Abstract
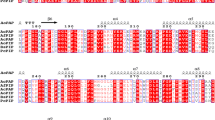
Prephenate dehydratase is a key regulatory enzyme in the phenylalanine-specific pathway of Corynebacterium glutamicum. PCR-based random mutagenesis and functional complementation were used to screen for m-fluorophenylalanine (mFP)-resistant mutants. Comparison of the amino acid sequence of the mutant prephenate dehydratases indicated that Ser-99 plays a role in the feedback regulation of the enzyme. When Ser-99 of the wild-type enzyme was replaced by Met, the specific activity of the mutant enzyme was 30% lower than that of the wild-type. The Ser99Met mutant was active in the presence of 50 μM phenylalanine, whereas the wild-type enzyme was not. The functional roles of the eight conserved residues of prephenate dehydratase were investigated by site-directed mutagenesis. Glu64Asp substitution reduced enzyme activity by 15%, with a 4.5- and 1.7-fold increase in K m and k cat values, respectively. Replacement of Thr-183 by either Ala or Tyr resulted in a complete loss of enzyme activity. Substitution of Arg-184 with Leu resulted in a 50% decrease of enzyme activity. The specific activity for Phe185Tyr was more than 96% lower than that of the wild-type, and the K m value was 26-fold higher. Alterations in the conserved Asp-76, Glu-89, His-115, and Arg-236 residues did not cause a significant change in the K m and k cat values. These results indicated that Glu-64, Thr-183, Arg-184, and Phe-185 residues might be involved in substrate binding and/or catalytic activity.


Similar content being viewed by others
References
Ahmad S, Jensen RA (1986) The evolutionary history of two bifunctional proteins that emerged in the purple bacteria. Trends Biochem Sci 11: 108–112
Ahmad S, Wilson AT, Jensen RA (1988) Chorismate mutase:prephenate dehydratase from Acinetobacter calcoaceticus. Purification, properties and immunological cross-reactivity. Eur J Biochem 116: 69–79
Baldwin GS, Davidson BE (1981) A kinetic and structural comparison of chorismate mutase/prephenate dehydratase from mutant strains of Escherichia coli K12 defective in the pheA gene. Arch Biochem Biophys 211:66-75
Baldwin GS, Mckenzie GH, Davidson BE (1981) The self-association of chorismate mutase/prephenate dehydratase from Escherichia coli K12. Arch Biochem Biophys 211: 76–85
Bathe B, Kalinowski J, Pühler A (1996) A physical and genetic map of the Corynebacterium glutamicum ATCC 13032 chromosome. Mol Gen Genet 252: 255–265
Byng GS, Jensen RA (1983) Impact of isozymes upon partitioning of carbon flow and regulation of aromatic biosynthesis in prokaryotes. In: Ratazzi MC, Scandalios JG, Whitt GS (eds) Isozymes, vol. 8, Alan R. Liss, New York, pp 115–140
Cadwell RC, Joyce CF (1992) Randomization of genes by PCR mutagenesis. PCR Methods Appl 2: 28–33
Chan MS, Hsu WH (1996) Cloning of m-fluorophenylalanine-resistant gene and mutational analysis of feedback-resistant prephenate dehydratase from Corynebacterium glutamicum. Biochem Biophys Res Comm 219: 537–542
Chook YM, Gray JV, Ke H, Lipscomb WN (1994) The monofunctional chorismate mutase from Bacillus subtilis. Structure determination of chorismate mutase and its complexes with a transition state analog and prephenate, and implications for the mechanism of the enzymatic reaction. J Mol Biol 240:476–500
Davidson BE, Blackburn EH, Dopheide TA (1972) Chorismate mutase-prephenate dehydratase from Escherichia coli K-12. I. Purification, molecular weight, and amino acid composition. J Biol Chem 247:4441–4446
Dopheide TA, Crewther P, Davidson BE (1972) Chorismate mutase-prephenate dehydratase from Escherichia coli K-12. II. Kinetic properties. J Biol Chem 247: 4447–4452
Dougherty DA (1996) Cation-pi interactions in chemistry and biology: a new view of benzene, Phe, Tyr, and Trp. Science 271:163–168
Euverink GJW, Wolters DJ, Dijkhuizen L (1995) Prephenate dehydratase of the actinomycete Amycolatopsis methanolica: purification and characterization of wild-type and deregulated mutant proteins. Biochem J 308: 313–320
Fazel RS, Jensen RA (1980) Regulation of prephenate dehydratase in coryneform species of bacteria by L-phenylalanine and by remote effectors. Arch Biochem Biophys 200: 165–176
Fermandez-Recio J, Romero A, Sancho J (1999) Energetics of a hydrogen bond (charged and neutral) and of a cation-pi interaction in apoflavodoxin. J Mol Biol 290:319–330
Follettie MT, Sinsky AJ (1986) Molecular cloning and nucleotide sequence of the Corynebacterium glutamicum pheA gene. J Bacteriol 167: 659–702
Gething MJH, Davidson BE (1976) Chorismate mutase/prephenate dehydratase from Escherichia coli K12. 2. Evidence for identical subunits catalyzing the two activities. Eur J Biochem 71: 327–336
