Abstract
Aims/hypothesis
Evidence continues to emerge detailing a fine-tuning of the regulation of metabolic processes and energy homeostasis by cell-autonomous circadian clocks. Pancreatic beta cell functional maturation occurs after birth and implies transcriptional changes triggered by a shift in the nutritional supply that occurs at weaning, enabling the adaptation of insulin secretion. So far, the developmental timing and exact mechanisms involved in the initiation of the circadian clock in the growing pancreatic islets have never been addressed.
Methods
Circadian gene expression was measured by quantitative RT-PCR in islets of rats at different postnatal ages up to 3 months, and by in vitro bioluminescence recording in newborn (10-day-old) and adult (3-month-old) islets. The effect of the microRNAs miR-17-5p and miR-29b-3p on the expression of target circadian genes was assessed in newborn rat islets transfected with microRNA antisense or mimic oligonucleotides, and luciferase reporter assays were performed on the rat insulin-secreting cell line INS832/13 to determine a direct effect. The global regulatory network between microRNAs and circadian genes was computationally predicted.
Results
We found up to a sixfold-change in the 24 h transcriptional oscillations and overall expression of Clock, Npas2, Bmal1, Bmal2, Rev-erbα, Per1, Per2, Per3 and Cry2 between newborn and adult rat islets. Synchronisation of the clock machinery in cultured islet cells revealed a delayed cell-autonomous rhythmicity of about 1.5 h in newborn compared with adult rats. Computational predictions unveiled the existence of a complex regulatory network linking over 40 microRNAs displaying modifications in their expression profiles during postnatal beta cell maturation and key core-clock genes. In agreement with these computational predictions, we demonstrated that miR-17-5p and miR-29b-3p directly regulated circadian gene expression in the maturing islet cells of 10-day-old rats.
Conclusions/interpretation
These data show that the circadian clock is not fully operational in newborn islets and that microRNAs potently contribute to its regulation during postnatal beta cell maturation. Defects in this process may have long-term consequences on circadian physiology and pancreatic islet function, favouring the manifestation of metabolic diseases such as diabetes.
Similar content being viewed by others
Introduction
Early postnatal life is a critical period for the development of the endocrine pancreas and involves the maturation of secretory and mitogenic processes. We and others have demonstrated that newborn beta cells are still immature immediately after birth [1,2,3]. They lack the unique and essential feature of mature beta cells: the capacity to secrete insulin in response to elevated glucose concentrations. In contrast, amino acid-stimulated insulin release and insulin biosynthesis are already fully operational. At the same time, newborn beta cells proliferate intensively within the expanding pancreas to achieve an appropriate beta cell mass that will then be maintained throughout adult life. We found that beta cell maturation is associated with a major reprogramming in gene expression. This process is at least in part governed by microRNAs (miRNAs), a class of small non-coding RNAs that inhibit the translation and/or stability of target mRNAs by pairing to their 3′ untranslated region (UTR) [1]. Indeed, we demonstrated that modifications in islet miRNA levels that take place during the postnatal period directly control the expression of key metabolic and cell cycle genes, driving the maturation of glucose-responsive adult beta cells.
Rodent and human pancreatic beta cells, like most cells in the body, possess self-sustained molecular clocks that coordinate the timing of metabolism throughout the day and modulate insulin secretion to maintain blood glucose homeostasis [4,5,6,7]. The circadian oscillators operative in beta cells and other peripheral cells are synchronised by systemic signals (e.g. neural and endocrine signals, fasting/feeding and metabolic cycles) controlled by a central pacemaker in the suprachiasmatic nucleus of the brain. This master clock, itself synchronised by daylight, maintains phase coherence among the body’s cellular clocks [8,9,10]. The clock machinery has a strong influence on gene expression programmes and orchestrates the timing of metabolic processes by regulating in a circadian manner about 10% of the transcripts [11, 12]. Cell-autonomous feedback loops repeated every 24 h are generated by the activation of the transcription factors circadian locomotor output cycles kaput (CLOCK) and brain and muscle ARNT-like 1 (BMAL1; also known as aryl hydrocarbon receptor nuclear translocator-like protein [ARNTL]) that trigger the expression of their own repressors, periods (PER1, 2 and 3) and cryptochromes (CRY1 and 2). CLOCK and BMAL1 also drive the rhythmic expression of the nuclear receptor subfamily 1 group D genes Rev-erbα (also known as Nr1d1) and Rev-erbβ (also known as Nr1d2) [9]. Moreover, the expression of clock genes is regulated at the post-transcriptional level by many other factors, including miRNAs [13].
