Abstract
Aims/hypothesis
Elucidating the molecular mechanisms of fat accumulation and its metabolic consequences is crucial to understanding and treating obesity, an epidemic disease. We have previously observed that Usp19 deubiquitinating enzyme-null mice (Usp19−/−) have significantly lower fat mass than wild-type (WT) mice. Thus, this study aimed to provide further understanding of the role of ubiquitin-specific peptidase 19 (USP19) in fat development, obesity and diabetes.
Methods
In this study, the metabolic phenotypes of WT and Usp19−/− mice were compared. The stromal vascular fractions (SVFs) of inguinal fat pads from WT and Usp19−/− mice were isolated and cells were differentiated into adipocytes in culture to assess their adipogenic capacity. Mice were fed a high-fat diet (HFD) for 18 weeks. Body composition, glucose metabolism and metabolic variables were assessed. In addition, following insulin injection, signalling activity was analysed in the muscle, liver and adipose tissue. Finally, the correlation between the expression of Usp19 mRNA and adipocyte function genes in human adipose tissue was analysed.
Result
Upon adipogenic differentiation, SVF cells from Usp19−/− failed to accumulate lipid and upregulate adipogenic genes, unlike cells from WT mice. Usp19−/− mice were also found to have smaller fat pads throughout the lifespan and a higher percentage of lean mass, compared with WT mice. When fed an HFD, Usp19−/− mice were more glucose tolerant, pyruvate tolerant and insulin sensitive than WT mice. Moreover, HFD-fed Usp19−/− mice had enhanced insulin signalling in the muscle and the liver, but not in adipose tissue. Finally, USP19 mRNA expression in human adipose tissue was positively correlated with the expression of important adipocyte genes in abdominal fat depots, but not subcutaneous fat depots.
Conclusions/interpretation
USP19 is an important regulator of fat development. Its inactivation in mice exerts effects on multiple tissues, which may protect against the negative metabolic effects of high-fat feeding. These findings suggest that inhibition of USP19 could have therapeutic potential to protect from the deleterious consequences of obesity and diabetes.
Similar content being viewed by others

Introduction
Obesity and the metabolic syndrome that accompanies it are serious global health concerns. Nearly two billion people are overweight or obese worldwide and obesity is a significant risk factor for the development of metabolic dysfunction including dyslipidaemia, hyperinsulinaemia and glucose intolerance [1]. The development of type 2 diabetes and the subsequent health complications, such as nephropathy, retinopathy and neuropathy, underlies the importance of solving the primordial problem of obesity. Understanding the molecular mechanisms of fat cell development and function will build an important scientific foundation to address the global epidemic of obesity.
A key regulator of many cellular processes is the ubiquitin proteasome system. The 8 kDa peptide ubiquitin is conjugated to proteins through the sequential action of ubiquitin activating, conjugating and ligating enzymes [2]. The length and the nature of the linkage of the ubiquitin chain can direct the fate of the targeted protein and many polyubiquitinated proteins are targeted to the 26S proteasome for degradation. Ubiquitination is a reversible process that is catalysed by deubiquitinating enzymes. There are ~90 deubiquitinating enzymes in the human genome that are divided into five families based on the homology of their catalytic domains [3, 4]. Ubiquitin-specific peptidase 19 (USP19) is a deubiquitinating enzyme in the ubiquitin-specific protease family. It is involved in many cellular processes including autophagy and immune response [5, 6], endoplasmic reticulum-associated degradation [7], misfolding associated protein secretion [8], cell proliferation [9] and hypoxia [10]. Most notably, USP19 plays an important role in muscle; USP19 regulates myogenic differentiation [11], and female muscle mass through oestrogen receptor-dependent mechanisms [12]. Importantly, Usp19 expression is induced in muscle under catabolic stimuli [13] and the inactivation of the gene in mice is protective against muscle wasting [14, 15].
Interestingly, Usp19−/− mice also appeared to have decreased fat mass. We therefore decided to characterize this in detail here and we hypothesised that USP19 may be important in processes such as fat development and accumulation, as well as obesity and its complications, such as diabetes. In this report, we explored whether USP19 is important for adipogenesis, whether Usp19−/−mice respond differently from wild-type (WT) mice when challenged with a high-fat diet (HFD) and, finally, whether USP19 might be involved in human adipose tissue function.
Methods
Animals
All animal studies were approved by the Animal Care Committees of McGill University or the McGill University Health Centre Research Institute. Animals were housed in high efficiency particulate air (HEPA)-filtered cages with a 12-h day/night cycle at 21°C with environmental enrichment and fed a normal diet composed of 32% protein, 14% fat and 54% carbohydrates as energy sources. The mice were generally housed in groups (maximum n = 5 per cage), but males were frequently isolated due to aggressive behaviour. Usp19−/− mice were generated in our laboratory, as previously described [15]. Heterozygous mice (C57BL/6 background) were mated to generate WT and Usp19−/− mice. All available animals were used for each experiment; thus, groups were assigned based on genotype, without randomisation. For HFD studies, male mice were fed a diet with 60% of energy from fat (Harlan Laboratories, Madison, WI, USA), from 8 weeks of age for 18 weeks. Each HFD experiment was repeated three times. Unless otherwise indicated, representative data from one experiment is presented in the results. Mice were subjected to body composition measurements by magnetic resonance imaging (MRI; EchoMRI, Houston, TX, USA) before and during the HFD.
