Abstract
Aims/hypothesis
While autoantibodies are traditional markers for type 1 diabetes development, we identified gut microbial biomarkers in 1-year-old infants associated with future type 1 diabetes up to 20 years before diagnosis.
Methods
Infants enrolled in the longitudinal general population cohort All Babies In Southeast Sweden (ABIS) provided a stool sample at a mean age of 12.5 months. Samples (future type 1 diabetes, n=16; healthy controls, n=268) were subjected to 16S ribosomal RNA (rRNA) sequencing and quantitative PCR. Microbial differences at the taxonomic and core microbiome levels were assessed. PICRUSt was used to predict functional content from the 16S rRNA amplicons. Sixteen infants, with a future diagnosis of type 1 diabetes at a mean age of 13.3±5.4 years, and one hundred iterations of 32 matched control infants, who remained healthy up to 20 years of age, were analysed.
Results
Parasutterella and Eubacterium were more abundant in healthy control infants, while Porphyromonas was differentially more abundant in infants with future type 1 diabetes diagnosis. Ruminococcus was a strong determinant in differentiating both control infants and those with future type 1 diabetes using random forest analysis and had differing trends of abundance when comparing control infants and those with future type 1 diabetes. Flavonifractor and UBA1819 were the strongest factors for differentiating control infants, showing higher abundance in control infants compared with those with future type 1 diabetes. Alternatively, Alistipes (more abundant in control infants) and Fusicatenibacter (mixed abundance patterns when comparing case and control infants) were the strongest factors for differentiating future type 1 diabetes. Predicted gene content regarding butyrate production and pyruvate fermentation was differentially observed to be higher in healthy control infants.
Conclusions/interpretation
This investigation suggests that microbial biomarkers for type 1 diabetes may be present as early as 1 year of age, as reflected in the taxonomic and functional differences of the microbial communities. The possibility of preventing disease onset by altering or promoting a ‘healthy’ gut microbiome is appealing.
Data availability
The forward and reverse 16S raw sequencing data generated in this study are available through the NCBI Sequence Read Archive under BioProject PRJNA875929. Associated sample metadata used for statistical comparison are available in the source data file. R codes used for statistical comparisons and figure generation are available at: https://github.com/PMilletich/T1D_Pipeline.
Graphical abstract

Similar content being viewed by others

Introduction
Type 1 diabetes, a heterogeneous, serious autoimmune disease associated with high morbidity and a shorter life expectancy [1], is characterised by the destruction of the insulin-producing beta cells of the pancreas. The aetiology of type 1 diabetes is complex, with suggested genetic and environmental factors [2]. Approximately 50% of the genetic risk of type 1 diabetes is conferred by the MHC class II HLA alleles [3], where different combinations of alleles result in varying risk. Most individuals with type 1 diabetes have no family history of the disease. Low genetic risk does not preclude the development of clinical type 1 diabetes. The incidence of type 1 diabetes varies during childhood, with a peak during puberty and earlier age at diagnosis and an increased incidence in populations with a lower genetic risk of type 1 diabetes [4,5,6]. Early childhood diet [7, 8], duration of breastfeeding [9], and antibiotic use early in life influence the gut microbiome [10] and have been proposed in the pathogenesis of type 1 diabetes.
The gut microbiome plays a critical role in differentiating immune cells by mediating proinflammatory Th17 cells and balancing Th1 and Th2 populations [11,12,13]. An altered gut microbiome may facilitate inflammation by decreased differentiation of CD8+ regulatory T cells, triggering an autoimmune response [13]. Dysbiosis early in life may result in abnormal immunoregulation [14, 15], predisposing a child to autoimmunity many years before the development of clinical disease [16].
Associations between the gut microbiome and autoimmunity have previously been described for type 1 diabetes [17,18,19,20]. Longitudinal studies have found a marked reduction in α diversity between seroconversion and type 1 diabetes diagnosis [21]. Additionally, matched case–control studies have found significant differences in gut microbiome composition in the first years of life [22, 23]. However, there are discrepancies in what constitutes core microbiome or dysbiosis related to type 1 diabetes.
Emerging evidence that gut microbiota contribute to the pathophysiology of type 1 diabetes suggests that gut dysbiosis is a central catalyst whereby bacterial metabolites disrupt the intestinal barrier function (i.e. give rise to ‘leaky gut’). In this environment, antigens then become dislocated into the circulation, activating the immune system and systemic inflammation [24]. This may initiate a cascade of autoimmune processes and beta cell damage through molecular mimicry from external antigens. Previous studies have found associations between increased Bacteroides-dominating communities and a decrease in butyrate-producing bacteria of the phylum Bacillota (previously Firmicutes), with early onset of autoantibody development [25, 26]. It is unknown whether these communities develop independently in these cohorts, although an imbalance in the ratio of Bacteroidetes to Firmicutes in the gut microbiome has been proposed in other pathologies such as obesity and inflammatory bowel disease [27].
Attempts to investigate environmental risk factors in large cohort studies have mainly focused on individuals with an increased genetic risk based on HLA alleles or a family history of type 1 diabetes [19, 28]. Generally, a reduced abundance of short-chain fatty acid (SCFA)-producing microbiota in children progressing to autoimmunity is implicated but no consistent differences in specific taxa across different high-risk populations and geographical regions have emerged. Studies restricted to those at genetic risk or with familial type 1 diabetes are limited in their potential to evaluate other factors that can protect against or trigger an autoimmune response in the general population. For example, it has been shown that the genetic risk HLA alleles of type 1 diabetes are associated with distinct changes in the human gut microbiome composition [29].