Studies of rhythmic gene expression based on the disruption of clock genes in liver, pancreas, muscle and adipose tissue suggest that peripheral clocks may play an important role in optimising metabolism by coordinating the function of key metabolic organs to nutrient availability and energy requirements [14,15,16,17]. Importantly, previous studies in humans and rodents favour the view that the synchronised rhythmicity of clock gene expression matures at rates that differ between tissues and is the result of specific changes occurring in the environmental conditions of each organ [18,19,20,21]. Indeed, in humans, plasma circadian variations of the glucocorticoid hormone appear after 3 months of age [22], which induces changes in the circadian gene expression phases in the liver, kidney and heart [23]. Consistently, a gradual completion of clock gene rhythmic expression during postnatal life has been reported in rat liver within 30 days after birth [24].
However, the establishment of circadian rhythms during pancreatic beta cell development and its underlying mechanisms have not previously been investigated. Hence, in this study, we determined the expression levels and circadian oscillation profiles of several core-clock genes in maturating beta cells. Moreover, we assessed the role of nutrition-driven islet miRNA changes in modulating circadian clockwork during postnatal development. We investigated the expression and rhythmic oscillations of circadian genes in newborn rat islets, and analysed expression of that miR-17-5p and miR-29b-3p throughout beta cell maturation, in relation to islet clock gene expression.
Methods
Animals
Male and pregnant female Sprague–Dawley rats were obtained from Janvier Laboratories (Le Genest-Saint-Isle, France) and housed under a 12 h/12 h light/dark cycle in a climate-controlled and pathogen-free facility. All procedures were performed in accordance with the Guide for the Care and Use of Laboratory Animals from the National Institutes of Health and were approved by the Swiss Research Councils and Veterinary Office (authorisation VD2608.1). At birth, litters were adjusted to 10–12 pups per dam to standardise the mothers’ milk availability. Newborn males and females were nursed until they were euthanised. At postnatal day (P)21, the pups were weaned on a standard chow diet containing 18.5% crude proteins, 4.5% fat and 54% carbohydrates.
Cell culture, transfection and lentiviral transduction
Pancreatic islets were isolated as described [25] by collagenase digestion followed by purification on Histopaque density gradients (Sigma-Aldrich, St Louis, MO, USA). The islets were cultured in RPMI 1640 Glutamax medium (Invitrogen, Carslbad, CA, USA) supplemented with 10% FCS (Amimed, London, UK), 50 U/ml penicillin, 50 μg/ml streptomycin, 1 mmol/l sodium pyruvate and 250 μmol/l HEPES. The rat insulin-secreting cell line INS832/13 [26], free from mycoplasma contamination, was cultured as described [27]. INS832/13 cells and dissociated rat islet cells were transfected using Lipofectamine 2000 (Invitrogen) [27]. To reduce miR-17-5p expression, approximately 250,000 cells were transfected with a specific single-stranded anti-miR (miScript miRNA inhibitor; Qiagen, Hilden, Germany), or with the miScript miRNA reference inhibitor (Qiagen) as a negative control. Overexpression of miR-17-5p and miR-29b-3p was achieved by transfecting cells with oligonucleotide duplexes (Eurogentec, Liege, Belgium; and Qiagen) using Lipofectamine 2000 (Invitrogen). A small interfering RNA (siRNA) directed against green fluorescent protein (GFP) was used as a negative control. The survival rate was over 90% in transfected cells. To monitor circadian bioluminescence, a Bmal1–luciferase (Bmal1–luc; Bmal1 is also known as Arntl) construct [28] was introduced into approximately 150,000 adherent cells by lentiviral transduction. To produce lentiviral particles, Bmal1–luc lentivectors were transfected into 293 T cells using the polyethylenimine method [5, 29]. Dissociated rat islet cells were transduced twice, with a multiplicity of infection of three for each transduction.