Histology
Fat tissue from the epididymal fat pad was fixed, embedded in paraffin, sectioned and stained with haematoxylin and eosin. Cross-sectional areas were measured by manually selecting an area of the section and tracing the outline of adipocytes in ImageJ or by the open source Adiposoft software plugin for ImageJ (https://imagej.nih.gov/ij/; version 1.45s) [16]. Five different fields per animal were used and ~50 adipocytes per field were manually counted or the entire field was counted with Adiposoft. The observer was blinded to the genotype during quantification.
Stromal vascular fraction isolation
The stromal vascular fraction (SVF) of the inguinal fat pad was isolated as follows: inguinal fat pads were removed from male WT or Usp19−/− mice and placed in 37°C DMEM/F12 media (Thermo Fisher Scientific, Waltham, MA, USA) before processing. Tissue from 3–5 fat pads was minced and digested for 1 h in collagenase solution, shaking at 37°C. Digested tissue was passed through a 240 μm filter to remove debris and undigested tissue. Cells were then centrifuged at 50 g for 5 min at room temperature to separate the adipocytes. The infranatant (below the floating adipocyte layer) containing the SVF cells was collected into a new tube and SVF cells were centrifuged at 500 g for 15 min. All but 5 ml of the media was removed and 10 ml of red blood cell lysis buffer (155 mmol/l NH4Cl, 10 mmol/l KHCO3, 10 mmol/l EDTA) was added, after which the solution was vortexed and incubated at room temperature for 5 min. DMEM/F12 was added to a total volume of 30 ml and cells were passed through a 20 μm filter to further purify the SVF cells. The cells were then centrifuged at 500 g for 15 min to pellet the SVF cells. These cells were resuspended in DMEM/F12 supplemented with 10% FBS (vol./vol.) and penicillin/streptomycin and plated at a density of 2.4 × 105 cells per well in a 6-well plate and grown until confluence. Cells were induced to differentiate with 1 μmol/l dexamethasone, 0.5 μmol/l 3-isobutyl-1-methylxanthine (IBMX), 200 μmol/l indometacin and 10 μg/μl insulin (all from Sigma-Aldrich, St. Louis, MO, USA), for 48 h. Differentiating cells were maintained in DMEM/F12 with 1 μg/μl insulin.
Oil Red O staining
Differentiated adipocytes were washed three times with cold PBS. Then, the cells were fixed for 15 min at room temperature in 4% (vol./vol.) paraformaldehyde in PBS. Cells were incubated for 5 min in 60% (vol./vol.) isopropanol and then incubated for 15 min with Oil Red O solution. Oil Red O solution was removed and cells were incubated again with 60% isopropanol. Finally, cells were rinsed three times with distilled water and staining was visualised.
Quantitative real-time PCR
RNA was prepared from cells or tissue by solubilisation in 4 mol/l guanidium isothiocyanate then ultracentrifugation through a CsCl cushion followed by phenol-chloroform extraction and RNA precipitation. RNA was quantified using a NanoDrop LITE spectrophotometer (Thermo Fisher Scientific). cDNA was synthesised from 1 μg of RNA using the High Capacity cDNA Reverse Transcriptase Kit (Applied Biosystems, Foster City, CA, USA). Quantitative real-time PCR (qPCR) analysis was done using SYBR-Green Master Mix (Applied Biosystems) on a ViiA7 thermal cycler with gene-specific primers for mouse Adipoq, Cebpa, Fabp4, Gapdh, Lep, Pparg and Usp19 and human ADIPOQ, FABP4, GAPDH, LEP, PPARG and USP19 (electronic supplementary material [ESM] Table 1). Differences in target gene expression were calculated using the ΔΔCt method.
Western blot
Tibialis anterior muscles, epididymal white adipose tissue (eWAT) and liver were homogenised in ice cold radioimmunoprecipitation assay (RIPA) buffer with protease and phosphatase inhibitor cocktails (Roche, Indianapolis, IN, USA). The homogenate was spun in a refrigerated centrifuge at 17,000 g for 15 min to clear the lysate. Protein concentration was determined by the BCA Micro Protein Assay (Thermo Fisher Scientific). Lysates (10 μg) were subjected to SDS-PAGE and transferred to 0.45 μm nitrocellulose membranes. Membranes were probed with primary antibodies (ESM Table 2) followed by secondary antibodies conjugated to horseradish peroxidase with binding detected using a chemiluminescent assay (Clarity ECL; Bio-Rad, Hercules, CA, USA). Membranes were visualised with a ChemiDoc Touch Camera (Bio-Rad) and the signals quantified using Image Lab software (Bio-Rad).
Glucose metabolism studies
Glucose tolerance tests (GTTs), insulin tolerance tests (ITTs) and pyruvate tolerance tests (PTTs) were performed in animals. HFD-fed mice were fasted for 16 h overnight for GTTs, PTTs and fasting insulin analysis, or for 4 h for ITTs. For GTTs, ITTs and PTTs, 2 mg/g glucose (Sigma-Aldrich), 1 U/kg crystalline zinc insulin (Humulin R; Lilly, Indianapolis, IN, USA) or 2 mg/g sodium pyruvate (Sigma-Aldrich), respectively, was administered to mice intraperitoneally. Blood was collected via tail-vein puncture and blood glucose was measured with an AccuChek glucometer (Roche) at various time points. For fasting insulin, blood was collected from the saphenous vein following overnight food deprivation and allowed to clot. Serum insulin levels were measured with an insulin single plex assay kit (Meso Scale Discovery, Rockville, MD, USA).
