Abstract
Summary
We performed a GWAS of trochanter and intertrochanter bone mineral density (BMD) in the Framingham Heart Study and replicated in three independent studies. Our results identified one novel locus around the associated variations at chromosomal region 3q13.32 and replicated two loci at chromosomal regions 3p21 and 8q24. Our findings provide useful insights that enhance our understanding of bone development, osteoporosis, and fracture pathogenesis.
Introduction
Hip trochanter (TRO) and intertrochanter (INT) subregions have important clinical relevance to subtrochanteric and intertrochanteric fractures but have rarely been studied by genome-wide association studies (GWASs).
Methods
Aiming to identify genomic loci associated with BMD variation at TRO and INT regions, we performed a GWAS utilizing the Framingham Heart Study (FHS, N = 6,912) as discovery sample and utilized the Women’s Health Initiative (WHI) African-American subsample (N = 845), WHI Hispanic subsample (N = 446), and Omaha osteoporosis study (N = 971), for replication.
Results
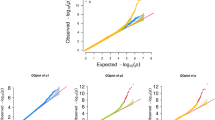
Combining the evidence from both the discovery and the replication samples, we identified one novel locus around the associated variations at chromosomal region 3q13.32 (rs1949542, discovery p = 6.16 × 10−8, replication p = 2.86 × 10−4 for INT-BMD; discovery p = 1.35 × 10−7, replication p = 4.16 × 10−4 for TRO-BMD, closest gene RP11-384F7.1). We also replicated two loci at chromosomal regions 3p21 (rs148725943, discovery p = 6.61 × 10−7, replication p = 5.22 × 10−4 for TRO-BMD, closest gene CTNNB1) and 8q24 (rs7839059, discovery p = 2.28 × 10−7, replication p = 1.55 × 10−3 for TRO-BMD, closest gene TNFRSF11B) that were reported previously. We demonstrated that the effects at both 3q13.32 and 3p21 were specific to the TRO, but not to the femoral neck and spine. In contrast, the effect at 8q24 was common to all the sites.
Conclusion
Our findings provide useful insights that enhance our understanding of bone development, osteoporosis, and fracture pathogenesis.



Similar content being viewed by others
References
Cummings SR, Melton LJ (2002) Epidemiology and outcomes of osteoporotic fractures. Lancet 359:1761–1767
Reginster JY, Burlet N (2006) Osteoporosis: a still increasing prevalence. Bone 38:S4–9
Peacock M, Turner CH, Econs MJ, Foroud T (2002) Genetics of osteoporosis. Endocr Rev 23:303–326
Richards JB, Rivadeneira F, Inouye M et al (2008) Bone mineral density, osteoporosis, and osteoporotic fractures: a genome-wide association study. Lancet 371:1505–1512
Styrkarsdottir U, Halldorsson BV, Gretarsdottir S et al (2008) Multiple genetic loci for bone mineral density and fractures. N Engl J Med 358:2355–2365
Duncan EL, Danoy P, Kemp JP et al (2011) Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLoS Genet 7:e1001372
Estrada K, Styrkarsdottir U, Evangelou E, et al. (2012) Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet 44(5):491–501
Guo Y, Tan LJ, Lei SF et al (2010) Genome-wide association study identifies ALDH7A1 as a novel susceptibility gene for osteoporosis. PLoS Genet 6:e1000806
Koller DL, Ichikawa S, Lai D et al (2010) Genome-wide association study of bone mineral density in premenopausal European-American women and replication in African-American women. J Clin Endocrinol Metab 95:1802–1809
Rivadeneira F, Styrkarsdottir U, Estrada K et al (2009) Twenty bone-mineral-density loci identified by large-scale meta-analysis of genome-wide association studies. Nat Genet 41:1199–1206
Xiong DH, Liu XG, Guo YF et al (2009) Genome-wide association and follow-up replication studies identified ADAMTS18 and TGFBR3 as bone mass candidate genes in different ethnic groups. Am J Hum Genet 84:388–398
Zhang L, Choi HJ, Estrada K et al (2014) Multistage genome-wide association meta-analyses identified two new loci for bone mineral density. Hum Mol Genet 23:1923–1933
Zheng HF, Forgetta V, Hsu YH et al (2015) Whole-genome sequencing identifies EN1 as a determinant of bone density and fracture. Nature 526:112–117
Davis TR, Sher JL, Horsman A, Simpson M, Porter BB, Checketts RG (1990) Intertrochanteric femoral fractures. Mechanical failure after internal fixation. J Bone Joint Surg Br 72:26–31
Parker MJ, Dutta BK, Sivaji C, Pryor GA (1997) Subtrochanteric fractures of the femur. Injury 28:91–95
Cupples LA, Arruda HT, Benjamin EJ et al (2007) The Framingham Heart Study 100K SNP genome-wide association study resource: overview of 17 phenotype working group reports. BMC Med Genet 8(Suppl 1):S1
The Women’s Health Initiative Study Group (1998) Design of the Women’s Health Initiative clinical trial and observational study. The Women’s Health Initiative Study Group. Control Clin Trials 19:61–109
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Zhang L, Li J, Pei YF, Liu Y, Deng HW (2009) Tests of association for quantitative traits in nuclear families using principal components to correct for population stratification. Ann Hum Genet 73:601–613
Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, Hurles ME, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467:1061–1073
Zhang L, Pei YF, Fu X, Lin Y, Wang YP, Deng HW (2014) FISH: fast and accurate diploid genotype imputation via segmental hidden Markov model. Bioinformatics 30:1876–1883
Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR (2010) MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 34:816–834
Willer CJ, Li Y, Abecasis GR (2010) METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26:2190–2191
Mumpower JL (1991) Risk, ambiguity, insurance, and the winner’s curse. Risk Anal 11:519–522
Li J, Ji L (2005) Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity 95:221–227
Ward LD, Kellis M (2012) HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 40:D930–934
Kundaje A, Meuleman W, Ernst J et al (2015) Integrative analysis of 111 reference human epigenomes. Nature 518:317–330
Consortium EP (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74
Lonsdale J, Thomas J, Salvatore M, Phillips R (2013) The Genotype-Tissue Expression (GTEx) project. Nat Genet 45:580–585
Westra HJ, Peters MJ, Esko T et al (2013) Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat Genet 45:1238–1243
Paternoster L, Lorentzon M, Lehtimaki T et al (2013) Genetic determinants of trabecular and cortical volumetric bone mineral densities and bone microstructure. PLoS Genet 9:e1003247
Suhre K, Shin SY, Petersen AK et al (2011) Human metabolic individuality in biomedical and pharmaceutical research. Nature 477:54–60
Cui W, Cuartas E, Ke J, Zhang Q, Einarsson HB, Sedgwick JD, Li J, Vignery A (2007) CD200 and its receptor, CD200R, modulate bone mass via the differentiation of osteoclasts. Proc Natl Acad Sci U S A 104:14436–14441
Sunaga N, Kohno T, Kolligs FT, Fearon ER, Saito R, Yokota J (2001) Constitutive activation of the Wnt signaling pathway by CTNNB1 (beta-catenin) mutations in a subset of human lung adenocarcinoma. Genes Chromosomes Cancer 30:316–321
Yu Y, Wu J, Wang Y, Zhao T, Ma B, Liu Y, Fang W, Zhu WG, Zhang H (2012) Kindlin 2 forms a transcriptional complex with beta-catenin and TCF4 to enhance Wnt signalling. EMBO Rep 13:750–758
Richards JB, Zheng HF, Spector TD (2012) Genetics of osteoporosis from genome-wide association studies: advances and challenges. Nat Rev Genet 13:576–588
Pei YF, Zhang L, Papasian CJ, Wang YP, Deng HW (2014) On individual genome-wide association studies and their meta-analysis. Hum Genet 133:265–279
Acknowledgments
This study was partially supported by the National Natural Science Foundation of China (31571291 and 31100902 to L.Z., 31501026 to Y.F.P., 81460223 to R.H.), the NIH (P50AR055081, R01AG026564, R01AR050496, RC2DE020756, R01AR057049, and R03TW008221 to H.W.D.), the Natural Science Foundation of Jiangsu Province of China (BK20150323 to Y.F.P.), the Franklin D. Dickson/Missouri Endowment and the Edward G. Schlieder Endowment (to H.W.D.), the Startup Funding Project of Soochow University (Q413900214 to L.Z. and Q413900114 to Y.F.P.), and a Project of The Priority Academic Program Development of Jiangsu Higher Education Institutions. Computing service was partially provided by the University of Shanghai for science and technology computing cluster.
The FHS is conducted and supported by the National Heart, Lung, and Blood Institute (NHLBI) in collaboration with Boston University (contract no. N01-HC-25195). This manuscript was not prepared in collaboration with investigators of the FHS and does not necessarily reflect the opinions or views of the FHS, Boston University, or NHLBI. Funding for SHARe Affymetrix genotyping was provided by NHLBI contract N02-HL-64278. SHARe Illumina genotyping was provided under an agreement between Illumina and Boston University. Funding support for the Framingham whole body and regional dual x-ray absorptiometry (DXA) dataset was provided by NIH grants R01 AR/AG 41398. The datasets used for the analyses described in this manuscript were obtained from dbGaP at http://www.ncbi.nlm.nih.gov/sites/entrez?db=gap through dbGaP accession phs000342.v14.p10.
The WHI program is funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, and the US Department of Health and Human Services through contracts N01WH22110, 24152, 32100-2, 32105-6, 32108-9, 32111-13, 32115, 32118-32119, 32122, 42107-26, 42129-32, and 44221. This manuscript was not prepared in collaboration with investigators of the WHI, has not been reviewed and/or approved by the Women’s Health Initiative (WHI), and does not necessarily reflect the opinions of the WHI investigators or the NHLBI. Funding for WHI SHARe genotyping was provided by NHLBI contract N02-HL-64278. The datasets used for the analyses described in this manuscript were obtained from dbGaP at http://www.ncbi.nlm.nih.gov/sites/entrez?db=gap through dbGaP accession phs000200.v10.p3.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
None
Additional information
Hong-Wen Deng and Lei Zhang jointly supervised this study.
Yu-Fang Pei and Zong-Gang Xie contributed equally to this work.
Rights and permissions
About this article
Cite this article
Pei, YF., Xie, ZG., Wang, XY. et al. Association of 3q13.32 variants with hip trochanter and intertrochanter bone mineral density identified by a genome-wide association study. Osteoporos Int 27, 3343–3354 (2016). https://doi.org/10.1007/s00198-016-3663-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00198-016-3663-y