Measurement of miRNA and mRNA expression
Total RNA was extracted with the miRNeasy kit (Qiagen). Mature miRNA and mRNA levels were assessed by quantitative RT-PCR (qRT-PCR) using the miScript II RT kit (Qiagen). miScript primer assays and primer sequences are provided in electronic supplementary material (ESM) Table 1. miRNA expression was normalised to the level of U6. mRNA expression was normalised to the amount of 18S. Pools of pancreatic islets from P1 (four pups), P5 (four pups) and P10 (two pups) were collected, while islets from P15, P20, P23, P31 and adult rats were extracted individually.
Luciferase assays
Luciferase reporter plasmids were generated by cloning about 200 nucleotides of the 3′ UTR of rat Clock, Npas2 (encoding neuronal PAS domain protein 2) and Per3 containing the putative binding sites of miR-17-5p or miR-29b-3p into the psiCHECK-1 vector (Promega, Madison, WI, USA). Luciferase activity was measured in around 180,000 INS832/13 cells with a dual luciferase reporter assay (Promega) 2 days after transfection. Renilla luciferase activity from the psiCHECK-1 constructs was normalised for transfection efficiency to the SV40-driven Firefly activity generated from a co-transfected PGL3 promoter vector (Promega).
In vitro synchronisation and circadian bioluminescence measurement
Adherent transduced islet cells were synchronised by a 1 h forskolin pulse (10 μmol/l; Sigma-Aldrich), and subjected to continuous bioluminescence recording with lumicycle equipment (made in house) in RPMI medium containing 100 μmol/l luciferin (d-luciferin 306–250; NanoLight Technology, Pinetop, AZ, USA) as previously described [5]. Photon counts from each well were integrated during 1 min over 24 min intervals. For detrended time series, raw luminescence signals were smoothened by a moving average with a window of 24 h, as previously described [7]. Average circadian amplitude and period length were calculated based on five consecutive peaks of detrended profiles starting from the second circadian cycle. The average timing of the positive peak phase was quantified starting from the second peak, relative to the 24 h day cycle [30].
Modelling of the core-clock mRNA–miRNA network during beta cell maturation
We selected 62 miRNAs that are differentially expressed between newborn and adult rats [1]. For each miRNA of interest, we retrieved the target genes from several sources (DIANA-microT, ElMMo, miRBase, miRanda, miRDB, PicTar, PITA and TargetScan) using the MultiMir R package (http://multimir.ucdenver.edu; accessed Jan 2017) [31]. The retrieved interactions were filtered using the list of core-clock genes investigated in this study. The resulting network was visualised with Cytoscape 3.0 (www.cytoscape.org/cy3.html) [32]. The information about the expression changes between the islets of newborn and adult rats mapped on the network was inferred from the microarray data for the miRNAs [1] and the qRT-PCR data (present study) for the core-clock genes. The colour of the edges of the network indicated the sign of the log fold change of each connected node. Anti-correlated interactions, suggesting a functional link between the miRNA and the mRNA, were highlighted.
Statistical analysis
Statistical differences were calculated using a Student’s t test for unpaired comparisons, or ANOVA followed by a post hoc Dunnett test for multiple comparisons, with a discriminating p value of 0.05 (GraphPad Prism, La Jolla, CA, USA). The amplitude differences between the consecutive circadian cycles in each profile were assessed using one-way ANOVA.
Results
Rhythmic transcriptional oscillation of circadian genes is not yet established in newborn islets
After birth, rat pancreatic beta cells are unable to secrete insulin specifically in response to glucose and undergo a postnatal maturation process resulting in the acquisition of a functional adult beta cell mass [1]. Since islets possess self-sustained molecular clocks that optimise cellular functions, including insulin secretion, we determined whether the islet circadian rhythmicity and its machinery differed between newborn and adult islets. For this purpose, we assessed the mRNA levels of several core circadian genes in newborn and adult rat islets across 24 h (Fig. 1). Interestingly, we found that the transcriptional oscillations of Npas2, Bmal1, Rev-erbα, Per1, Per2, Per3 and Cry2 over 24 h were strikingly attenuated in P10 compared with 3-month-old adult islets (Fig. 1b, c, e–h, j). Data are expressed as means ± SD.