Metabolic studies
Animals were housed in TSE PhenoMaster (TSE Systems, Bad Homburg, Germany) metabolic cages for 1 week of acclimatisation, followed by 1 week of measurements including food and water intake, locomotor activity measured by infrared light beams, and indirect calorimetry. Each study was carried out once, with the number of mice indicated in the figure legends.
Human tissue correlation analysis
Human mesenteric, omental and abdominal subcutaneous adipose tissue samples were obtained from the biobank at Institut universitaire de cardiologie et de pneumologie de Québec (IUCPQ), where written informed consent was obtained from the study participants. Female participants (BMI ranges: 35–40 kg/m2 and 60–69 kg/m2) were paired for age and date of bariatric surgery and participants with diabetes were excluded. Study approval was obtained from the Research Ethics Boards of the McGill University Health Centre and the IUCPQ.
Statistical analysis
Data are presented as mean ± SEM unless otherwise stated. Statistical analysis was performed using Graphpad Prism 6 software (www.graphpad.com/scientific-software/prism/; version 6.0c). Analysis was carried out using the unpaired student’s t test if comparing a single variable or a two-way ANOVA with group (genotype) by time interaction if comparing multiple independent variables. Energy expenditure was analysed by direct normalisation to tissue mass and also by ANCOVA, with lean mass plus 0.2 times fat mass as a covariate to control for the different metabolic capacities of lean and fat tissue. The ANCOVA analysis of energy expenditure was provided by the National Institute of Diabetes and Digestive Kidney Diseases (NIDDK) National Mouse Metabolic Phenotyping Centers (MMPC) using the Energy Expenditure Analysis page (www.mmpc.org/shared/regression.aspx, accessed 5 May 2018) and supported by grants DK076169 and DK115255. Bivariate correlation between the expression of genes was tested using the Pearson correlation coefficient. Significance was set at p < 0.05.
Results
Usp19 −/− mice have less white adipose tissue than WT mice
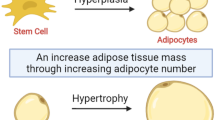
On a normal diet, male Usp19−/− mice had significantly smaller epididymal fat pads compared with WT mice (Fig. 1a, b). After normalisation to body weight, epididymal fat pads were still significantly smaller in Usp19−/− mice (ESM Fig. 1a). This decrease in fat mass was consistent throughout the lifespan from as early as 21 days to as late as 24 months of age. Total body fat mass in Usp19−/− mice, as measured by MRI, was also significantly lower than in WT (ESM Fig. 1b). The inguinal subcutaneous fat was also smaller in the Usp19−/− mice at 8 weeks of age (ESM Fig. 1c), while brown adipose tissue weight was similar between WT and Usp19−/− mice (ESM Fig. 1d). Fat tissue expansion is the result of two processes: adipocyte hypertrophy and/or adipocyte hyperplasia. To determine if Usp19−/− mice have a defect resulting in adipocyte hypertrophy or hyperplasia, we assessed the size of the adipocytes in the fat pads of WT and Usp19−/− mice. Epididymal adipocytes in Usp19−/− mice had significantly smaller cross-sectional areas from 6 months of age (Fig. 1c, d). Interestingly, the adipocyte size was similar between WT and Usp19−/− mice at an early age (2 months; Fig. 1c, d), yet the fat pad was significantly smaller at this age (Fig. 1b), an indication that fewer cells were present. These findings suggested that USP19 may be modulating fat cell development.
Usp19−/− mice have less fat than WT mice. (a) Representative images of epididymal fat pads from male WT and Usp19−/− mice at 2 months of age. Scale bar, 1 cm. (b) Epididymal fat mass at 0.75, 2, 6 and 24 months of age in WT and Usp19−/− mice. 0.75 months, n=9; 2 months, n=5; 6 months, WT n=10 and Usp19−/− n=8; 24 months, WT n=15 and Usp19−/−, n=12. (c) Representative images of epididymal fat sections from WT and Usp19−/− mice at 2, 6 and 24 months of age. Scale bar, 50 μm. (d) Cross-sectional area (CSA) of adipocytes in the epididymal fat pad (number of mice: 2 months, n=4; 6 months, n=4; 24 months, WT n=6 and Usp19−/− n=3); n>250 adipocytes per animal from five different fields. *p<0.05, **p<0.01
USP19 is required for normal adipogenesis
Adipogenesis is regulated by a transcriptional cascade hinging on the positive regulatory loop between the transcription factors peroxisome proliferator-activated receptor γ (PPARγ) and CCAAT/enhancer-binding protein α (CEBPα), the transcriptional activities of which also drive the expression of adipocyte-specific genes [17]. To further dissect the role of USP19 in adipogenesis, we isolated the SVF from the inguinal fat pads of WT and Usp19−/− mice and differentiated them into adipocytes in vitro. Usp19−/− cells failed to accumulate lipid, unlike WT cells, as shown by marked reductions in staining by Oil Red O, which labels neutral lipids (Fig. 2a). The differentiated cells from Usp19−/− mice expressed significantly lower levels of the key regulator of adipogenesis, Pparg, as well as genes important for adipocyte function, such as Fabp4, Adipoq and Lep (Fig. 2b). Additionally, the cells isolated from the Usp19−/− mice expressed significantly lower levels of PPARγ protein (Fig. 2c [quantification not shown]). Therefore, it may be concluded that USP19 is required for robust adipogenic differentiation of cells in the SVF.
