Abstract
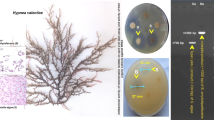
Amoxicillin-resistant bacteria were isolated using selective enrichment procedure. The morphological, biochemical and molecular characterization based on 16S rRNA gene sequencing and phylogenetic analysis of the bacterial strain WA5 confirmed that the strain belongs to the genus Stenotrophomonas. The bacteria were named as Stenotrophomonas sp. strain WA5 (MK110499). Substantial growth was seen in M9 minimal media supplemented with 5 mg L−1 of amoxicillin as a sole source of carbon and energy. RNA yield was also observed to be decreased in the presence of amoxicillin. Amoxicillin (5 mg L−1)-induced alteration is seen on bacterial protein profile and unique polypeptide bands were seen to be induced in the presence of amoxicillin, the bands were subjected to trypsin digestion, and LC–MS/MS analysis showed that the bands belong to the family of DNA-dependent RNA polymerase subunit β (rpoC). Plasmid DNA isolation indicated the presence of antibiotic-resistant genes being harboured by the plasmid.





Similar content being viewed by others
References
Baquero F, Martínez JL, Cantón R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Berendonk TU, Manaia CM, Merlin C, Fatta-Kassinos D, Cytryn E, Walsh F, Bürgmann H, Sørum H, Norström M, Pons MN, Kreuzinger N (2015) Tackling antibiotic resistance: the environmental framework. Nat Rev Microbiol 13:307–310
Bovio E, Gnavi G, Prigione V, Spina F, Denaro R, Yakimov M, Calogero R, Crisafi F, Varese GC (2017) The culturable mycobiota of a mediterranean marine site after an oil spill: isolation, identification and potential application in bioremediation. Sci Total Environ 576:310–318
Fair RJ, Tor Y (2014) Antibiotics and bacterial resistance in the 21st century. Perspect Med Chem 6:25–64
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Forni C, Cascone A, Cozzolino S, Migliore L (2001) Drugs uptake and degradation by aquatic plants as a bioremediation technique. Minerva Biotecnologica 13:151–152
Ghauch A, Tuqan A, Assi HA (2009) Antibiotic removal from water: elimination of amoxicillin and ampicillin by microscale and nanoscale iron particles. Environ Pollut 157:1626–1635
Goldstein BP (2014) Resistance to rifampicin: a review. J Antibiot 67:625–630
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kumar M, Jaiswal S, Sodhi KK, Shree P, Singh DK, Agrawal PK, Shukla P (2019a) Antibiotics bioremediation: perspectives on its ecotoxicity and resistance. Environ Int 124:448–461
Kumar M, Sodhi KK, Singh P, Agrawal PK, Singh DK (2019b) Synthesis and characterization of antibiotic-metal complexes [FeCl3 (L1) 2H2O and Ni (NO3) 2 (L2) 2H2O] and enhanced antibacterial activity. Environ Nanotechnol Monit Manage 11:100209
Oteo J, Saez D, Bautista V, Fernández-Romero S, Hernández-Molina JM, Pérez-Vázquez M, Campos J (2013) Carbapenemase-producing Enterobacteriaceae in Spain in 2012. Antimicrob Agents Chemother 57:6344–6347
Reddy AVB, Yusop Z, Jaafar J, Reddy YVM, Aris AB, Majid ZA, Talib J, Madhavi G (2016) Recent progress on Fe-based nanoparticles: Synthesis, properties, characterization and environmental applications. J Environ Chem Eng 4:3537–3553
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sharma S, Bhattacharya A (2017) Drinking water contamination and treatment techniques. Appl Water Sci 7:1043–1067
Singer AC, Shaw H, Rhodes V, Hart A (2016) Review of antimicrobial resistance in the environment and its relevance to environmental regulators. Front microbiol 7:1728
Singh H, Rathore RS, Singh S, Cheema PS (2011) Comparative analysis of cultural isolation and PCR based assay for detection of Campylobacter jejuni in food and faecal samples. Braz J Microbiol 42:181–186
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci 101:11030–11035
Williams ST, Sharpe ME, Holt JG (eds) (1989) Bergey’s Manual of Systematic Bacteriology, 1st edn, vol 4. Williams and Wilkins, Baltimore
Xu M, Zhou YN, Goldstein BP, Jin DJ (2005) Cross-resistance of Escherichia coli RNA polymerases conferring rifampin resistance to different antibiotics. J Bacteriol 187:2783–2792
Acknowledgements
Financial assistance provided by NASF research grant (project entitled “Bioremediation of chemical contaminants and their complexes present in drainage water with high dynamic flux used for irrigation in urban and peri urban agriculture”), sanction no. NASF/CA-6030/2017-18 is highly acknowledged. The authors Kushneet Kaur Sodhi and Mohit Kumar highly acknowledges the University Grant Commission (UGC), Government of India for providing the stipend.
Funding
This study was supported by the funding agency, National Agricultural Science Fund, Indian Council of Agricultural Research, Delhi, India.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflicts of interest.
Human and animals rights
The study is not related with animals or humans.
Informed consent
Not applicable.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sodhi, K.K., Kumar, M., Balan, B. et al. Isolation and characterization of amoxicillin-resistant bacteria and amoxicillin-induced alteration in its protein profiling and RNA yield. Arch Microbiol 202, 225–232 (2020). https://doi.org/10.1007/s00203-019-01737-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-019-01737-6