Rhythmic expression profile of genes involved in the circadian clock in newborn and adult islets. The islets of 10-day-old (black circles) and adult (white circles) rats were isolated and the RNA extracted at 4 h intervals across the 12 h/12 h light/dark cycle. Transcript expression was determined by qRT-PCR. Fold-change values are displayed as relative abundance to that of 18S amplified within the same sample. Two-way ANOVA was used to determine variance with respect to time and group (mean ± SD, n = 4 rats per group per time point, *p < 0.05 between groups)
These findings can be explained either by an intrinsic incapacity of the genes to oscillate in neonatal islets or by the lack of synchronisation between newborn cells. To investigate the cell-autonomous properties of the circadian clocks in newborn islets, the oscillation profile of the bioluminescent circadian reporter Bmal1–luc was recorded in cultured P10 and adult islet cells, following in vitro synchronisation. When synchronised by forskolin, both P10 and adult islet cells exhibited self-sustained circadian oscillations of Bmal1–luc expression (Fig. 2a, b), but the characteristics of these profiles were distinct (Fig. 2c–e). Newborn islet cells displayed an earlier first peak (Fig. 2c; 5.51 ± 1.31 h in P10 islet cells compared with 10.10 ± 0.74 h in adult cells) and a circadian phase delay of almost 4 h (Fig. 2d; 12.68 ± 1.37 h in P10 vs 8.79 ± 0.64 h in adults). Moreover, circadian period length was about 1.5 h longer in P10 islet cells (25.08 ± 0.16 h vs 23.81 ± 0.22 h in adults; Fig. 2e). The average amplitude of Bmal1–luc oscillations in P10 islet cells tended to be lower compared with their adult counterparts, but this did not reach statistical significance (Fig. 2f; 0.19 ± 0.03 h vs 0.27 ± 0.046, p = 0.248). However, desynchronisation was faster in P10 than adult islet cells, as shown by a decreased circadian amplitude starting from the fifth cycle compared with the second cycle in P10 cells (Fig. 2g; p = 0.0061). Taken together, these data suggest that although the rhythmic oscillations of Bmal1–luc reporter were observed in P10 islet cells synchronised in vitro, their properties differed significantly from those of their adult counterparts.
Newborn and adult islet cells synchronised in vitro exhibit distinct circadian oscillation profiles. Pancreatic islets were isolated from P10 and 3-month-old adult rats. Islet cells were transduced with lentiviral particles expressing Bmal1–luc, synchronised by a forskolin pulse, and bioluminescence was recorded. (a) Raw data profiles (counts/min) from a parallel recording of islet cells derived from P10 (red line) and adult (blue line) animals. (b) Average detrended bioluminescence profiles for n = 3 independent experiments (one pooled litter of 10–12 pups per experiment) for P10 islet cells (red line), and n = 3 independent experiments (one animal per experiment) for adult islet cells (blue line). (c) Zenith (peak) time for the first peak of Bmal1–luc detrended bioluminescence (b) following forskolin synchronisation. (d) Average circadian phase of three consecutive peaks starting from the second peak and normalised to a 24 h day period. (e) Circadian period length. (f) Circadian amplitude, averaged for all the cycles starting from the second cycle, or (g) presented individually for each cycle (black bars, P10 islet cells; white bars, adult cells). Data in (c–g) are expressed as mean ± SEM, n = 3 independent experiments (one pooled litter of 10–12 pups per experiment) for P10 islet cells, and n = 3 independent experiments (one animal per experiment) for adult islet cells. *p < 0.05 unpaired, two-tailed Student’s t test. (g) One-way ANOVA was applied to the P10 and adult graphs separately, indicating a significant (p = 0.0061) decrease in amplitude in P10, but not in adult (p = 0.5390), islet cells
Islet circadian genes are differentially expressed during postnatal maturation
As beta cell postnatal maturation involves drastic changes in the expression of numerous islet genes, mainly involved in stimulus–secretion coupling and cell cycling [1], we determined whether the expression of circadian genes also differed between newborn and adult islets. qRT-PCR measurements revealed that half of the tested genes displayed different levels in adult compared with immature newborn islets (Fig. 3a). The expression of Clock, Npas2, Bmal1 and Bmal2 were higher in fully mature islet cells, whereas the levels of Rev-erbα and Per3 were lower. We next examined the kinetics of the changes in expression of these circadian genes by measuring their levels at different time points during postnatal life. We observed that immediately after birth the expression profiles were stable, becoming modified after the third postnatal week (Fig. 3b–g), i.e. at the time the islet cells became fully functional. Indeed, we previously demonstrated that the nutritional shift occurring at weaning (postnatal day 21) leads to the acquisition of a mature insulin secretory phenotype. This adaptation ensures the release of appropriate amounts of insulin in response to an increased carbohydrate dietary intake [1]. These data reveal that the adult circadian gene expression profile in islets is established once beta cell maturation has occurred.