USP19 is required for normal adipogenesis. Cells from the SVF from inguinal fat pads from male WT and Usp19−/− mice were isolated and differentiated in culture. (a) Representative brightfield images of differentiation of cells from WT and Usp19−/− mice and image of Oil Red O staining on day 9 of differentiation. Scale bar, 200 μm. (b) Relative expression of adipogenic genes on day 9 of WT and Usp19−/− SVF cell differentiation, calculated as fold change compared with WT; n=3 independent experiments with four biological replicates per experiment. (c) Cells were harvested at day 9 of differentiation and immunoblotted for USP19, PPARγ isoforms (PPARγ1 and PPARγ2) and tubulin; n=3. **p<0.01
Usp19 −/− mice have an altered energy balance, favouring a leaner phenotype on an HFD
To test whether absence of USP19 would prevent the mice from developing obesity, we challenged the mice with an HFD containing 60% of the energy as fat. We found that Usp19−/− mice weighed less than WT mice over the 18 weeks that mice were fed an HFD (Fig. 3a). To understand what may be contributing to this difference in weight gain, the mice were housed in metabolic cages while indirect calorimetry, activity and food intake were measured. Usp19−/− mice had significantly increased activity levels when compared with WT mice, when fed either a normal diet (ESM Fig. 2a) or an HFD (Fig. 3b). Similar increases in activity were observed in the Usp19−/− mice during both the light and the dark cycle. Additionally, Usp19−/− mice consumed more food than WT mice when fed a normal diet (ESM Fig. 2b), but similar amounts of food when HFD-fed (Fig. 3c). Oxygen consumption and energy expenditure were similar between WT and Usp19−/− mice (Fig. 3d, f). When normalised to body weight, however, oxygen consumption and energy expenditure were significantly increased in the Usp19−/− mice (Fig. 3e, g). Since normalisation to body weight does not take into account differences in body composition between the two genotypes, we performed additional analyses. When normalised to lean mass + (0.2 × fat mass), which reflects the major contribution of skeletal muscle to energy expenditure with a minor contribution from fat, oxygen consumption and energy expenditure remained elevated in the Usp19−/− mice when compared with WT mice (Fig. 3e, g). When fed a normal diet, there were no significant differences in oxygen consumption and energy expenditure between Usp19−/− and WT mice (ESM Fig. 2c, d). Finally, since this normalisation may still not optimally correct for body composition [18], we performed an ANCOVA examining the relationship between energy expenditure and lean mass + (0.2 × fat mass) in WT and Usp19−/− mice. This analysis did not reveal a statistically significant increase in energy expenditure in the Usp19−/− mice, as compared with WT, neither when fed the normal diet nor the HFD. However, considering the borderline non-significant p values obtained from this analysis (p = 0.1; ESM Fig. 3a, b) and the much larger sample sizes usually required for such analyses, a difference between the genotypes remains possible. Taken together with the observations that normal diet-fed Usp19−/− mice have higher food intake but decreased body weight and higher activity than WT mice, these findings can be used to argue that these mice have higher energy expenditure. The respiratory exchange ratio (RER) was not significantly different between WT and Usp19−/− mice on an HFD (Fig. 3h) and lower than on normal diet (ESM Fig. 2e), indicating a shift towards fat oxidation when either genotype was fed an HFD. To assess the body composition of the mice, they were subjected to MRI scanning. Interestingly, the Usp19−/− mice had a significantly lower percentage of fat mass and a higher percentage of lean mass before and during HFD, as compared with WT mice under the same conditions (Fig. 3i, j).
Usp19−/− mice have an altered energy balance favouring a leaner phenotype when fed an HFD. Male WT and Usp19−/− mice were fed an HFD (60% of energy as fat). (a) Body weight of WT and Usp19−/− mice over 18 weeks; n=17; p<0.0001, ANOVA. (b) After 5 weeks on an HFD, activity counts were measured hourly for 48 h and (c) total food intake over 7 days was measured; n=8. (d) Calorimetry was performed and oxygen consumption was measured; n=8. (e) Oxygen consumption was normalised to body weight or lean mass + (0.2 × fat mass); n=8. (f) Energy expenditure was calculated in kJ per day; n=8. (g) Energy expenditure was normalised to body weight or lean mass + (0.2 × fat mass); n=8. (h) Respiratory exchange ratio was calculated by dividing carbon dioxide production by oxygen consumption; n=8. (i) Fat mass and (j) lean mass, as measured by MRI, of WT and Usp19−/− mice before HFD feeding and at 3 and 13 weeks after HFD feeding, presented as a percentage of total body weight; n=8. Circles, WT; triangles, Usp19−/−. BW, body weight; EE, energy expenditure; FM, fat mass; LM, lean mass; RER, respiratory exchange ratio. *p<0.05, **p<0.01
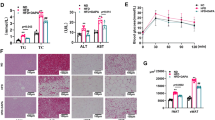
At euthanasia after HFD, as compared with WT mice, Usp19−/− mice had significantly lower body weight and decreased body length (ESM Fig. 4a). Usp19−/− mice also had smaller subcutaneous and brown adipose tissue fat pads, but the difference in epididymal fat pads did not achieve statistical significance (p = 0.09) (ESM Fig. 4c). Adipocyte size was significantly smaller in the eWAT and subcutaneous white adipose tissue (scWAT) depots of the Usp19−/− mice, as compared with WT mice (ESM Fig. 5a, b). Consistent with the MRI results indicating a leaner body, Usp19−/− mice had significantly larger tibialis anterior muscles than WT mice. In addition, the mean gastrocnemius muscle mass was larger compared with WT mice, but this did not reach statistical significance (p = 0.09) (ESM Fig. 4d). Usp19−/− mice also had significantly smaller livers and hearts than WT mice (ESM Fig. 4e). Therefore, the overall body composition of Usp19−/− mice at euthanasia is consistent with a leaner body, with larger muscles and less fat, as compared with WT mice.