To further elucidate the role of the early microbiome in the pathology of type 1 diabetes, this study evaluates differences in the gut microbiome of infants in a general population, characterising signatures that diverge between those with a diagnosis of type 1 diabetes up to 20 years later and matched healthy control infants without this diagnosis.
Methods
Study design and sample collection
This study is based on the longitudinal, general population cohort All Babies In Southeast Sweden (ABIS). Families of children born in the Swedish counties of Östergötland, Småland, Blekinge and Öland between 1 October 1997 and 1 October 1999 were invited to join the study. Of 21,700 children born, 17,055 participated at birth (78.6%) and 13,886 (64.0%) at 1 year of age. Parents completed questionnaires at the time of birth and when the infant reached 1 year of age, and filled in a diary during the first year of life. The collected material includes information about pregnancy, factors during the first year of life such as nutrition, and information on parental lifestyle such as smoking and alcohol use.
Stool samples were collected from infants’ diapers at approximately 1 year by sterile spatula and tube and immediately frozen. Samples collected at home were transported frozen to the WellBaby Clinic. The samples were then stored dry in a −80°C freezer in Linköping and later transported frozen to the University of Florida. The viability of samples was confirmed previously by the ability to isolate and culture non-spore-forming, facultative anaerobic strains of Bifidobacterium [29].
Ethical considerations
The ABIS project has been approved by the Research Ethics Committees of the Faculty of Health Science at the University of Linköping, Linköping, Sweden (ref 1997/96287 and 2003/03-092) and the Medical Faculty at the University of Lund, Lund, Sweden (Dnr 99227 and Dnr 99321). Participating families gave informed consent after oral and written information and the opportunity to watch a video of the study. The microbiome analysis performed at the University of Florida has been approved by the University of Florida’s Institutional Review Board as an exempt study (IRB201800903).
HLA genotyping
Sequence-specific hybridisation with lanthanide-labelled oligonucleotide probes was used to determine HLA genotype by typing for HLA-DQB1 and informative DQA1 and DRB1 alleles. HLA-DR/DQ genotypes associated with risk and protection were defined according to the presence of common European HLA-DR-DQ haplotypes associated with the risk of autoimmunity. Infants were categorised into four risk groups according to HLA genotype [30]. Risk-associated HLA haplotype alleles were defined as (DR3)-DQA1*05-DQB1*02 and (DR4)-DQA1*03-DQB1*02 (DR3/4), while protective HLA haplotypes were defined as (DR15)-DQB1*0602, (DR13)-DQB1*0603, (DR5)-DQA1*05-DQB1*0301 and (DR7)-DQA1*0201-DQB1-0603.
A high genetic risk is defined by the presence of two increased risk-associated haplotypes.
Increased risk is defined by the presence of one risk-associated allele and a neutral haplotype.
Neutral risk is defined by either of the risk-associated haplotypes in combination with one of the protective haplotypes or two neutral haplotypes.
Low genetic risk is defined by the presence of one or two protective haplotypes.
Diagnosis
The Swedish National Patient Register [31] and the Swedish pediatric diabetes quality register SWEDIABKIDS [32] have been used to identify children with type 1 diabetes, with new incidences reported annually. A prescription of insulin in the Swedish National Drug Prescription Register [33] validated the diagnosis. As of the latest update in December 2020, 167 of the initial 17,055 participants have developed type 1 diabetes (0.98%). A diagnosis of other autoimmune diseases or neurodevelopmental disorders was confirmed by the Swedish National Diagnosis Registry.
Microbiome analysis
DNA extraction, 16S ribosomal RNA (rRNA) barcoded PCR and V3-V4 16S sequencing using Illumina Miseq 2×300 bp were performed as described previously [29]. Bacterial quantification through universal 16S rRNA primers was performed [34] and reads were merged and filtered [35]. In summary, forward and reverse fastq reads were merged, ambiguous primers were removed, and sequences were filtered and classified using SILVA version 138 before processing in R (version 4.2.2) [36, 37]. Nomenclature was established by sorting amplicon sequence variants (ASVs) by known genus and appending unique sequential numbers to the genus (and species, as available).
For this investigation, 1598 infants possessed more than 1000 16S rRNA copies/g of stool (74,018±33,485). Of these, 177 infants with other autoimmune or neurodevelopmental diagnoses were excluded. In the remaining 1421 infants, 16 received a future type 1 diabetes diagnosis. ASVs with fewer than five reads in five or more infants, as well as those ASVs lacking a known genus using SILVA classification, were removed. The remaining 1669 ASVs were conglomerated into 199 genera using the tax_glom function [38].
Random permutations for matched case–control iterations
Factors differentiating infants with a future type 1 diabetes diagnosis were determined with χ2 tests and p values computed by Monte Carlo simulation. Factors impacting binomial β diversity were tested using the permutational multivariate analysis of variance (PERMANOVA) test with 999 permutations [39]. The p values were corrected for false discovery rate (FDR) using the Benjamini–Hochberg method.
