Abstract
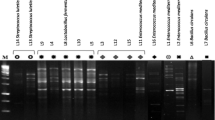
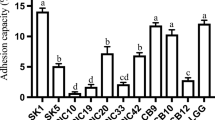
Consumer’s vigilance towards health-promoting foods beyond only taste and nutrition has increased the recognition for probiotic products. In the present study, various parameters have been studied to define the probiotic properties of cultures isolated from different fermented products. Around 118 samples were selectively screened for antimicrobial compound (AMC) producing isolates by overlay-plate assay using Micrococcus luteus ATCC9341. Among 134 zone producing isolates, 48 cultures showing Gram-positive, catalase negative, non-spore forming and non-motile rods and cocci were selected. Subsequently, 18 strains were chosen based on non-hemolytic, absence of biogenic amine production, gelatinase and lecithinase negative trait for safer isolates. These were identified by biochemical assays and then subjected to RAPD-PCR. The selected cultures DB-1aa, DB-b2-15b, Cu2-PM7, Cu3-PM8 and IB-pM15 were identified by 16S rDNA sequencing as Enterococcus durans, Enterococcus faecium, Lactobacillus plantarum, and two Lactobacillus fermentum, respectively. Several in vitro experiments were carried out including acid and bile tolerance, survival under simulated gastrointestinal condition, adhesion assay to evaluate the probiotic potential of the isolates. In addition, the isolates were studied for competent properties such as antibacterial, antioxidant activity, and enzyme production for their functional application. The results of the study prove the efficiency of selected isolates as potential probiotic cultures and hence can be recommended for application in any functional food formulations.




Similar content being viewed by others
References
Altschul SF, Maddan TL, Schaffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bauer RW, Kirby MDK, Sherris JC et al (1996) Antibiotic susceptibility testing by standard single disc diffusion method. Am J Clin Pathol 45:493–496
Chang M, Chang HC (2012) Development of a screening method for biogenic amine producing Bacillus spp. Int J Food Microbiol 153:269–274
Del Re B, Sgorbati B, Miglioli M et al (2000) Adhesion, autoaggregation and hydrophobicity of 13 strains of Bifidobacterium longum. Lett Appl Microbiol 31:438–442
Devi SM, Halami PM (2011) Detection and characterization of pediocin PA-1/AcH like bacteriocin producing lactic acid bacteria. Curr Microbiol 62:181–185
Devi SM, Archer AC, Halami PM (2015) Screening, characterization and in vitro evaluation of probiotic properties among lactic acid bacteria through comparative analysis. Probiotics Antimicrob Proteins 7(3):181–192
Dunne C, O’Mahony L, Murphy L et al (2001) In vitro selection criteria for probiotic bacteria of human origin: correlation with in vivo findings. Am J Clin Nutr 73:386S–392S
FAO/WHO (2001) Report of a Joint FAO/WHO expert consultation on evaluation of health nutritional properties of probiotics in food including powder milk with live lactic acid bacteria. World Health Organization and Food and Agriculture Organization of the United Nations, London, Ontario, Canada
Grosu-Tudor SS, Stancu MM, Pelinescu D et al (2014) Characterization of some bacteriocins produced by lactic acid bacteria isolated from fermented foods. World J Microbiol Biotechnol 30:2459–2469
Guan X, Xu Q, Zheng Y et al (2017) Screening and characterization of lactic acid bacterial strains that produce fermented milk and reduce cholesterol levels. Braz J Microbiol 48:730–739
Gusils C, Gonzalez S, Oliver G (1999) Some probiotic properties of chicken lactobacilli. Can J Microbiol 45:981–987
Handa S, Sharma N (2016) In vitro study of probiotic properties of Lactobacillus plantarum F22 isolated from chhang—a traditional fermented beverage of Himachal Pradesh, India. J Genetic Engineer Biotechnol 14:91–97