Usp19 −/− mice have improved glucose homeostasis
Based on the increased activity levels and leaner body composition of Usp19−/− mice compared with WT mice, we evaluated fasting glucose and insulin levels in HFD-fed mice. Usp19−/− mice had significantly lower fasting insulin levels as compared with WT mice, despite similar fasting glucose levels (Fig. 4a, b), suggesting that Usp19−/− mice are more insulin sensitive. To assess this further, we challenged these mice with an ITT. Indeed, upon insulin administration, we saw more profound blood glucose lowering in Usp19−/− mice than WT mice (Fig. 4c). Increased insulin sensitivity should result in an increased ability to lower blood glucose when challenged with a glucose load. Indeed, Usp19−/− mice were able to lower blood glucose levels more effectively than WT mice (Fig. 4d). Another component of overall blood glucose homeostasis is glucose production from the liver, which is under the regulation of several hormones including insulin. To test whether Usp19−/− mice were also more insulin sensitive in the liver, pyruvate, a substrate for gluconeogenesis, was administered to the mice and their blood glucose levels were monitored over time. Usp19−/− mice showed a lower level of glucose production compared with WT mice during the PTT (Fig. 4e), consistent with increased insulin sensitivity in the liver. Thus, Usp19−/− mice have improved glucose metabolism when fed an HFD.
Usp19−/− mice have improved glucose homeostasis. Male WT and Usp19−/− mice were fasted overnight for 16 h. (a) Fasting insulin levels (n=10) and (b) blood glucose levels (n=10) were assessed. (c) ITTs were performed by injecting mice with 1 U/kg insulin intraperitoneally and measuring blood glucose over time. Values normalised to blood glucose at time 0 are shown; WT, n=22 and Usp19−/−, n=20; p<0.01, ANOVA, pooled from two experiments. AUC was calculated over the first 30 min after insulin administration. (d) GTTs were performed by injecting mice with 2 mg/g d-glucose intraperitoneally and measuring blood glucose over time; WT, n=16, Usp19−/−, n=13; p<0.0001, ANOVA. AUC was calculated over the entire 120 min period. (e) PTTs were performed by injecting mice with 2 mg/g sodium pyruvate intraperitoneally and measuring blood glucose over time; WT, n=12 and Usp19−/−, n=16; p<0.0001, ANOVA. AUC was calculated over the entire 120 min period. *p<0.05, **p<0.01
Insulin signalling is enhanced in Usp19 −/− mouse muscle and liver
To explore the cellular physiology of improved glucose homeostasis in Usp19−/− mice, we assessed the activation of Akt–mechanistic target of rapamycin (mTOR) signalling pathway in insulin sensitive tissues: the muscle, liver and epididymal adipose tissue. HFD-fed Usp19−/− mice had increased levels of p-Akt and p-S6K (also known as p70/S6K) in the muscle, as compared with WT mice (Fig. 5a). They also had increased p-Akt content in the liver (Fig. 5b), but no changes were detected in levels of these proteins in adipose tissue (Fig. 5c). This suggests that the ability of USP19 to maintain better glucose homeostasis is due to increased insulin signalling in the muscle and the liver, but possibly not in visceral adipose tissue.
Insulin signalling is enhanced in Usp19−/− mouse muscle and liver. WT and Usp19−/− mice were injected with 1 U/kg insulin 20 min prior to euthanasia. (a) Tibialis anterior muscle (WT, n=12 and Usp19−/−, n=11), (b) liver (WT, n=11 and Usp19−/−, n=11) and (c) eWAT (WT, n=8 and Usp19−/−, n=5) samples were homogenised. Homogenates were analysed by western blot for the indicated components of the insulin signalling pathway. The graphs show quantification of insulin-stimulated phosphorylation of Akt on the threonine 308 residue normalised to the total amount of Akt (p-Akt/Akt) and of p-S6K normalised to the total amount of S6K (p-S6K/S6K), relative to WT. −/−, Usp19−/− mice. Circles, WT; triangles, Usp19−/−. **p<0.01
USP19 mRNA expression in human mesenteric and omental adipose tissue is positively correlated with adipogenic gene expression
Finally, we investigated whether the USP19 modulation of fat accumulation that we observed in mice may occur in humans. We obtained human mesenteric, omental and abdominal subcutaneous adipose tissue from two groups of female participants, with BMI in the ranges of 35–40 kg/m2 or 60–69 kg/m2, who had undergone bariatric surgery. Participants were paired for age and date of bariatric surgery, while participants with diabetes were excluded. Fasting plasma glucose, total cholesterol and triacylglycerol levels were similar between groups (ESM Table 3). There was no difference in USP19 mRNA expression between the two groups (those with high BMI [35–40 kg/m2] or very high BMI [60–69 kg/m2]) in any fat depot (ESM Fig. 6). Since we observed that Usp19−/− mice have a lower adipogenic capacity, we next asked whether human adipose tissue with higher USP19 expression might be more adipogenic than adipose tissue with lower USP19 expression. To address this, we explored whether USP19 mRNA expression correlates with the mRNA expression of genes important for adipocyte differentiation and function (PPARG, FABP4, ADIPOQ and LEP). We found that USP19 expression was significantly positively correlated with the expression of PPARG, FABP4 and ADIPOQ, but not LEP, in the mesenteric and omental fat biopsies (Fig. 6a–h). Interestingly, these positive correlations were not observed in the subcutaneous fat biopsies (Fig. 6i–l). Thus, USP19 might be involved in adipogenesis in human visceral adipose tissue depots, but not subcutaneous depots.