A subset of control infants (n=268) was identified by matching to case infants on geographical region, presence of siblings at birth, residence type, duration of total breastfeeding and month of stool collection (ANOVA: p≤0.001, padj≤0.01). On average, there were 14.5±19.09 control infants per case infant. Case infants with missing data from the diary were matched using the remaining variables. The month of stool collection was matched with a range of ±1 month for each case.
From the resulting sample of 268 healthy control infants, 100 iterations of 32 randomly selected control cohorts were generated. This iterative process allowed accounting for the inherent variability of control infants within a general population cohort without the risk of overpowering the results. The imbalance of control infants was mitigated while still balancing for the most significant influences of gut microbial diversity.
Statistical analysis
Pairwise comparisons of α diversity between infants with future type 1 diabetes (n=16) and control infants without future type 1 diabetes (n=268) were performed against both genera and ASVs using the default parameter s of the R functions plot_richness, ggplot2, and stat_compare_means [40, 41].
Prevalence filtering of the core microbiome
The Prevalence Interval for Microbiome Evaluation (PIME) R package was employed on each of the 100 case–control group comparisons using either ASVs or genera to obtain the core taxa representing the future type 1 diabetes or control groups [42].
The first ten iterations of the total abundance of case–control cohorts were used as a model for the remaining iterations. The initial out-of-bag error (OOB), without prevalence filtering, for genera ranged from 31.3% to 41.2%, with a mean of 36.4% (data not shown). A prevalence threshold of 70% was selected as it reduced the OOB to a mean of 2.5% while retaining an average of 33 of the original 199 genera (16.6%) (electronic supplementary material [ESM] Table 1). At the ASV level, the initial OOB, without prevalence filtering, ranged from 31.3% to 41.2%, averaging 35% (data not shown). A prevalence threshold of 50% was selected as it reduced the OOB to a mean of 4.0% while retaining an average of 61 of the original 1669 ASVs (3.7%) (ESM Table 2). After filtering for prevalence, binomial distances for the tenth iteration were visualised using Principal Coordinate Analysis (PCoA) [41].
Taxa found to be in the core microbiome in at least half of the iterations and with a positive mean decrease accuracy (MDA) in both the case and control groups were further assessed. The average MDA was determined through PIME and the difference in mean abundance between the case and the control subset was calculated for each iteration at each taxon.
Differential abundance analysis
Differential abundances of microbes, present in either 10% (genus-level analysis) or 20% (ASV-level analysis) of future type 1 diabetes or iterative control cohorts, were assessed with the binomial distribution model R package, DESeq2 [43]. The estimateSizeFactor() function was first used with the ‘poscount’ type, allowing for zeroes. A local fit type for the Wald test was used, without Cook’s distance filtering. After each of the 100 iterations of DESeq2, significance values were adjusted through default Benjamini–Hochberg method. Taxa were deemed significant to this investigation if they appeared in at least half of the iterations of case–control matching with adjusted p value (padj) <0.05. The distribution of Log2FoldChange was depicted using ggplot2 [40].
Results
Description of cohort
As of December 2020, 167 children in the ABIS cohort have developed type 1 diabetes (Fig. 1). Of these, stool samples at 1 year of age were available for 16 children; five had a high HLA risk for type 1 diabetes, five had an increased risk, four had a neutral risk and two had a decreased risk (see Genetic risk definition text box). There was a slight overrepresentation of boys to girls (10 vs 6), representative of the whole ABIS cohort (91 boys vs 76 girls with type 1 diabetes diagnosis, χ2 p=0.49). The mean age at diagnosis was 13 years, with a median of 14 years. The youngest age at type 1 diabetes diagnosis was 1 year and 4 months, and the oldest was 21 years and 4 months (Table 1).

Factors associated with future type 1 diabetes or microbiome composition
The HLA haplotype DR4-DQ8 was more prevalent in infants with future type 1 diabetes (62.5% vs 25.9%, p=0.003, padj=0.04) (ESM Table 3). Infants with future type 1 diabetes were 4.4 times more likely to have DR4-DQ8, and 2.58 times more likely to have DR3-DQ2.5 (p=0.007 and p=0.07, respectively) (Table 2). In contrast, the protective allele DR15-DQ602 was not significantly different when comparing the groups (p=0.15).
The most significant confounders of binomial β diversity included geographical region, presence of siblings at birth, residence type, duration of total breastfeeding and age, in months, at stool collection (ANOVA: p≤0.001 and padj≤0.01) (ESM Table 4). These factors were used to select control infants (n=268) and remove outliers. Additional confounders identified included the infant’s biological sex, both parents living abroad during infancy, mode of delivery, maternal risk factors during pregnancy and dietary factors during infancy (ANOVA: p≤0.05 and padj≤0.05).
Genera microbiome signatures
Despite the absence of a difference in α diversity of genera between control infants and those with a future diagnosis of type 1 diabetes (observed, p=0.82; Shannon, p=0.25) (ESM Fig. 1a), distinct clustering was observed after supervised learning through PIME (ESM Fig. 2). The core microbiome was estimated by filtering genera to a 70% prevalence threshold in either iterative controls or cases.
