Tenkanen M (1861) Master E (2017) Functional comparison of versatile carbohydrate esterases from families CE1, CE6 and CE16 on acetyl-4-O-methylglucuronoxylan and acetyl-galactoglucomannan. Biochim Biophys Acta Gen Subj 9:2398–2405
Mansour AA, Da Costa A, Arnaud T, Lu-Chau TA, Fdz-Polanco M, Moreira MT, Cacho Rivero JA (2016) Review of lignocellulolytic enzyme activity analyses and scale-down to microplate-based assays. Talanta 150:629–637
McCarthy T, Hanniffy O, Savage A, Tuohy MG (2003) Catalytic properties and mode of action of three endo-β-glucanases from Talaromyces emersonii on soluble β-1, 4-and β-1, 3, 1, 4-linked glucans. Int J Biol Macromol 33:141–148
Molinatto G, Puopolo G, Sonego P, Moretto M, Engelen K, Viti C, Ongena M, Pertot I (2016) Complete genome sequence of Bacillus amyloliquefaciens subsp. plantarum S499, a rhizobacterium that triggers plant defences and inhibits fungal phytopathogens. J Biotechnol 238:56–59
Mu YY, Qi WP, Zhang T, Zhang JY, Mao SY (2021) Gene function adjustment for carbohydrate metabolism and enrichment of rumen microbiota with antibiotic resistance genes during subacute rumen acidosis induced by a high-grain diet in lactating dairy cows. J Dairy Sci 104(2):2087–2105
Niazi A, Manzoor S, Asari S, Bejai S, Meijer J, Bongcam-Rudloff E (2014) Genome analysis of Bacillus amyloliquefaciens Subsp. plantarum UCMB5113: a rhizobacterium that improves plant growth and stress management. PLoS One 9(8):e104651.
Pan HQ, Li QL, Hu JC (2017) The complete genome sequence of Bacillus velezensis 9912D reveals its biocontrol mechanism as a novel commercial biological fungicide agent. J Biotechnol 247:25–28
Pardo M, Ward M, Bains S, Molina M, Blackstock W, Gil C, Nombela C (2000) A proteomic approach for the study of Saccharomyces cerevisiae cell wall biogenesis. Electrophoresis 21(16):3396–3410
See-Too WS, Chua KO, Lim YL, Chen JW, Convey P, Mohd Mohidin TB, Yin WF, Chan KG (2017) Complete genome sequence of Planococcus donghaensis JH1T, a pectin-degrading bacterium. J Biotechnol 252:11–14
Sharma A, Tewari R, Rana SS, Soni R, Soni SK (2016) Cellulases: Classification, Methods of Determination and Industrial Applications. Appl Biochem Biotechnol 179(8):1346–1380
Sluiter A, Hames B, Ruiz R, Scarlata C (2012) Sluiter J, Templeton D, et al. Determination of structural carbohydrates and lignin in biomass. Version 2012. National Renewable Energy Laboratory, USA.
Sun SN, Cao XF, Zhang XM, Xu F, Sun RC, Jones GL (2014) Characteristics and enzymatic hydrolysis of cellulose-rich fractions from steam exploded and sequentially alkali delignified bamboo (Phyllostachys pubescens). Bioresour Technol 163:377–380
Teng D, Wang JH, Fan Y, Yang YL, Tian ZG, Luo J, Yang GP, Zhang F (2006) Cloning of β-1, 3–1, 4-glucanase gene from Bacillus licheniformis EGW039 (CGMCC 0635) and its expression in Escherichia coli BL21 (DE3). Appl Microbiol Biotechnol 72:705–712
Thompson AJ, Speciale G, Iglesias-Fernández J, Hakki Z, Belz T, Cartmell A, Spears RJ, Chandler E, Temple MJ, Stepper J, Gilbert HJ, Rovira C, Williams SJ, Davies GJ (2015) Evidence for a boat conformation at the transition state of GH76 α-1,6-mannanases–key enzymes in bacterial and fungal mannoprotein metabolism. Angew Chem Int Ed Engl 54(18):5378–5382
Torpenholt S, Le Nours J, Christensen U, Jahn M, Withers S, Ostergaard PR, Borchert TV, Poulsen JC, Lo Leggio L (2011) Activity of three β-1,4-galactanases on small chromogenic substrates. Carbohydr Res 346(13):2028–2033
Vanholme B, Jacob J, Cannoot B, Gheysen G, Haegeman A (2009) Arabinogalactan endo-1,4-β-galactosidase: a putative plant cell wall-degrading enzyme of plant-parasitic nematodes. Nematology 11(5):739–747
Van Soest PJ, Robertson JB, Lewis BA (1991) Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J Dairy Sci 74(10):3583–3597
Wi SG, Lee DS, Nguyen QA, Bae HJ (2017) Evaluation of biomass quality in short-rotation bamboo (Phyllostachys pubescens) for bioenergy products. Biotechnol Biofuels 10:127
Woo HL, Hazen TC, Simmons BA, DeAngelis KM (2014) Enzyme activities of aerobic lignocellulolytic bacteria isolated from wet tropical forest soils. Syst Appl Microbiol 37(1):60–67
Yang W, Bai Y, Yang P, Luo H, Huang H, Meng K, Shi P, Wang Y, Yao B (2015) A novel bifunctional GH51 exo-α-l-arabinofuranosidase/endo-xylanase from Alicyclobacillus sp. A4 with significant biomass-degrading capacity. Biotechnol Biofuels 8:197
Yin Y, Mao X, Yang J, Chen X, Mao F, Xu Y (2012) dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40(Web Server issue):W445-451
Yuan Z, Wen Y (2017) Evaluation of an integrated process to fully utilize bamboo biomass during the production of bioethanol. Bioresour Technol 236:202–211
Yuan Z, Wen Y, Kapu NS (2018) Ethanol production from bamboo using mild alkaline pre-extraction followed by alkaline hydrogen peroxide pretreatment. Bioresour Technol 247:242–249
Zhang J, Siika-Aho M, Tenkanen M, Viikari L (2011) The role of acetyl xylan esterase in the solubilization of xylan and enzymatic hydrolysis of wheat straw and giant reed. Biotechnol Biofuels 4(1):60
Zhao L, Cao GL, Wang AJ, Ren HY, Dong D, Liu ZN, Guan XY, Xu CJ, Ren NQ (2012) Fungal pretreatment of cornstalk with Phanerochaete chrysosporium for enhancing enzymatic saccharification and hydrogen production. Bioresour Technol 114:365–369
Zhu D, Tanabe SH, Xie C, Honda D, Sun J, Ai L (2014) Bacillus ligniniphilus sp. Nov., an alkaliphilic and halotolerant bacterium isolated from sediments of the South China Sea. Int J Syst Evol Microbiol 64(Pt 5):1712–1717
Acknowledgements
The data were analyzed on the free online platform of Majorbio Cloud Platform (www.majorbio.com).
Funding
This work was supported by Sichuan science and technology program (2019YFG0139) and Scientific Research Foundation of Leshan Normal University (XJR19001).
Author information
Authors and Affiliations
Contributions
CBL designed and performed the experiments; HT, LZ and YQL wrote the manuscript; LZ, LL, YQL, XWY and HT analyzed the data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This study does not include any experimental procedure performed on humans or animals.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tang, H., Zheng, L., Li, Y. et al. Comparative genomic and secretomic characterisation of endophytic Bacillus velezensis LC1 producing bioethanol from bamboo lignocellulose. Arch Microbiol 203, 3089–3099 (2021). https://doi.org/10.1007/s00203-021-02306-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02306-6