Abstract
We present a texture analysis methodology that combined uncommitted machine-learning techniques and partial least square (PLS) in a fully automatic framework. Our approach introduces a robust PLS-based dimensionality reduction (DR) step to specifically address outliers and high-dimensional feature sets. The texture analysis framework was applied to diagnosis of knee osteoarthritis (OA). To classify between healthy subjects and OA patients, a generic bank of texture features was extracted from magnetic resonance images of tibial knee bone. The features were used as input to the DR algorithm, which first applied a PLS regression to rank the features and then defined the best number of features to retain in the model by an iterative learning phase. The outliers in the dataset, that could inflate the number of selected features, were eliminated by a pre-processing step. To cope with the limited number of samples, the data were evaluated using Monte Carlo cross validation (CV). The developed DR method demonstrated consistency in selecting a relatively homogeneous set of features across the CV iterations. Per each CV group, a median of 19 % of the original features was selected and considering all CV groups, the methods selected 36 % of the original features available. The diagnosis evaluation reached a generalization area-under-the-ROC curve of 0.92, which was higher than established cartilage-based markers known to relate to OA diagnosis.




Similar content being viewed by others
References
Abdi, H.: Partial least squares (pls) regression. Encycl. Soc. Sci. Res. Methods Ed. 21(4), 1–7 (2003)
Alsberg, B.K., Kell, D.B., Goodacre, R.: Variable selection in discriminant partial least-squares analysis. Analyt. Chem. 70(19), 4126–4133 (1998)
Barker, M., Rayens, W.: Partial least squares for discrimination. J. Chemom. 17(3), 166–173 (2003)
Beyer, K.S., Goldstein, J., Ramakrishnan, R., Shaft, U.: When is “nearest neighbor” meaningful? In: Proceedings of the 7th International Conference on Database Theory, ICDT ’99, pp. 217–235. Springer, London, UK (1999)
Branden, K.V., Hubert, M.: Robustness properties of a robust partial least squares regression method. Analytica Chimica Acta 515(1), 229–241 (2004)
Bryan, K., Brennan, L., Cunningham, P.: Metafind: A feature analysis tool for metabolomics data. BMC Bioinform. 9(1), 470 (2008)
Burman, P.: A comparative study of ordinary cross-validation, v-fold cross-validation and the repeated learning-testing methods. Biometrika 76(3), 503–514 (1989)
Chong, I.G., Jun, C.H.: Performance of some variable selection methods when multicollinearity is present. Chemom. Intell Lab. Syst. 78(1–2), 103–112 (2005)
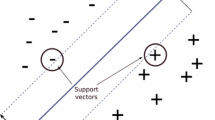
Cortes, C., Vapnik, V.: Support-vector networks. Mach. Learn. 20(3), 273–297 (1995)
Dam, E.B., Loog, M., Christiansen, C., Byrjalsen, I., Folkesson, J., Nielsen, M., Qazi, A., Pettersen, P.C., Garnero, P., Karsdal, M.A.: Identification of progressors in osteoarthritis by combining biochemical and MRI-based markers. Arthr. Res. Ther. 11(4), R115 (2009)
Daszykowski, M., Heyden, Y.V., Walczak, B.: Robust partial least squares model for prediction of green tea antioxidant capacity from chromatograms. J. Chromatogr. A 1176(1–2), 12–18 (2007)
Daszykowski, M., Kaczmarek, K., Heyden, Y.V., Walczak, B.: Robust statistics in data analysis—a review: basic concepts. Chemom. Intell. Lab. Syst. 85(2), 203–219 (2007)
Florack, L.M.J., Haar Romeny, BMt, Koenderink, J.J., Viergever, M.A.: The Gaussian scale-space paradigm and the multiscale local jet. Int. J. Comput. Vis. 18(1), 61–75 (1996)
Folkesson, J., Dam, E.B., Olsen, O.F., Pettersen, P.C., Christiansen, C.: Segmenting articular cartilage automatically using a voxel classification approach. IEEE Trans. Med. Imaging 26, 106–115 (2007)
Furlanello, C., Serafini, M., Merler, S., Jurman, G.: Entropy-based gene ranking without selection bias for the predictive classification of microarray data. BMC Bioinform. 4(1), 54 (2003)
Guyon, I., Weston, J., Barnhill, S., Vapnik, V.: Gene selection for cancer classification using support vector machines. Mach. Learn. 46(1-3), 389–422 (2002)
Herlidou, S., Rolland, Y., Bansard, J.Y., Le Rumeur, E., Certaines, J.D.: Comparison of automated and visual texture analysis in mri: Characterization of normal and diseased skeletal muscle. Magn. Resonan. Imaging 17(9), 1393–1397 (1999)