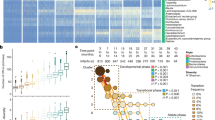
Seventeen core genera demonstrated a positive MDA for differentiating the future type 1 diabetes and iterative control cohorts. Ruminococcus was a key factor for differentiating both case and control infants, Flavonifractor and UBA1819 were the strongest factors for differentiating control infants, and Alistipes and Fusicatenibacter were the strongest factors for differentiating infants with future type 1 diabetes (Fig. 2a,b).
Significant differences in the core genera between future type 1 diabetes (n=16) and iterative control cohorts (n=32). (a, b) MDA from iterations of PIME at 70% prevalence thresholding through total (a) and relative (b) abundance. (c, d) Respective genera identified through PIME with distribution of differences in average total (c) and relative (d) abundance across all 100 iterations. Boxplots show the median and IQR for each group, with circles representing outliers for each respective group. (e, f) Differentially abundant genera (DESeq2 padj<0.05) with minimum 10% prevalence in either control or case infants using (e) total or (f) relative abundance. Positive values are more abundant in future type 1 diabetes, negative values are more abundant in control iterations
Bacteroides, Enterococcus, Gemella, Hungatella and TM7x had higher total and relative abundance in infants with a future diagnosis of type 1 diabetes. Alistipes, Anaerostipes, Eggerthella, Flavonifractor and Ruminococcaceae UBA1819 had higher total and relative abundance in control infants (Fig. 2c,d). Agathobacter, Blautia and Fusicatenibacter had a higher total abundance in control infants but a higher relative abundance in infants with a future type 1 diabetes diagnosis (ESM Fig. 3a). Alternatively, Romboutsia, Roseburia and Ruminococcus had higher total abundance in infants with a future type 1 diabetes diagnosis but higher relative abundance in control infants (ESM Fig. 3a).
Outside of the core microbiome analysis, differentially abundant bacteria were identified using DESeq2 with a 10% prevalence threshold applied (padj<0.05). Porphyromonas was higher in both total and relative abundance in infants with future type 1 diabetes, while Eubacterium and Parasutterella had a higher total and relative abundance in control infants (Fig. 2e,f). Prevalence differences of key genera are shown in ESM Fig. 4a, ASVs composing key genera are shown in ESM Table 5.
ASV microbiome signatures
The case and control groups demonstrated similar α diversity within ASVs (observed, p=0.56; Shannon, p=0.51) (ESM Fig. 1b). Yet, distinct clustering was observed in the core microbiome after filtering to a 50% prevalence threshold (ESM Fig. 2h).
Ten core ASVs were most significant in differentiating between cases and controls. Agathobacter-434 had the highest MDA score for both groups, Anaerostipes-747 had the next highest MDA for control infants, while Lachnospira-5640 had the next highest for infants with a future type 1 diabetes diagnosis (Fig. 3a,b). Agathobacter-434 had a higher total and relative abundance in infants with future type 1 diabetes. Anaerostipes-747, Eggerthella lenta-3665, Faecalibacterium praustnitzii-4451 and Veillonella atypica-10087 had higher total and relative abundance in control infants (Fig. 3c,d). Agathobacter-387 and Lachnospira-5640 had higher total abundance in control infants but higher relative abundance in infants with future type 1 diabetes. Additionally, Anaerostipes hadrus-719, Veillonella-10427 and Veillonella atypica-10084 had a higher total abundance in the future type 1 diabetes group but a higher relative abundance in the control group (ESM Fig. 3b).
Core ASV differences between future type 1 diabetes (n=16) and iterative control cohorts (n=32). (a, b) MDA from iterations of PIME at 50% prevalence thresholding through total (a) and relative (b) abundance. (c, d) Respective ASVs identified through PIME with the distribution of differences in average total (c) and relative (d) abundance across all 100 iterations. Boxplots show the median and IQR for each group, with circles representing outliers for each respective group. (e, f) Differentially abundant genera (DESeq2 padj<0.05) with minimum 10% prevalence in either iterative control or case infants using (e) total or (f) relative abundance. Positive values are more abundant in future type 1 diabetes, negative values are more abundant in control iterations
Differentially abundant ASVs were identified after applying a 20% prevalence filter in either the case or iterative control cohort (DESeq2 padj<0.05) (Fig. 3e,f). Two ASVs, Clostridium sensu stricto 1 butyricum-2752 and Terrisporobacter-9845, had higher total and relative abundance in infants with future type 1 diabetes. Eisenbergiella massiliensis-3699, Enterococcus-3895, Enterococcus-4905, Erysipelatoclostridium-3992 and Veillonella atypica-10085 had higher total and relative abundance in control infants.
ASV 16S rRNA sequences and BLAST [44] classifications can be found in ESM Table 6, and prevalence differences of key ASVs are shown in ESM Fig. 4b.
Functional differences
In addition to taxonomical differences in the gut microbiome, functional differences were also found (Picrust2: Wilcox p≤0.05, padj>0.1) (Fig. 4) based on predictions from the 16S rRNA data. Control infants (n=268) possessed higher predicted expression of acetyl-CoA fermentation to butanoate II (Cohen’s d=0.511), pyruvate fermentation to acetone (Cohen’s d=0.417), cob(II)yrinate a,c-diamide biosynthesis I (early cobalt insertion) (Cohen’s d=0.447) and nitrate reduction VI (assimilatory) (Cohen’s d=0.552).