Modifications of islet clock gene expression throughout postnatal life. (a) qRT-PCR was performed using islet RNAs from 10-day-old vs 3-month-old rats isolated between 8:00 and 10:00 hours. The expression of the indicated genes in adult islets is presented as the ratio to the level measured in newborn islets (dotted line) and represent the mean ± SD of four rats per group. *p < 0.05 vs newborn rats, ANOVA. (b–g) qRT-PCR was performed using samples from rats at the indicated ages. Data are mean ± SD (n = 4 per group). Statistical difference from 10-day-old rats was assessed by one-way ANOVA with a Dunnett post hoc test: *p˂0.05. Npas2 expression: p = 0.06 between P10 and adult islets
miRNAs regulate circadian gene expression
Nutrient-induced postnatal miRNA modifications promote beta cell maturation by regulating the expression of metabolic and cell cycle genes [1]. To further determine the contribution of miRNAs to the regulation of islet circadian genes in postnatal life, we used the computational algorithms miRSystem and TargetScan to search for miRNAs targeting the 3′ UTR of circadian mRNAs. We included in our computational analysis 17 miRNAs that we had previously investigated for their role in beta cell maturation [1]. The compiled results revealed that the predicted targets of miR-17-5p and miR-29b-3p were enriched in genes belonging to the circadian machinery. To experimentally verify the computational predictions, we mimicked in islet cells obtained from 10-day-old rats the reduction of miR-17-5p (ESM Fig. 1a) and the upregulation of miR-29-3p (ESM Fig. 1b) that occurred during the postnatal maturation of beta cells. The decrease in miR-17-5p resulted in an increase of Clock and Npas2 mRNA levels (Fig. 4a, b). Dual luciferase reporter assays in the rat beta cell line INS832/13 confirmed that miR-17-5p directly targeted the 3′ UTR of Clock but not Npas2 (Fig. 4c; ESM Fig. 1c). Thus, the effect of miR-17-5p on Npas2 expression is most likely to be indirect. As predicted by the computational programs, the increase in miR-29b-3p in newborn islet cells resulted in a reduction in Per3 expression (Fig. 4d) and suppressed the luciferase activity produced from the Per3–3′ UTR construct (Fig. 4e; ESM Fig. 1d). These findings confirm that miR-29b-3p directly inhibited the expression of Per3, the antagonistic regulator of the CLOCK–BMAL1 heterodimer complex. We then assessed whether the expression of miR-17-5p and miR-29b-3p followed an oscillatory pattern across 24 h. As expected, the levels of miR-17-5p were lower and those of miR-29b-3p higher at all time points in adult islets [1]. However, none of the miRNAs displayed a circadian oscillatory pattern (Fig. 4f, g).