Ho WY, Beh BK, Lim KL et al (2015) Antioxidant and hepatoprotective effects of the food seasoning curry leaves Murraya Koenigii (L.) Spreng (Rutaceae). RSC Adv 5:100589–100597. https://doi.org/10.1039/C5RA19154H
Huang DJ, Ou BX, Prior RL (2005) The chemistry behind antioxidant capacity assays. J Agric Food Chem 53:1841–1856
Huidrom S, Sharma N (2018) Isolation and screening of novel isolates of Bifidobacteria from human milk as potential probiotic with antidiarrheal activity. Ann Appl Microbiol Biotechnol J 2(1):1007–1015
Islam KB, Fukiya S, Hagio M et al (2011) Bile acid is a host factor that regulates the composition of the cecal microbiota in rats. Gastroenterol 141:1773–1781
Jacobsen CN, Rosenfeldt Nielsen V, Hayford AE et al (1999) Screening of probiotic activities of forty-seven strains of Lactobacillus spp. by in vitro techniques and evaluation of the colonization ability of five selected strains in humans. Appl Environ Microbiol 65:4949–4956
Jacobsen L, Wilcks A, Hammer K, Huys G, Gevers D, Andersen SR (2007) Horizontal transfer of tet(M) and erm(B) resistance plasmids from food strains of Lactobacillus plantarum to Enterococcus faecalis JH2-2 in the gastrointestinal tract of gnotobiotic rats. FEMS Microbiol Ecol 59:158–166
Jatmiko YD, Howarth GS, Barton MD (2017) Assessment of probiotic properties of lactic acid bacteria isolated from Indonesian naturally fermented milk. In: AIP conference proceedings 1908, 050008; https://doi.org/10.1063/1.5012732
Kanmani P, Kumar RS, Yuvaraj N et al (2013) Probiotics and its functionality valuable products—a review. Crit Rev Food Sci Nut 53:641–658
Kaushik JK, Kumar A, Duary RK et al (2009) Functional and probiotic attributes of an indigenous isolate of Lactobacillus plantarum. PLoS ONE 4:1–11
Kos B, Suskovic J, Vukovic S et al (2003) Adhesion and aggregation ability of probiotic strain Lactobacillus acidophilus M92. J Appl Microbiol 94:981–987
Kumar S, Nei M, Dudley J et al (2008) Mega: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief Bioinform 9:299–306
Lin M, Chang F (2000) Antioxidative effect of intestinal bacteria Bifidobacterium longum ATCC 15708 and Lactobacillus acidophilus ATCC 4356. Dig Dis Sci 45:1617–1622
Lin MY, Yen CL (1999) Antioxidant ability of lactic acid bacteria. J Agri Food Chem 47(4):1460–1466
Liu C, Pan T (2010) In Vitro effects of lactic acid bacteria on cancer cell viability and antioxidant activity. J Food Drug Anal 18:77–86
Lopes MFS, Simoes AP, Tenreiro R et al (2007) Activity and expression of a virulence factor, gelatinase in dairy enterococci. Int J Food Microbiol 112(3):208–214
Martinez-Cayuela M (1995) Oxygen free radicals and human disease. Biochimie 77:147–161
Maryam AS, Abubakr (2018) Antimicrobial activities of lactic acid bacteria strains isolated from human breast milk against human pathogenic strains. Int J Clin Dev Anat 4(1):27–31. https://doi.org/10.11648/j.ijcda.20180401.14
Mashak K (2016) Antimicrobial activity of lactobacillus isolated from kashk-e zard and tarkhineh, two Iranian traditional fermented foods. Int J Enteric Pathog 4:e34692
Mathara JM, Schillinger U, Kutima PM et al (2008) Functional properties of Lactobacillus plantarum strains isolated from Maasai traditional fermented milk products in Kenya. Curr Microbiol 56:315–321
Mauriello G, De Luca E, La Storia A et al (2005) Antimicrobial activity of a nisin-activated plastic film for food packaging. Lett Appl Microbiol 41:464–469
Mora D, Fortina MG, Parini C et al (2000) Genomic sub-populations within the species Pediococcus acidilactici detected multilocus typing analysis: relationships between pediocin AcH/PA-1 producing and non-producing strains. Microbiology 146:2027–2038
Nishiyama K, Nakazato A, Ueno S et al (2015) Cell surface-associated aggregation-promoting factor from Lactobacillus gasseri SBT2055 facilitates host colonization and competitive exclusion of Campylobacter jejuni. Mol Microbiol 98:712–726