Functional pathway differences between control infants (n=268) and those with future type 1 diabetes (n=16) inferred by 16S amplicons through PICRUSt. Pathway expression was transformed to relative abundance for each infant, checked for normality, and then assessed for significance using either t test or Wilcoxon test. p values adjusted through FDR were non-significant (NS). Boxplots show the median and IQR for each group, with circles representing outliers for each respective group
Discussion
This general population cohort study identified several potential early bacterial biomarkers for the future onset of type 1 diabetes. One hundred iterations of 32 control infants were compared with 16 infants diagnosed with future type 1 diabetes at a mean age of 13.3±5.4 years. Despite the considerable period between stool collection and diagnosis of diabetes, compositional and functional differences in gut microbiota were observed when comparing healthy infants and those with future type 1 diabetes.
To avoid progression to symptomatic type 1 diabetes, tools to predict future diabetes before or during the first stage of the disease must be developed. Beta cell autoantibodies are rarely detected before the age of 6 months, while the peak incidence of IAA is between 9 and 24 months and that for GADA is 36 months. Gut microbial biomarkers at 12 months would benefit the prediction opportunity well before the onset of multiple autoantibodies. Although gut diversity did not significantly differ between case and control infants, perhaps due to the transient nature of this period of gut microbiome development, taxonomical differences were observed.
Standard methods for microbial community analysis typically focus exclusively on differences in relative abundance of the microbes. The microbial load on the gastrointestinal tract can often not be accurately shown through relative abundance alone. This prompted us to assess both total and relative abundance, should both copy number and relative composition of the gut play a role for the bacterium in question. While not statistically significant, we observed an increase in bacterial load in infants with a future diagnosis of type 1 diabetes (mean copies of 16S per gram of stool: 6.59×105–5.68×108 in infants with future type 1 diabetes; 2.7×102–9.01×108 in control infants), possibly explaining some of the differences between total and relative abundance patterns. This warrants further investigation into how these bacterial loads impact the gut microbiome.
Core genera more abundant in control infants were primarily Firmicutes, such as Anaerostipes, Flavonifractor, Ruminococcaceae UBA1819 and Eubacterium, which have been associated with health [24, 45, 46]. Genera more abundant in future type 1 diabetes consisted of Firmicutes (Enterococcus, Gemella and Hungatella), as well as Bacteroides (Bacteroides and Porphyromonas). Genera contributing the most to core microbiome differentiation (i.e. with a higher MDA score but inconsistent patterns of abundance between cases and controls), such as Fusicatenibacter, Granulicatella, Roseburia and Ruminococcus, all belong to Firmicutes as well. Fusicatenibacter, a major contributor in core microbiome analysis, is of particular interest for its ability to create SCFAs and proinflammatory metabolites such as succinate [47]. Ruminococcus, a genus with the highest MDA score for both control infants and infants with future type 1 diabetes, has been associated with increased GADA production and inflammation [48, 49].
The pathological contribution of the differences that were observed in this investigation may be explained, in part, by the influence of differences in bacterial metabolism. The increased abundance of Firmicutes, primary producers of butyrate in the gut, that we observed in control infants parallels the increase in predicted acetyl-CoA fermentation to butanoate II, an ester of butyrate. Butyrate promotes intestinal homeostasis by inhibiting proinflammatory mediators and increasing epithelial barrier function [45]. Additional SCFAs are generated by fermentation of pyruvate to acetone, another predicted pathway higher in the ABIS control cohort. These predicted pathway differences confirm previous studies of at-risk populations describing a decrease in butyrate-producing bacteria with early onset of autoantibody development. Impairment in epithelial barrier function, resulting from reduced butyrate production, could prime an infant to the faulty immune activation that is responsible for autoimmune disorders such as type 1 diabetes by dislocation of antigens into the systemic circulation [24,25,26].
Furthermore, SCFA metabolites interact with T cell immunometabolism, a possible link to reported findings of intestinal inflammation in children with type 1 diabetes, signifying activation of mucosal innate and adaptive immunity combined with impaired induction of regulatory T cells in the small intestine [50]. Regulatory T cells participate in immune system tolerance to the body’s antigens and ingested antigens. When impaired, antigens may elicit an inappropriate immune response, such as a humoral response (Th2) to food allergens, through the secretion of IL-4, IL-5 and IL-13. Autoimmune diseases are associated with a cell-mediated (Th1) response through IL-2 and IFN-γ [51]. Regulatory T cell dysfunction is necessary for disease but other factors responsible for Th1 or Th2 imbalance determine the progression to autoimmunity or allergy. Altered gut microbiome metabolism, through regulatory T cell impairment and a Th1 overbalance, could explain the pathophysiological mechanism of previously proposed environmental factors, like dietary antigens and enterovirus infection, and autoimmunity in type 1 diabetes. Not surprisingly, duration of total breastfeeding and other dietary and environmental factors were found to be confounders of binomial β diversity in this study. Thus, control infants were carefully selected accordingly to be better able to determine the microbiome composition and associated functional SCFA pathways of 1-year-old infants with a future type 1 diabetes diagnosis.