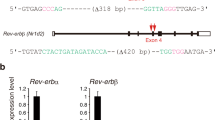
miR-17-5p and miR-29b-3p regulation of clock genes and oscillation profiles. (a, b, d) Dispersed 10-day-old rat islet cells were transfected with anti-miRNAs to inhibit (a, b), or with oligonucleotide mimics to overexpress (d), the miRNAs. An miRNA reference inhibitor (Qiagen) and an siRNA against GFP were used respectively as negative controls (ctrl). mRNA levels assessed by qRT-PCR are shown as fold changes (n = 4–6). Data are mean ± SD. *p˂0.05, Student’s t test. (c, e) Direct interaction of miR-17-5p (c) and miR-29b-3p (e) with their putative targets was assessed by luciferase reporter assays in INS832/13 cells. Vectors containing the 3′ UTR of Clock, Npas2 and Per3 were co-transfected with either a control oligonucleotide or oligonucleotides mimicking the miRNAs of interest. An empty vector was used as the control. Data are mean ± SD (n = 4), and statistical significance was calculated by ANOVA. (f, g) Level of miR-17-5p and miR-29b-3p in 10-day-old (black circles) and adult (white circles) rat islets isolated at 4 h intervals across the 12 h/12 h light/dark cycle was determined by qRT-PCR. The data (mean ± SD, n = 4 per group per time point; *p < 0.05 between groups) are displayed as abundance relative to that of U6 amplified within the same sample
Besides miR-17-5p and miR-29b-3p, several other miRNAs displaying expression changes during postnatal beta cell maturation are predicted to target at least one core-clock gene. To obtain a comprehensive picture of the potential impact of miRNAs on the expression of core-clock components, we used a computational approach to search for all the predicted interactions between the 62 miRNAs displaying expression changes during the functional maturation of beta cells [1] and the 12 clock genes investigated in this study (Fig. 1). MiRNAs and core-clock genes were found to be connected by a complex regulatory network (Fig. 5), with more than 40 miRNAs showing anti-correlated expression changes with their respective targets. As expected, most miRNAs were predicted to regulate the level of multiple clock genes, and several clock genes were found to be targeted by multiple miRNAs. Of note, 20 different miRNAs displaying reduced expression changes upon weaning were found to potentially contribute to the observed rise in the level of Clock in mature beta cells (Fig. 5). These findings indicate that the islet circadian gene expression profile is probably not determined by a single miRNA but results from the cooperative action of multiple miRNAs involved in the functional maturation of beta cells.
Core-clock mRNA–miRNA network during beta cell maturation. Potential interactions between miRNAs displaying expression changes during postnatal beta cell maturation and circadian clock genes were computationally predicted as described in the experimental procedures. Green and red symbols indicate up- and downregulation (log2) between adult and newborn rat islets, respectively. Opposite expression changes between the miRNAs and their putative targets suggestive of functional interactions are depicted with thick blue lines, while changes in the same direction are shown with pink lines. Clock genes are shown with square symbols, while miRNAs are presented as diamond symbols
Discussion
Emerging evidence indicates that the cell-autonomous circadian clocks operating in the endocrine pancreas play an essential role in coordinating insulin secretion and islet cell transcriptome in a cyclic-dependent manner [5, 6, 33,34,35,36]. However, the rhythmicity and expression profiles of clock genes in pancreatic islets and the mechanisms regulating them throughout postnatal beta cell maturation have so far not been investigated. The present study highlights drastic differences in the expression and temporal profiles of the core-clock genes between 10-day-old and 3-month-old rat islets. Importantly, we found that miR-17-5p and miR-29b-3p directly regulated the core circadian genes Clock and Per3 throughout postnatal islet maturation, and are likely to be involved in establishing a functional autonomous molecular clock. Our data suggest that the regulatory network connecting miRNAs and clock genes is most probably far more complex, and the establishment of a fully operational circadian clock is unlikely to be explained by single miRNA–target associations. Indeed, several other miRNAs involved in the functional maturation of beta cells are predicted to regulate at least one of the core-clock genes and display anti-correlated expression changes with their putative targets. Additional studies will be needed to experimentally verify these computational predictions and to precisely delineate the interconnection between miRNAs and the circadian clock in the context of postnatal beta cell maturation.