USP19 mRNA expression is positively correlated with adipogenic gene expression in human adipose tissue. (a–d) Mesenteric white adipose tissue, (e–h) omental white adipose tissue and (i–l) scWAT RNA expression was analysed. Relative USP19 expression is plotted against the expression of PPARg, FABP4, ADIPOQ or LEP. Pearson correlation analysis was performed; r and p values are presented in the figure. n=12 participants
Discussion
In this report, we show for the first time that the deubiquitinating enzyme USP19 is important in adipose tissue development and has profound effects on glucose metabolism. Inactivation of Usp19 reduced body weight, which was mainly due to decreased fat mass since lean body mass was proportionally higher in the Usp19−/− mice (Fig. 3). Importantly, these effects persisted throughout life (Fig. 1). At a physiological level, Usp19−/− mice manifested improved systemic insulin sensitivity and glucose tolerance (Fig. 4). Furthermore, these effects were not only present on a normal diet, but persisted on extended high-fat feeding, indicating the possibility of protection against the development of type 2 diabetes.
USP19 exerts these effects through actions in multiple tissues. Usp19−/− mice have a defect in fat accumulation throughout their lifespan. The reduced fat mass in Usp19−/− mice was reflected by decreased sizes of both subcutaneous and visceral adipose depots, but not in the thermogenic brown adipose tissue, which was unchanged in the Usp19−/− mice. This decrease in the subcutaneous depot can be ascribed, in part, to decreased adipogenic capacity of progenitors (Fig. 2). Inactivation of Usp19 prevented the upregulation of key adipogenic genes, including Pparγ. A recent publication shows that USP19 is downregulated during adipogenesis in mouse embryonic fibroblasts, but does not further explore the role of USP19 in adipogenesis [19]. This defect in adipogenesis was most apparent at 2 months of age when the fat pad in the Usp19−/− mice was decreased in size but the adipocyte area was similar. At 6 months of age or later, adipocyte area was decreased in the Usp19−/− mice suggesting that they also developed a defect in fat accumulation later in life.
The reduced adipogenic capacity in Usp19−/− mice with reductions in subcutaneous fat mass and key genes regulating insulin sensitivity and glucose homeostasis (leptin and adiponectin) may have been used to predict that Usp19−/− mice would have systemic insulin resistance [20, 21]. scWAT is believed to ‘trap’ excess dietary fat, protecting visceral fat depots and peripheral tissues, such as muscle, liver and pancreas, from the excess lipid influx that promotes lipotoxicity and associated defects in insulin secretion and action [20]. However, Usp19−/− mice manifested improved glucose homeostasis. On HFD feeding, Usp19−/− mice showed improved glucose tolerance and better blood-glucose lowering capabilities when insulin was administered, consistent with enhanced insulin signalling in muscle. Additionally, hepatic glucose production from the gluconeogenic substrate pyruvate was significantly lower in the Usp19−/− mice, suggesting enhanced insulin signalling in the liver. Indeed, this was demonstrated in vivo where insulin administration in the Usp19−/− mice led to increased levels of p-Akt in muscle and liver and p-S6K in muscle, which are key nodes in insulin signalling. Thus, loss of USP19 results in positive effects on insulin sensitivity in liver and muscle that can mitigate possible negative effects arising from decreased subcutaneous fat.
The Usp19−/− mice also manifested significantly increased physical activity (Fig. 3 and ESM Fig. 2) and increased energy expenditure when fed an HFD (Fig. 3g), which also may have contributed to the positive metabolic phenotype. The reasons for these findings are unclear, but since Usp19−/− mice are whole-body USP19 deficient, we cannot exclude an effect in the brain leading to altered behaviour and increased locomotor activity. This increased activity can lead to increased oxidation of fuel substrates, thereby reducing the need for fat storage in white adipose tissue and protecting peripheral tissues from lipid accumulation and lipotoxicity. This may explain why, despite higher or equal food intake on a normal diet and HFD, respectively, Usp19−/− mice had lower fat mass depot size and lower body weight. Moreover, increased physical activity is also associated with increased insulin sensitivity [22, 23]. However, it is unlikely that the enhanced physical activity explains all the beneficial metabolic effects observed in the Usp19−/− mice, as cell autonomous effects of USP19 have also been observed. We have previously shown that silencing Usp19 in skeletal muscle of WT mice leads to protection from muscle wasting due to alterations in protein turnover [14]. In addition, silencing or gene inactivation of Usp19 in muscle cells has been shown to promote their differentiation in vitro [11] or enhance insulin signalling [15], respectively. Moreover, in the present work, when SVF cells were isolated from fat from WT and Usp19−/− mice and cultured, we observed a significant decrease in adipogenesis and fat accumulation (Fig. 2). In the future, tissue-specific knockouts will help tease out the tissue intrinsic metabolic roles of USP19.