By nature of 16S rRNA classification, a single genus is typically characterised by multiple ASVs. As expected with bacterial strains, ASVs with significant findings in a particular genus may differ in directional abundance. This could explain discrepancies that are observed at the genus level. For instance, Agathobacter had higher total abundance in control infants but had a higher relative abundance in case infants. This could be in part due to strain-level differences, as Agathobacter-387 was directionally split across analyses while Agathobacter-434 was observed to have higher total and relative abundance in case infants. Anaerostipes, with a higher total and relative abundance in control infants, contained two significant core ASVs, separated by only two nucleotides in the 16S rRNA: Anaerostipes-747, which was more abundant in control infants; and Anaerostipes hadrus 719, which had split patterns of abundance. Whether these strains behave differently in the microbiome is unknown but this is an area for future investigation. The same observation was made for Eggerthella and Eggerthella lenta-3665. The contribution made by Eggerthella lenta-3665 to the core microbiome differences (i.e. the MDA score) is much higher than was observed at the genus level, suggesting that the six other Eggerthella ASVs could be mitigating the impact of the genus. These examples demonstrate the importance of investigating strain-level differences in the gut microbiome, especially with the potential variability of bacterial function that is observed within a given genus.
A major strength of this investigation is the opportunity to study the gut microbiome at infancy in a non-HLA-restricted general population. Furthermore, extensive questionnaires allow for study of and matching on environmental factors, particularly those known to influence the microbiota or type 1 diabetes risk independently. While the sample size of the type 1 diabetes group is relatively small (~1% of the cohort), as expected in a general population cohort, the iterative process of matching in this investigation was employed to mitigate inherent differences in other factors mediating the control group. The fact that significant differences in the microbiota were observed, notwithstanding the iterative matching criteria, is a strength.
In conclusion, although the mean age at which type 1 diabetes was diagnosed was more than a decade after sample collection, at 1 year of age distinct microbial signatures were identified, with parallel observations in reduced predicted bacterial SCFA pathways. The autoimmune processes usually begin long before the onset of overt symptoms of type 1 diabetes [52], illustrating how differences in microbiome composition this early in life could shed important light on the complex interaction between the developing immune system, environmental exposures in childhood, and autoimmunity. The possibility of preventing disease onset by altering or promoting a ‘healthy’ gut microbiome is appealing.
Data availability
The forward and reverse 16S raw sequencing data generated in this study are available through the NCBI Sequence Read Archive under BioProject PRJNA875929. Associated sample metadata used for statistical comparison are available in the source data file. R codes used for statistical comparisons and figure generation are available at: https://github.com/PMilletich/T1D_Pipeline.
Abbreviations
- ABIS:
-
All Babies In Southeast Sweden
- ASV:
-
Amplicon sequence variant
- FDR:
-
False discovery rate
- MDA:
-
Mean decrease accuracy
- OOB:
-
Out-of-bag error
- PIME:
-
Prevalence interval for microbiome evaluation
- rRNA:
-
Ribosomal RNA
- SCFA:
-
Short-chain fatty acid
References
Rawshani A, Sattar N, Franzén S et al (2018) Excess mortality and cardiovascular disease in young adults with type 1 diabetes in relation to age at onset: a nationwide, register-based cohort study. Lancet 392(10146):477–486. https://doi.org/10.1016/S0140-6736(18)31506-X
Rewers M, Ludvigsson J (2016) Environmental risk factors for type 1 diabetes. Lancet 387(10035):2340–2348. https://doi.org/10.1016/S0140-6736(16)30507-4
Pociot F, Lernmark Å (2016) Genetic risk factors for type 1 diabetes. Lancet 387(10035):2331–2339. https://doi.org/10.1016/S0140-6736(16)30582-7
Berhan Y, Waernbaum I, Lind T, Möllsten A, Dahlquist G, Swedish Childhood Diabetes Study Group (2011) Thirty years of prospective nationwide incidence of childhood type 1 diabetes. Diabetes 60(2):577–581. https://doi.org/10.2337/db10-0813
Rogers MAM, Kim C, Banerjee T, Lee JM (2017) Fluctuations in the incidence of type 1 diabetes in the United States from 2001 to 2015: a longitudinal study. BMC Med 15(1):199. https://doi.org/10.1186/s12916-017-0958-6
Harjutsalo V, Sjöberg L, Tuomilehto J (2008) Time trends in the incidence of type 1 diabetes in Finnish children: a cohort study. Lancet 371(9626):1777–1782. https://doi.org/10.1016/S0140-6736(08)60765-5
Wahlberg J, Vaarala O, Ludvigsson J, the ABIS-study group (2006) Dietary risk factors for the emergence of type 1 diabetes-related autoantibodies in 2½-year-old Swedish children. Br J Nutr 95(3):603–608. https://doi.org/10.1079/BJN20051676