We demonstrated that the circadian expression patterns of clock components are not fully established in immature newborn islets. Of note, although in vivo circadian oscillations in newborn islet cells were strongly dampened, oscillations of the circadian reporter could still be induced in vitro in P10 islet cells by forskolin synchronisation (compare Fig. 1 with Fig. 2). These findings suggest that the lack of oscillations observed in newborns is a consequence of the desynchronisation of the cells rather than an intrinsic incapacity of clock genes to undergo circadian expression changes. However, the oscillatory profiles observed in vitro have distinct characteristics in P10 cells compared with their adult counterparts. These changes comprised the phase advance of the first circadian peak (Fig. 2c), which may suggest an altered immediate resetting response. The differences between P10 and adult islet cell clocks were not limited to the kinetics of the first circadian cycle, but were extended to the circadian cycle characteristics such as phase, amplitude and period length (Fig. 2d–f), including a faster decrease in the amplitude of the P10 oscillations, which may suggest more rapid desynchronisation for these cells following the initial pulse. In agreement with our observations, the reported phase shift of the circadian oscillator Per1 in rat liver, thyroid and pineal gland throughout development [19] may highlight a ubiquitous delay of the clock gene oscillations in the maturating peripheral tissues after birth. Despite the fact the circadian clock machinery is already expressed at a cellular level at very early developmental stages—although initially only at low basal levels—it was proposed that circadian rhythm ontogenesis is strongly influenced by changes in the environmental conditions and in organ function occurring during development [19, 37, 38]. It is well established that both mature beta and alpha cells display cell-autonomous circadian rhythms of core-clock and functional metabolic genes [6, 7, 39]. Consistently, we observed that forskolin-induced oscillations of clock genes are elicited ex vivo despite the absence of in vivo circadian rhythms in newborn islets. These data were obtained in whole islets. Hence, these changes may occur and have an effect on one or more islet cell types. Nevertheless, beta cells constitute the main cell type in both P10 and adult islets, suggesting that modifications in the circadian gene oscillation profiles and overall mRNA levels contribute to beta cell maturation. To further elucidate the mechanism of beta cell clock maturation, in vitro experiments implying single-cell analysis of P10 and adult islet circadian bioluminescence profiles will be required.
Taken together, our results suggest that the initiation of islet circadian rhythms might rely on environmental factors, comprising blood-borne signals. Although further investigations are needed to determine the potential involvement of hormonal factors and neural inputs in the establishment of circadian rhythms, our study reveals that miRNAs are part of the mechanisms contributing to the post-transcriptional control of clock genes in immature islets. We have shown previously that miRNA modifications are driven by the shift in food supply occurring at weaning when high-fat maternal milk is replaced by a carbohydrate-enriched diet. In view of this, it is tempting to speculate that the nutritional switch is responsible not only for islet miRNA changes and the subsequent functional beta cell maturation, but also for adaptive circadian clock ontogenesis. Indeed, the attenuated clock gene expression and phase in the islets during early postnatal life suggest that the circadian regulation of insulin secretion is not yet necessary for maintaining blood glucose homeostasis as newborn animals fed a low-carbohydrate diet are normoglycaemic. Accordingly, disruption of the circadian rhythmicity of clock genes induces beta cell dysfunction, impairment of glucose metabolism and the release of inappropriate amounts of insulin [33, 36]. Before weaning, the low glucose responsiveness of newborns is thought to be the consequence of reduced glucose oxidation and metabolite production leading to an imbalanced NADPH/NADP ratio [40]. NADPH and the subsequent activation of essential transcription factors is a strong regulator of clock gene activation and has the potential to phase-shift circadian rhythms [41, 42]. This supports our hypothesis that circadian clock ontogenesis is rather needed once the beta cells have to face the rise in insulin needs required to match the increased carbohydrate supply, and hence the islet clock is required to coordinate beta cell function with circadian energy homeostasis.
Our study also reveals that the expression of core-clock genes in postnatal rat islets reaches levels similar to those of adults within 30 days of age. The differential rhythmicity and expression of clock genes observed in immature compared with adult islets have been suggested to apply also to other metabolic organs in early life. Indeed, previous studies have reported a gradual development of the circadian clock during the postnatal period, with temporal differences in the appearance of clock amplitude oscillations between the central nervous system and heart in mice [43]. This raises the hypothesis that the time frame of islet clock establishment might be cell type-specific and directly related to the capacity of the beta cells to release insulin in response to nutrients, in particular to glucose, allowing the increased insulin needs caused by food transition to be met.