We did not observe any difference in Usp19 mRNA expression in human adipose tissue between two groups of participants with distinct BMI ranges. However, the human samples used in this study came from participants who have high BMI, who were subdivided into high (35–40 kg/m2) and very high (60–69 kg/m2) BMI categories. It remains to be seen if USP19 expression would be correlated with BMI if samples from a complete range of BMIs were available. We did find that USP19 mRNA expression was positively correlated with some important adipocyte genes, including PPARg, FABP4 and ADIPOQ. Intriguingly, this correlation was most evident in the omental and mesenteric adipose tissue depots. Studies indicate that the abdominal adipose tissue depots (omental and mesenteric) are more associated with insulin resistance and metabolic dysfunction during obesity [24,25,26]. This suggests that higher USP19 expression results in more adipogenic adipose tissue, concordant with the results we have observed in the mouse, where loss of USP19 resulted in fewer and smaller adipocytes and increased whole body insulin sensitivity.
Other deubiquitinating enzymes have been implicated in adipogenesis. Ubiquitin-specific peptidase 7 (USP7) regulates the ubiquitination of the histone acetyltransferase TIP60, which acts as a positive regulator of PPARγ transcriptional activity in adipogenesis [27]. In addition, pre-adipocytes and mouse embryonic fibroblast cells isolated from Uchl3−/− mice have an impaired capacity for adipogenesis [28]. Although Uchl3−/− mice manifest decreased visceral and subcutaneous fat mass and resistance to obesity on high-fat feeding (as with our Usp19−/− mice), there was no difference in glucose tolerance, only a small effect on insulin sensitivity upon high-fat feeding and markedly decreased muscle mass [29]. This contrasts with the markedly improved glucose homeostasis, positive effects on muscle mass and insulin signalling in muscle and liver in Usp19−/− mice reported here, in spite of a decreased subcutaneous fat mass. The direct substrate(s) that USP19 acts on to exert these metabolic processes are still unknown. Identification of these substrates could lead to the development of novel inhibitors of USP19 with therapeutic potential to target several metabolic tissues and, thereby, provide protection from the deleterious metabolic consequences of obesity, including diabetes and its complications.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its supplementary information files.
Abbreviations
- eWAT:
-
Epididymal white adipose tissue
- GTT:
-
Glucose tolerance test
- HFD:
-
High-fat diet
- ITT:
-
Insulin tolerance test
- IUCPQ:
-
Institut universitaire de cardiologie et de pneumologie de Québec
- MRI:
-
Magnetic resonance imaging
- PPARγ:
-
Peroxisome proliferator-activated receptor γ
- PTT:
-
Pyruvate tolerance tests
- scWAT:
-
Subcutaneous white adipose tissue
- SVF:
-
Stromal vascular fraction
- USP19:
-
Ubiquitin-specific peptidase 19
- WT:
-
Wild-type
References
Malik VS, Willett WC, Hu FB (2013) Global obesity: trends, risk factors and policy implications. Nat Rev Endocrinol 9(1):13–27. https://doi.org/10.1038/nrendo.2012.199
Varshavsky A (2012) The ubiquitin system, an immense realm. Annu Rev Biochem 81(1):167–176. https://doi.org/10.1146/annurev-biochem-051910-094049
Coyne ES, Wing SS (2016) The business of deubiquitination - location, location, location. F1000Res 5:163
Clague MJ, Barsukov I, Coulson JM, Liu H, Rigden DJ, Urbe S (2013) Deubiquitylases from genes to organism. Physiol Rev 93(3):1289–1315. https://doi.org/10.1152/physrev.00002.2013
Jin S, Tian S, Chen Y et al (2016) USP19 modulates autophagy and antiviral immune responses by deubiquitinating Beclin-1. EMBO J 35(8):866–880. https://doi.org/10.15252/embj.201593596
Cui J, Jin S, Wang RF (2016) The BECN1-USP19 axis plays a role in the crosstalk between autophagy and antiviral immune responses. Autophagy 12(7):1210–1211. https://doi.org/10.1080/15548627.2016.1173801
Hassink GC, Zhao B, Sompallae R et al (2009) The ER-resident ubiquitin-specific protease 19 participates in the UPR and rescues ERAD substrates. EMBO Rep 10(7):755–761. https://doi.org/10.1038/embor.2009.69
Lee JG, Takahama S, Zhang G, Tomarev SI, Ye Y (2016) Unconventional secretion of misfolded proteins promotes adaptation to proteasome dysfunction in mammalian cells. Nat Cell Biol 18(7):765–776. https://doi.org/10.1038/ncb3372
Lu Y, Bedard N, Chevalier S, Wing SS (2011) Identification of distinctive patterns of USP19-mediated growth regulation in normal and malignant cells. PLoS One 6(1):e15936. https://doi.org/10.1371/journal.pone.0015936
Altun M, Zhao B, Velasco K et al (2012) Ubiquitin-specific protease 19 (USP19) regulates hypoxia-inducible factor 1alpha (HIF-1alpha) during hypoxia. J Biol Chem 287(3):1962–1969. https://doi.org/10.1074/jbc.M111.305615
Wiles B, Miao M, Coyne E, Larose L, Cybulsky AV, Wing SS (2015) USP19 deubiquitinating enzyme inhibits muscle cell differentiation by suppressing unfolded-protein response signaling. Mol Biol Cell 26(5):913–923. https://doi.org/10.1091/mbc.e14-06-1129