Virtanen SM, Nevalainen J, Kronberg-Kippilä C et al (2012) Food consumption and advanced β cell autoimmunity in young children with HLA-conferred susceptibility to type 1 diabetes: a nested case-control design. Am J Clin Nutr 95(2):471–478. https://doi.org/10.3945/ajcn.111.018879
Sadauskaitė-Kuehne V, Ludvigsson J, Padaiga Ž, Jašinskienė E, Samuelsson U (2004) Longer breastfeeding is an independent protective factor against development of type 1 diabetes mellitus in childhood. Diabetes Metab Res Rev 20(2):150–157. https://doi.org/10.1002/dmrr.425
Clausen TD, Bergholt T, Bouaziz O et al (2016) Broad-spectrum antibiotic treatment and subsequent childhood type 1 diabetes: a nationwide danish cohort study. PLoS ONE 11(8):e0161654. https://doi.org/10.1371/journal.pone.0161654
Zheng D, Liwinski T, Elinav E (2020) Interaction between microbiota and immunity in health and disease. Cell Res 30(6):492–506. https://doi.org/10.1038/s41422-020-0332-7
Wen L, Ley RE, Volchkov PYU et al (2008) Innate immunity and intestinal microbiota in the development of Type 1 diabetes. Nature 455(7216):1109–1113. https://doi.org/10.1038/nature07336
Arpaia N, Campbell C, Fan X et al (2013) Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504(7480):451–455. https://doi.org/10.1038/nature12726
Olszak T, An D, Zeissig S et al (2012) Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336(6080):489–493. https://doi.org/10.1126/science.1219328
Hansen CHF, Nielsen DS, Kverka M et al (2012) Patterns of early gut colonization shape future immune responses of the host. PLoS ONE 7(3):e34043. https://doi.org/10.1371/journal.pone.0034043
Paun A, Danska JS (2016) Modulation of type 1 and type 2 diabetes risk by the intestinal microbiome: Role of gut microbiome in diabetes. Pediatr Diabetes 17(7):469–477. https://doi.org/10.1111/pedi.12424
Davis-Richardson AG, Ardissone AN, Dias R et al (2014) Bacteroides dorei dominates gut microbiome prior to autoimmunity in Finnish children at high risk for type 1 diabetes. Front Microbiol 5:678. https://doi.org/10.3389/fmicb.2014.00678
Endesfelder D, zu Castell W, Ardissone A et al (2014) Compromised gut microbiota networks in children with anti-islet cell autoimmunity. Diabetes 63(6):2006–2014. https://doi.org/10.2337/db13-1676
Stewart CJ, Ajami NJ, O’Brien JL et al (2018) Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562(7728):583–588. https://doi.org/10.1038/s41586-018-0617-x
Brown CT, Davis-Richardson AG, Giongo A et al (2011) Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS ONE 6(10):e25792. https://doi.org/10.1371/journal.pone.0025792
Kostic AD, Gevers D, Siljander H et al (2015) The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe 17(2):260–273. https://doi.org/10.1016/j.chom.2015.01.001
Gülden E, Wong FS, Wen L (2015) The gut microbiota and type 1 diabetes. Clin Immunol 159(2):143–153. https://doi.org/10.1016/j.clim.2015.05.013
de Goffau MC, Fuentes S, van den Bogert B et al (2014) Aberrant gut microbiota composition at the onset of type 1 diabetes in young children. Diabetologia 57(8):1569–1577. https://doi.org/10.1007/s00125-014-3274-0
Mokhtari P, Metos J, Anandh Babu PV (2021) Impact of type 1 diabetes on the composition and functional potential of gut microbiome in children and adolescents: possible mechanisms, current knowledge, and challenges. Gut Microbes 13(1):1926841. https://doi.org/10.1080/19490976.2021.1926841
Endesfelder D, Engel M, Davis-Richardson AG et al (2016) Towards a functional hypothesis relating anti-islet cell autoimmunity to the dietary impact on microbial communities and butyrate production. Microbiome 4(1):17. https://doi.org/10.1186/s40168-016-0163-4
Vaarala O (2012) Gut microbiota and type 1 diabetes. Rev Diabet Stud 9(4):251–259. https://doi.org/10.1900/RDS.2012.9.251
Stojanov S, Berlec A, Štrukelj B (2020) The influence of probiotics on the firmicutes/bacteroidetes ratio in the treatment of obesity and inflammatory bowel disease. Microorganisms 8(11):1715. https://doi.org/10.3390/microorganisms8111715
Honkanen H, Oikarinen S, Nurminen N et al (2017) Detection of enteroviruses in stools precedes islet autoimmunity by several months: possible evidence for slowly operating mechanisms in virus-induced autoimmunity. Diabetologia 60(3):424–431. https://doi.org/10.1007/s00125-016-4177-z
Russell JT, Roesch LFW, Ördberg M et al (2019) Genetic risk for autoimmunity is associated with distinct changes in the human gut microbiome. Nat Commun 10(1):3621. https://doi.org/10.1038/s41467-019-11460-x
Ilonen J, Kiviniemi M, Lempainen J et al (2016) Genetic susceptibility to type 1 diabetes in childhood - estimation of HLA class II associated disease risk and class II effect in various phases of islet autoimmunity: HLA class II effects in T1D. Pediatr Diabetes 17:8–16. https://doi.org/10.1111/pedi.12327
Socialstyrelsen (2022) National Patient Register. Available from https://www.socialstyrelsen.se/en/statistics-and-data/registers/national-patient-register/. Accessed 6 Jul 2022
Eeg-Olofsson K, Linder E, National Diabetes Registry. Nationella DiabetesRegistret. Available from https://www.ndr.nu/#/english. Accessed 6 Jul 2022
Socialstyrelsen (2022) National Prescribed Drug Register. Available from https://www.socialstyrelsen.se/en/statistics-and-data/registers/national-prescribed-drug-register/. Accessed 6 Jul 2022