Nivoliez A, Veisseire P, Alaterre E et al (2015) Influence of manufacturing processes on cell surface properties of probiotic strain Lactobacillus rhamnosus Lcr35®. Appl Microbiol Biotechnol 99:399–411
Oh NS, Lee JY, Oh S et al (2016) Improved functionality of fermented milk is mediated by the synbiotic interaction between Cudrania tricuspidata leaf extract and Lactobacillus gasseri strains. Appl Microbiol Biotechnol 100(13):5919–5932
Ouoba LII, Lei V, Jensen LB (2008) Resistance of potential probiotic lactic acid bacteria and bifidobacteria of African and European origin to antimicrobials: Determination and transferability of the resistance genes to other bacteria. Int J Food Microbiol 121(2):217–224
Otero MC, Ocana VS, Macias ENM (2004) Bacterial surface characteristics applied to selection of probiotic microorganisms. Method Mol Biol 268:435–440
Ouwehand AC, Kirjavainen PV, Shortt C et al (1999) Probiotics: mechanisms and established effects. Int Dairy J 9(1):43–52
Peiniz S, Andreazza R, Anghinoni T et al (2014) Probiotic potential, antimicrobial and antioxidant activities of Enterococcus durans strain LAB18s. Food Control 37:251–256
Prasad J, Gill H, Smart J et al (1998) Selection and characterization of Lactobacillus and Bifidobacterium strains for use as probiotics. Int Dairy J 8:993–1002
Raghavendra P, Halami PM (2009) Screening, selection and characterization of phytic acid degrading lactic acid bacteria from chicken intestine and their probiotic properties. Int J Food Microbiol 133:129–134
Rahman SMK (2015) Probiotic properties analysis of isolated lactic acid bacteria from buffalo milk. Arch Clin Microbiol 7:1
Rastogi S, Mittal V, Singh A (2019) In vitro evaluation of probiotic potential and safety assessment of Lactobacillus mucosae strains isolated from Donkey’s lactation. Probiotics Antimicrob Proteins. https://doi.org/10.1007/s12602-019-09610-0
Reid G, Hass J, Sebulsdy MT et al (2003) Potential use of probiotics in clinical practice. Clin Microbiol Rev 16:658–672
Rosenberg M, Gutnick D, Rosenberg E (1980) Adherence of bacteria to hydrocarbons: a simple method for measuring cell-surface hydrophobicity. FEMS Microbiol Lett 9:29–33
Rossini K, Noreña CPZ, Olivera FC et al (2009) Casein peptides with inhibitory activity on lipid oxidation in beef homogenates and mechanically deboned poultry meat. LWT Food Sci Technol 42:862–867
Rowan NJ, Deans K, Anderson JG et al (2001) Putative virulence factor expression by clinical and food isolates of Bacillus spp. after growth in reconstituted infant milk formulae. Appl Environ Microbiol 67(9):3873–3881
Sahadeva RPK, Leong SF, Chua KH et al (2011) Survival of commercial probiotic strains to pH and bile. Int Food Res J 18(4):1515–1522
Sarem F, Sarem-Damerdji LO, Nicolas JP (1996) Comparison of the adherence of three Lactobacillus strains to Caco-2 and Int-407 human intestional cell lines. Lett Appl Microbiol 22:439–442
Schär-Zammaretti P, Ubbink J (2003) The cell wall of lactic acid bacteria: surface constituents and macromolecular conformations. Biophys J 85:4076–4092
Shahbazi S, Nateghi L, Aghababyan A (2016) Effect of fatty acids on hydrophobicity of the cell membrane of Lactobacillus species. Appl Food Biotechnol 3:194–200
Sharaf EF, El-Sayed WS, Abosaif RM (2014) Lecithinase-producing bacteria in commercial and home-made foods: evaluation of toxic properties and identification of potent producers. J Taibah Univ Sci 8(3):207–215
Shehata MG, El Sohaimy SA, El-Sahn MA et al (2016) Screening of isolated potential probiotic lactic acid bacteria for cholesterol lowering property and bile salt hydrolase activity. Ann Agri Sci 61(1):65–75
Son SH, Jeon HL, Yang SJ et al (2017) Probiotic lactic acid bacteria isolated from traditional Korean fermented foods based on ß-glucosidase activity. Food Sci Biotechnol 27:123–129
