Abstract
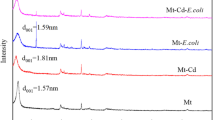
A heavy metal resistant strain, Pseudomonas stutzeri (MTCC 101) has been investigated for its cadmium tolerance properties along with its antibiotic resistance. The organism could tolerate cadmium up to 1,200 μg/mL with LD50 value 700 μg/mL. The gene(s) involved in such high resistance appear(s) to be induced in the presence of the metal. Increasing concentrations of cadmium successively prolonged the lag phase of growth with delayed attainment of the stationary phase. Transmission electron microscope and scanning electron microscope–energy dispersive analysis of X-ray spectroscope analysis showed cadmium adsorption on the bacterial surface with morphological distortion. Atomic absorption spectrometric study corroborated this data, showing highest cadmium accumulation in the cell wall fraction of the bacteria. Additionally, the cell wall fraction showed synthesis of new proteins when grown under metal stress.







Similar content being viewed by others
References
Abdelatey LM, Khalil WKB, Ali TH, Mahrous KF (2011) Heavy metal resistance and gene expression analysis of metal resistance genes in gram positive and gram negative bacteria present in Egyptian soil. J Appl Sci Environ Sanit 6:201–211
Antony R, Sujith PP, Fernandes SO, Verma P, Khedekar VD, Bharathi PAL (2011) Cobalt immobilization by manganese oxidizing bacteria from the Indian ridge system. Curr Microbiol 62:840–849
Austin CB, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14:176–182
Basu M, Paul AK (2008) Influence of environmental factors on the uptake of chromium by Pseudomonas stutzeri TEM-317 isolated from tannery sludge. J Mycopathol Res 46:289–295
Chovanova K, Sladekova D, Kmet V, Proksova M, Harichova J, Puskarova A, Polek B, Ferianc P (2004) Identification and characterization of eight cadmium resistant bacterial isolates from a cadmium-contaminated sewage sludge. Biologia, Bratislava 59:817–827
CLSI (2010) Performance standards for antimicrobial susceptibility testing; twentieth informational supplement, Ed. 20, vol 30, no. 1
Goulhen F, Gloter A, Guyot F, Bruschi M (2006) Cr(VI) detoxification by Desulfovibrio vulgaris strain hildenborough: microbe–metal interactions studies. Appl Microbiol Biotechnol 71:892–897
Haritha A, Sagar KP, Tiwari A, Kiranmayi P, Rodrigue A, Mohan PM, Singh SS (2009) MrdH, a novel metal resistance determinant of Pseudomonas putida KT2440, is flanked by metal-inducible mobile genetic elements. J Bacteriol 191:5976–5987
Higham DP, Sadler PJ, Scawen MD (1985) Cadmium resistance in Pseudomonas putida: growth and uptake of cadmium. J Gen Microbiol 131:2539–2544
Lu LT, Chang IC, Hsiao TY, Yu YH, Ma HW (2007) Identification of pollution source of cadmium in soil: application of material flow analysis and a case study in Taiwan. Environ Sci Pollut Res Int 14:49–59
Maagd RA, Lugtenberg B (1986) Fractionation of Rhizobium leguminosarum cells into outer membrane, cytoplasmic membrane, periplasmic, and cytoplasmic components. J Bacteriol 167:1083–1085
Massadeh AM, Al-Momani FA, Haddad HI (2005) Removal of lead and cadmium by halophilic bacteria isolated from the Dead Sea shore, Jordan. Biol Trace Elem Res 108:259–269
Mejare M, Bulow L (2001) Metal-binding proteins and peptides in bioremediation and phytoremediation of heavy metals. Trends Biotechnol 19:67–73
Nandakumar MP, Shen J, Raman B, Marten MR (2003) Solubilization of trichloroacetic acid (TCA) precipitated microbial proteins via NaOH for two-dimensional electrophoresis. J Proteome Res 2:89–93
Parungao MM, Tacata PS, Tanayan CRG, Trinidad LC (2007) Biosorption of copper, cadmium and lead by copper-resistant bacteria isolated from Mogpog River, Marinduque. Philipp J Sci 136:155–165
Ramos L, Fernández MA, González MJ, Hernández LM (1999) Heavy metal pollution in water, sediments, and earthworms from the Ebro River, Spain. Bull Environ Contam Toxicol 63:305–311
Satarug S, Baker JR, Urbenjapol S, Haswell-Elkins M, Reilly PEB, Williams DJ, Moore MR (2003) A global perspective on cadmium pollution and toxicity in non-occupationally exposed population. Toxicol Lett 137:65–83
Silver S, Phung T (2005) A bacterial view of the periodic table: genes and proteins for toxic inorganic ions. J Ind Microbiol Biotechnol 32:587–605
Singh V, Chauhan PK, Kanta R, Dhewa T, Kumar V (2010) Isolation and characterization of Pseudomonas resistant to heavy metal contaminants. Int J Pharm Sci Rev Res 3:164–167
Teitzel GM, Parsek MR (2003) Heavy metal resistance of biofilm and planktonic Pseudomonas aeruginosa. Appl Environ Microbiol 69:2313–2320
Watcharamusika A, Prapagdeea B, Thavipokeb P, Boontanonb N (2008) The role of exopolymers in protection of Ralstonia sp., a cadmium-resistant bacterium, from cadmium toxicity. Environ Asia 2:37–42
Acknowledgments
Financial support from Barrackpore Rastraguru Surendranath College, West Bengal, India, Grant # 157-BT(Estt.)/RD-11/10, Department of Biotechnology (DBT), West Bengal, India and technical support from DR. B. Mondal of Bidhan Chandra Krishi Vidyalaya, West Bengal, India, DR. W. Ghosh of Bose Institute, West Bengal, India and P. Chowdhury of Barrackpore Rastraguru Surendranath College have been greatly acknowledged. The authors are also thankful to Dr. M.K. Ghosh, CSIR-IICB, Kolkata and Prof. A.K. Paul of University of Calcutta for his constant support and guidance during the compilation of the work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Deb, S., Ahmed, S.F. & Basu, M. Metal Accumulation in Cell Wall: A Possible Mechanism of Cadmium Resistance by Pseudomonas stutzeri . Bull Environ Contam Toxicol 90, 323–328 (2013). https://doi.org/10.1007/s00128-012-0933-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00128-012-0933-z