Hagino H, Nakatama K (1974) L-Phenylalanine production by an analogue-resistant mutant of Corynebacterium glutamicum. Agric Biol Chem 38:157–161
Ikeda M, Katsumata R (1992) Metabolic engineering to produce tyrosine or phenylalanine in a tryptophan-producing Corynebacterium glutamicum strain. Appl Environ Microbiol 58: 781–785
Jetten MS, Sinsky AJ (1995) Recent advances in the physiology and genetics of amino acid-producing bacteria. Crit Rev Biotechnol 15:73–103
Kuninka A (1986) Nucleic acid, nucleotides and related compounds. In: Rehm HJ, Reed G (eds) Biotechnology vol. 4. Microbial production. Neenbeem VCH Verlagsyese II Schaff, Weinheim, Germany pp 71–117
Kunkel TA, Roberts JD, Zakour RA (1987) Rapid and efficient site-specific mutagenesis without phenotype selection. Methods Enzymol 154: 367–38
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685
Lee S, Lin X, McMurray J, Sun G (2002) Contribution of an active site cation-pi interaction to the spectroscopic properties and catalytic function of protein tyrosine kinase Csk. Biochemistry 41:12107–12114
Liebl W, Ehrmann M, Ludwig W, Schleifer KH (1991) Transfer of Brevibacteria divaricatum DSM 20297, Brevibacteria flavum DSM 20412 and DSM 1412 and Corynebacterium lilium DSM 20137 to Corynebacterium glutamicum and their distinction by rRNA gene restriction patterns. Int J Syst Bacteriol 41: 225–260
Nelms J, Edwards RM, Warwick J, Fotheringham I (1992) Novel mutations in the pheA gene of Escherichia coli K-12 which result in highly feedback inhibition-resistant variants of chorismate mutase/prephenate dehydratase. Appl Environ Microbiol 58:2592–2598
Pierson DL, Jensen RA (1974) Metabolic interlock: control of an interconvertible prephenate dehydratase by hydrophobic amino acid in Bacillus subtilis. J Mol Biol 252: 5839–5846
Pohnert G, Zhang S, Husain A, Wilson DB, Ganem B (1999) Regulation of phenylalanine biosynthesis. Studies on the mechanism of phenylalanine binding and feedback inhibition in the Escherichia coli P-protein. Biochemistry 38:12212–12217
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laborotary manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York pp17.2–17.44
Scrutton NS, Raine AR (1996) Cation-pi bonding and amino-aromatic interactions in the biomolecular recognition of substituted ammonium ligands. Biochem J 319:1-8
Shiio I, Sugimota S (1979) Two components of chorismate mutase in Brevibacterium flavum. J Biochem (Tokyo) 86: 17–25
Studier FW, Moffatt BA (1986) Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol 189:113–130
Sugimoto S, Shiio I (1974) Regulation of prephenate dehydratase in Brevibacterium flavum. J Biochem (Tokyo) 76:1103–1111
Ting AY, Shin I, Lucero C, Schultz G (1998) Energetic analysis of an engineered cation-π interaction in staphylococcal nuclease. J Am Chem Soc 120:7135–7136
Xia T, Zhao G, Jensen RA (1992) Loss of allosteric control but retention of the bifunctional catalytic competence of a fusion protein formed by excision of 260 base pairs from the 3′ terminus of pheA from Erwinia herbicola. Appl Environ Microbiol 58:2792–2798
Xue Y, Lipscomb WN (1995) Location of the active site of allosteric chorismate mutase from Saccharomyces cerevisiae, and comments on the catalytic and regulatory mechanisms. Proc Natl Acad Sci USA 92:10595–10598
Yanofsky C (1988) Transcription attenuation. J Biol Chem 263: 609–612
Yoshinga F, Nakamori S (1983) Production of amino acids. In: Hermann KM, Somerville KL (eds) Amino acid, biosynthesis and genetic regulation. Addison Wesley, New York, pp 310–355
Zamir LO, Tiberio R, Jensen RA (1983) Differential acid-catalyzed aromatization of prephenate, arogenate, and spiroarogenate. Tetrahedron Lett 24:2815-2818
Zhang S, Pohnert G, Kongsaeree P, Wilson DB, Clardy J, Ganem B (1998) Chorismate mutase-prephenate dehydratase from Escherichia coli. Study of catalytic and regulatory domains using genetically engineered proteins. J Biol Chem 273:6248–6253
Zhang S, Wilson DB, Ganem B (2000) Probing the catalytic mechanism of prephenate dehydratase by site-directed mutagenesis of the Escherichia coli P-protein dehydratase domain. Biochemistry 39:4722–4728
Acknowledgements
This work was supported by grants NSC88–2311-B-005-030 and NSC89–2316-B-005–018 from the National Science Council of the Republic of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hsu, SK., Lin, LL., Lo, HH. et al. Mutational analysis of feedback inhibition and catalytic sites of prephenate dehydratase from Corynebacterium glutamicum . Arch Microbiol 181, 237–244 (2004). https://doi.org/10.1007/s00203-004-0649-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-004-0649-5