Soomro A, Masud T, Anwaar K (2002) Role of lactic acid bacteria (LAB) in food preservation and human health—a review. PJN 1(1):20–24. https://doi.org/10.3923/pjn.2002.20.24
Sullivan A, Nord CE (2002) The place of probiotics in human intestinal infections. Int J Antimicrob Agents 20(5):313–319. https://doi.org/10.1016/S0924-8579(02)00199-1
Talib N, Mohamad NE, Yeap SK et al (2019) Isolation and characterization of Lactobacillus spp. From kefir samples in Malaysia. Molecules 24(14):2606
Tulumoglu S, Yuksekdag ZN, Beyatli Y et al (2013) Probiotic properties of lactobacilli species isolated from children’s feces. Anaerobe 24:36–42
Tuo Y, Yu H, Ai L et al (2013) Aggregation and adhesion properties of 22 Lactobacillus strains. J Dairy Sci 96:4252–4257
Valentao P, Fernandes E, Carvalho F et al (2002) Antioxidatives properties of cardoon (Cynara cardunculus L.) infusion against superoxide radical, hydroxyl radical, and hypochlorous acid. J Agri Food Chem 50:4989–4993
Virtanen T, Pihlanto A, Akkanen S et al (2007) Development of antioxidant activity in milk whey during fermentation with lactic acid bacteria. J Appl Microbiol 102(1):106–115
Wang B, Wei H, Yuan J et al (2008) Identification of a surface protein from Lactobacillus reuteri JCM1081 that adheres to porcine gastric mucin and human enterocyte-like HT-29 Cells. Curr Microbiol 57:33–38
Waters CM, Antiporta MH, Murray BE et al (2003) Role of the Enterococcus faecalis GelE protease in determination of cellular chain length, supernatant pheromone levels, and degradation of fibrin and misfolded surface proteins. J Bacteriol 185:3613–3623
Wescombe PA, Hale JD, Heng NC et al (2012) Developing oral probiotics from Streptococcus salivarius. Future Microbiol 7(12):1355–1371
Wu Q, Shah NP (2014) Effects of elaidic acid, a predominant industrial trans fatty acid, on bacterial growth and cell surface hydrophobicity of lactobacilli. J Food Sci 79:M2485–M2490
Wu S, Sun MZ, Zhang C et al (2014) Antioxidant properties of Lactobacillus and its protecting effects to oxidative stress Caco-2 cells. The J Ani Plant Sci 24(6):1766–1771
Xie J, Zhang R, Shang C et al (2009) Isolation and characterization of a bacteriocin produced by an isolated Bacillus subtilis LFB112 that exhibits antimicrobial activity against domestic animal pathogens. Afr J Biotechnol 8(20):5611–5619
Xing J, Wang G, Zhang Q et al (2015) Determining antioxidant activities of Lactobacilli cell-free supernatants by cellular antioxidant assay: a comparison with traditional methods. PLoS ONE 10(3):e0119058
Yong L, Yong Z, Yan B et al (2010) Study on cell surface properties and inhibitory effects on pathogens of four probiotic strains. J Chin Institute Food Sci Technol 10:28–34
Zárate G, Chaia AP, González S et al (2000) Viability and β-galactosidase activity of dairy propionibacteria subjected to digestion by artificial gastric and intestinal fluids. J Food Prot 63:1214–1221
Acknowledgements
This work was support by University Grants Commission for the award of NFSC research fellowship
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

203_2020_2037_MOESM1_ESM.jpg
Representative plates of (a) LAB isolation on MRS media and (b) Selection of zone producer by over-laying with Micrococcus luteus. (JPG 41 kb)

203_2020_2037_MOESM2_ESM.jpg
Representative pictures showing (a) Hemolytic activity (b) Lecithinase activity (c) Gelatin hydrolysis and (d) Biogenic amine production. (JPG 55 kb)
Rights and permissions
About this article
Cite this article
Bindu, A., Lakshmidevi, N. Identification and in vitro evaluation of probiotic attributes of lactic acid bacteria isolated from fermented food sources. Arch Microbiol 203, 579–595 (2021). https://doi.org/10.1007/s00203-020-02037-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02037-0