Herlidou, S., Grebe, R., Meyer, M.E.: Influence of age and osteoporosis on calcaneus trabecular bone structure: a preliminary in vivo mri study by quantitative texture analysis. Magn. Resonan. Imaging 22(2), 237–243 (2004)
Hoskuldsson, A.: Analysis of latent structures in linear models. J. Chemom. 17(1), 630–645 (2003)
Igel, C., Heidrich-Meisner, V., Glasmachers, T.: Shark. J. Mach. Learn. Res. 9, 993–996 (2008)
de Jong, S.: Simpls: an alternative approach to partial least squares regression. Chemom. Intell. Lab. Syst. 18(3), 251–263 (1993)
Kellgren, J.H., Lawrence, J.S.: Radiological assessment of osteo-arthrosis. Annals Rheum. Dis. 16(4), 494–502 (1957)
Kemsley, E.K.: Discriminant analysis of high-dimensional data: a comparison of principal components analysis and partial least squares data reduction methods. Chemom. Intell. Lab. Syst. 33(1), 47–61 (1996)
Kovalev, V.A., Kruggel, F., von Cramon, D.: Gender and age effects in structural brain asymmetry as measured by mri texture analysis. NeuroImage 19(3), 895–905 (2003)
Kruger, U., Zhou, Y., Wang, X., Rooney, D., Thompson, J.: Robust partial least squares regression: part i, algorithmic developments. J. Chemom. 22(1), 1–13 (2008)
Krzanowski, W.J. (ed.): Principles of Multivariate Analysis: A User’s Perspective. Oxford University Press Inc., New York (1988)
Eriksson, L., Johansson, E., Kettaneh-Wold, N., Trygg, J., Wikstrom, C., Wold, S: Multi- and Megavariate Data Analysis: Basic Principles and Applications. vb. 1. Umetrics AB (2006)
Li, H., Liang, Y., Xu, Q., Cao, D.: Key wavelengths screening using competitive adaptive reweighted sampling method for multivariate calibration. Analytica Chimica Acta 648(1), 77–84 (2009)
Liitiainen, E., Corona, F., Lendasse, A.: On the curse of dimensionality in supervised learning of smooth regression functions. Neural Process. Lett. 34, 133–154 (2011)
Lindgren, F., Geladi, P., Rannar S., et al.: Interactive variable selection (ivs) for pls. part 1: theory and algorithms. J. Chemom. 8, 349–363 (1994)
Liu, H., Motoda, H.: Feature Extraction, Construction and Selection: A Data Mining Perspective. Kluwer Academic Publishers, Norwell (1998)
Mehmood, T., Martens, H., Saebo, S., Warringer, J., Snipen, L.: A partial least squares based algorithm for parsimonious variable selection. Algorithms Mol. Biol. 6(1), 27 (2011)
Ramirez, J., Gorriz, J., Segovia, F., Chaves, R., Salas-Gonzalez, D., Lopez, M., Alvarez, I., Padilla, P.: Computer aided diagnosis system for the alzheimer’s disease based on partial least squares and random forest spect image classification. Neurosci. Lett. 472(2), 99–103 (2010)
Rodgers, J.L., Nicewander, W.A.: Thirteen ways to look at the correlation coefficient. Am. Stat. 42(1), 59 (1988)
Schad, L.R., Blml S, Z.I.: Mr tissue characterization of intracranial tumors by means of texture analysis. Magnetic Resonan. Imaging 11(6), 889–896 (1993)
Sørensen, L., Shaker, S.B., de Bruijne, M.: Quantitative analysis of pulmonary emphysema using local binary patterns. IEEE Trans. Med. Imaging 29(2), 559–569 (2010)
Tuceryan, M., Jain, A.K.: Texture analysis. In: The Handbook of Pattern Recognition and Computer Vision, 2nd edn., pp. 207–248. World Scientific Publishing Co., Singapore (1998)
Wang, X., Tang, X.: Experimental study on multiple lda classifier combination for high dimensional data classification
Weickert, J.: Anisotropic Diffusion in Image Processing. B.G. Teubner, Stuttgart (1998)
Wold, S., Sjostrom, M., Eriksson, L.: Pls-regression: a basic tool of chemometrics. Chemom. Intell. Lab. Syst. 58(2), 109–130 (2001)
Wold, S., Trygg, J.: The pls method—partial least squares projections to latent structures—and its applications in industrial rdp (research, development, and production). In: PLS in Industrial RPD for Prague, vol. 1, pp. 1–44 (2004)
Acknowledgments
We thankfully acknowledge the funding from the Danish Research Foundation (Den Danske Forskningsfond) supporting this work and the Center for Clinical and Basic Research for providing scans and radiographic readings. Christian Igel gratefully acknowledges support from the Danish National Advanced Technology Foundation through project “Personalized breast cancer screening”.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marques, J., Igel, C., Lillholm, M. et al. Linear feature selection in texture analysis - A PLS based method. Machine Vision and Applications 24, 1435–1444 (2013). https://doi.org/10.1007/s00138-012-0461-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00138-012-0461-1