Russell JT, Lauren Ruoss J, de la Cruz D et al (2021) Antibiotics and the developing intestinal microbiome, metabolome and inflammatory environment in a randomized trial of preterm infants. Sci Rep 11(1):1943. https://doi.org/10.1038/s41598-021-80982-6
Milletich PL, Ahrens AP, Russell JT et al (2022) Gut microbiome markers in subgroups of HLA class II genotyped infants signal future celiac disease in the general population: ABIS study. Front Cell Infect Microbiol 12:920735. https://doi.org/10.3389/fcimb.2022.920735
Quast C, Pruesse E, Yilmaz P et al (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):D590–D596. https://doi.org/10.1093/nar/gks1219
R Core Team (2022) R: A language and environment for statistical computing. Version 4.2.2. Available from https://www.r-project.org/
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8(4):e61217. https://doi.org/10.1371/journal.pone.0061217
Oksanen J, Simpson GL, Blanchet FG et al (2022) vegan: community ecology package. Version 2.6-4. Available from https://CRAN.R-project.org/package=vegan
Kassambara A (2020) ggpubr: “ggplot2” based publication ready plots. Version 0.5.0. https://CRAN.R-project.org/package=ggpubr
Wickham H, Chang W, Henry L et al (2022) ggplot2: create elegant data visualisations using the grammar of graphics. Version 3.4.0. https://ggplot2.tidyverse.org
Roesch LFW, Dobbler PT, Pylro VS, Kolaczkowski B, Drew JC, Triplett EW (2020) PIME: a package for discovery of novel differences among microbial communities. Mol Ecol Resour 20(2):415–428. https://doi.org/10.1111/1755-0998.13116
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Muñoz-Tamayo R, Laroche B, Walter É et al (2011) Kinetic modelling of lactate utilization and butyrate production by key human colonic bacterial species: Modelling butyrate production by colonic bacterial species. FEMS Microbiol Ecol 76(3):615–624. https://doi.org/10.1111/j.1574-6941.2011.01085.x
Carlier J-P, Bedora-Faure M, K’ouas G, Alauzet C, Mory F (2010) Proposal to unify Clostridium orbiscindens Winter et al. 1991 and Eubacterium plautii (Séguin 1928) Hofstad and Aasjord 1982, with description of Flavonifractor plautii gen. nov., comb. nov., and reassignment of Bacteroides capillosus to Pseudoflavonifractor capillosus gen. nov., comb. nov. Int J Syst Evol Microbiol 60(3):585–590. https://doi.org/10.1099/ijs.0.016725-0
Takada T, Kurakawa T, Tsuji H, Nomoto K (2013) Fusicatenibacter saccharivorans gen. nov., sp. nov., isolated from human faeces. Int J Syst Evol Microbiol 63(Pt_10):3691–3696. https://doi.org/10.1099/ijs.0.045823-0
Luo S, Yue T, Liu Z et al (2022) Gut microbiome and metabolic activity in type 1 diabetes: an analysis based on the presence of GADA. Front Endocrinol 13:938358. https://doi.org/10.3389/fendo.2022.938358
Zhou H, Zhao X, Sun L et al (2020) Gut microbiota profile in patients with type 1 diabetes based on 16S rRNA gene sequencing: a systematic review. Dis Markers 2020:1–11. https://doi.org/10.1155/2020/3936247
Vaarala O, Atkinson MA, Neu J (2008) The “Perfect Storm” for type 1 diabetes. Diabetes 57(10):2555–2562. https://doi.org/10.2337/db08-0331
van Oosterhout AJM (2005) Th1/Th2 paradigm: not seeing the forest for the trees? Eur Respir J 25(4):591–593. https://doi.org/10.1183/09031936.05.00014105
Ziegler AG, Rewers M, Simell O et al (2013) Seroconversion to multiple islet autoantibodies and risk of progression to diabetes in children. JAMA 309(23):2473. https://doi.org/10.1001/jama.2013.6285
Acknowledgements
We thank I. Johansson, G. Smolinska and Å. Faresjö from Linköping University for their help with sample collection and distribution and for providing authors with register data.
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Contribution statement
PLM performed data generation and statistical analysis. All authors contributed to data analysis and data interpretation. MB and PLM drafted the manuscript. JL created ABIS, designed the study, and carried out sample collection and processing. All authors were involved in manuscript revisions and approval of the final revision. EWT is responsible for the integrity of the work as a whole.
Funding
ABIS was supported by the following funding bodies: Barndiabetesfonden (Swedish Child Diabetes Foundation); Swedish Council for Working Life and Social Research (grant/award numbers FAS2004-1775, FAS2004–1775); Swedish Research Council (grant/award numbers K2005-72X-11242-11A, K2008-69X-20826-01-4, K2008-69X-20826-01-4); Östgöta Brandstodsbolag; Medical Research Council of Southeast Sweden (FORSS); JDRF Wallenberg Foundation (grant/award no. K 98-99D-12813-01A); and ALF and LFoU grants from Region Östergötland and Linköping University, Sweden. The microbiome analysis was supported by an Innovation Award (1-INO-2018-637-A-N) from JDRF to EWT.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Malin Bélteky and Patricia L. Milletich share first authorship.
Supplementary information
ESM
(PDF 4520 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bélteky, M., Milletich, P.L., Ahrens, A.P. et al. Infant gut microbiome composition correlated with type 1 diabetes acquisition in the general population: the ABIS study. Diabetologia 66, 1116–1128 (2023). https://doi.org/10.1007/s00125-023-05895-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-023-05895-7