Ogawa M, Kitakaze T, Harada N, Yamaji R (2015) Female-specific regulation of skeletal muscle mass by USP19 in young mice. J Endocrinol 225(3):135–145. https://doi.org/10.1530/JOE-15-0128
Combaret L, Adegoke OA, Bedard N, Baracos V, Attaix D, Wing SS (2005) USP19 is a ubiquitin-specific protease regulated in rat skeletal muscle during catabolic states. Am J Physiol Endocrinol Metab 288(4):E693–E700. https://doi.org/10.1152/ajpendo.00281.2004
Bedard N, Jammoul S, Moore T et al (2015) Inactivation of the ubiquitin-specific protease 19 deubiquitinating enzyme protects against muscle wasting. FASEB J 29(9):3889–3898. https://doi.org/10.1096/fj.15-270579
Coyne ES, Bedard N, Wykes L et al (2018) Knockout of USP19 deubiquitinating enzyme prevents muscle wasting by modulating insulin and glucocorticoid signaling. Endocrinology 159(8):2966–2977. https://doi.org/10.1210/en.2018-00290
Galarraga M, Campion J, Munoz-Barrutia A et al (2012) Adiposoft: automated software for the analysis of white adipose tissue cellularity in histological sections. J Lipid Res 53(12):2791–2796. https://doi.org/10.1194/jlr.D023788
Rosen ED, Walkey CJ, Puigserver P, Spiegelman BM (2000) Transcriptional regulation of adipogenesis. Genes Dev 14(11):1293–1307
Tschop MH, Speakman JR, Arch JR et al (2011) A guide to analysis of mouse energy metabolism. Nat Methods 9(1):57–63. https://doi.org/10.1038/nmeth.1806
Lim KH, Choi JH, Park JH et al (2016) Ubiquitin specific protease 19 involved in transcriptional repression of retinoic acid receptor by stabilizing CORO2A. Oncotarget 7(23):34759–34772. https://doi.org/10.18632/oncotarget.8976
Faraj M, Lu HL, Cianflone K (2004) Diabetes, lipids, and adipocyte secretagogues. Biochem Cell Biol 82(1):170–190. https://doi.org/10.1139/o03-078
Galic S, Oakhill JS, Steinberg GR (2010) Adipose tissue as an endocrine organ. Mol Cell Endocrinol 316(2):129–139. https://doi.org/10.1016/j.mce.2009.08.018
Balkau B, Mhamdi L, Oppert JM et al (2008) Physical activity and insulin sensitivity: the RISC study. Diabetes 57(10):2613–2618. https://doi.org/10.2337/db07-1605
Richter EA, Garetto LP, Goodman MN, Ruderman NB (1982) Muscle glucose metabolism following exercise in the rat: increased sensitivity to insulin. J Clin Invest 69(4):785–793. https://doi.org/10.1172/JCI110517
Preis SR, Massaro JM, Robins SJ et al (2010) Abdominal subcutaneous and visceral adipose tissue and insulin resistance in the Framingham heart study. Obesity (Silver Spring) 18(11):2191–2198. https://doi.org/10.1038/oby.2010.59
Wajchenberg BL, Giannella-Neto D, da Silva ME, Santos RF (2002) Depot-specific hormonal characteristics of subcutaneous and visceral adipose tissue and their relation to the metabolic syndrome. Horm Metab Res 34(11/12):616–621. https://doi.org/10.1055/s-2002-38256
Tchernof A, Despres JP (2013) Pathophysiology of human visceral obesity: an update. Physiol Rev 93(1):359–404. https://doi.org/10.1152/physrev.00033.2011
Gao Y, Koppen A, Rakhshandehroo M et al (2013) Early adipogenesis is regulated through USP7-mediated deubiquitination of the histone acetyltransferase TIP60. Nat Commun 4(1):2656. https://doi.org/10.1038/ncomms3656
Suzuki M, Setsuie R, Wada K (2009) Ubiquitin carboxyl-terminal hydrolase l3 promotes insulin signaling and adipogenesis. Endocrinology 150(12):5230–5239. https://doi.org/10.1210/en.2009-0332
Setsuie R, Suzuki M, Kabuta T et al (2009) Ubiquitin C-terminal hydrolase-L3-knockout mice are resistant to diet-induced obesity and show increased activation of AMP-activated protein kinase in skeletal muscle. FASEB J 23(12):4148–4157. https://doi.org/10.1096/fj.09-132217
Acknowledgements
The authors thank the team at the IUPCQ tissue bank for providing the human adipose tissue samples and M. Kokoeva (Department of Medicine, McGill University Health Centre, Montréal, QC, Canada) for the metabolic studies. The authors also thank M. Plourde (Department of Medicine, McGill University, Montréal, QC, Canada) for excellent technical assistance.
Funding
This work was supported by grants from the Canadian Institutes of Health Research (SSW, MOP 82734) and from the Canadian Cancer Society Research Institute Innovation Grant (SSW no. 703394).
Author information
Authors and Affiliations
Contributions
ESC, MF, AT and SSW conceptualised and designed the studies. ESC, NB, YJG and SSW executed the experiments and analysed the data. ESC and SSW wrote the original draft. ESC, NB, YJG, MF, AT, SSW edited and revised the manuscript. ESC, NB, YJG, MF, AT, SSW approved the final version of this manuscript. SSW is the guarantor of this work.
Corresponding author
Ethics declarations
SSW receives funding from ALMAC Discovery for work on USP19 that is not related to the studies presented in this manuscript. AT receives research funding from Johnson & Johnson and Medtronic for studies unrelated to this manuscript.
Electronic supplementary material
ESM
(PDF 1114 kb)
Rights and permissions
About this article
Cite this article
Coyne, E.S., Bédard, N., Gong, Y.J. et al. The deubiquitinating enzyme USP19 modulates adipogenesis and potentiates high-fat-diet-induced obesity and glucose intolerance in mice. Diabetologia 62, 136–146 (2019). https://doi.org/10.1007/s00125-018-4754-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-018-4754-4