The islet miRNA changes taking place during postnatal beta cell maturation, which we detailed in a previous study [1], are unlikely to be dictated by the central clock pacemaker in the suprachiasmatic nucleus, which is itself activated by light exposure [44]. In contrast, miRNA modifications occur between postnatal days P20 and P31, when the transition from fat-enriched maternal milk to a high-carbohydrate diet takes place. In line with this, Sladek and colleagues showed that the amplitude of the oscillations of clock genes in rat liver are not yet established at the age of 20 days but achieve a mature state only at around P30 [24]. Similarly, the circadian expression of intestinal disaccharidase enzymes achieves the levels observed in adult rats only several days after weaning [45]. Previous findings suggest that the establishment of clock gene expression and rhythmicity in peripheral tissues such as the endocrine pancreas is entrained by the periodicity of meals, which is acquired during adulthood, rather than by light [46]. Indeed, photo-entrainment is already functional at birth [47], whereas the rhythmicity of timely regulated food intake seems to be established after the shift to solid food, reinforcing the need to achieve a regulated insulin release in a circadian manner after the transition of nutritional supplies that occurs at weaning. Food-anticipatory activities are defined as food-seeking behaviours in anticipation of mealtimes, including physiological and hormonal activations in response to daily temporal windows of feeding time [48, 49]. Notably, global and brain-specific deletions of the clock component Rev-erbα have been reported to impair the 24 h pattern of food entrainment in mice [50]. We could speculate that, in newborns, the lack of a regulated temporal pattern of food intake justifies the absence of a circadian control of beta cell secretory activity. This raises the possibility that circadian clock entrainment and feeding time dictate the already described phasic insulin release from adult islets [7].
In conclusion, we have revealed the contribution of miRNAs to circadian clock regulation in pancreatic islets during the postnatal period. The suggested plethora of known and yet-to-be-discovered signalling pathways linking the chronobiology of insulin secretion with miRNAs indicates that we have just begun to scratch the surface of understanding the interplay between the circadian clock and postnatal beta cell maturation.
Abbreviations
- BMAL:
-
Brain and muscle ARNT-like
- CLOCK:
-
Circadian locomotor output cycles kaput
- Bmal1–luc:
-
Bmal1–luciferase
- GFP:
-
Green fluorescent protein
- miRNA:
-
microRNA
- P:
-
Postnatal day
- qRT-PCR :
-
Quantitative RT-PCR
- siRNA:
-
Small interfering RNA
- UTR :
-
Untranslated region
References
Jacovetti C, Matkovich SJ, Rodriguez-Trejo A, Guay C, Regazzi R (2015) Postnatal beta-cell maturation is associated with islet-specific microRNA changes induced by nutrient shifts at weaning. Nat Commun 6:8084
Pildes RS, Hart RJ, Warrner R, Cornblath M (1969) Plasma insulin response during oral glucose tolerance tests in newborns of normal and gestational diabetic mothers. Pediatrics 44:76–83
Rozzo A, Meneghel-Rozzo T, Delakorda SL, Yang SB, Rupnik M (2009) Exocytosis of insulin: in vivo maturation of mouse endocrine pancreas. Ann N Y Acad Sci 1152:53–62
Rakshit K, Qian J, Colwell CS, Matveyenko AV (2015) The islet circadian clock: entrainment mechanisms, function and role in glucose homeostasis. Diabetes Obes Metab 17(Suppl 1):115–122
Saini C, Petrenko V, Pulimeno P et al (2016) A functional circadian clock is required for proper insulin secretion by human pancreatic islet cells. Diabetes Obes Metab 18:355–365
Perelis M, Marcheva B, Ramsey KM et al (2015) Pancreatic beta cell enhancers regulate rhythmic transcription of genes controlling insulin secretion. Science 350:aac4250
Pulimeno P, Mannic T, Sage D et al (2013) Autonomous and self-sustained circadian oscillators displayed in human islet cells. Diabetologia 56:497–507
Schibler U, Gotic I, Saini C et al (2015) Clock-talk: interactions between central and peripheral circadian oscillators in mammals. Cold Spring Harb Symp Quant Biol 80:223–232
Asher G, Schibler U (2011) Crosstalk between components of circadian and metabolic cycles in mammals. Cell Metab 13:125–137
Dibner C, Schibler U (2015) Circadian timing of metabolism in animal models and humans. J Intern Med 277:513–527
Duffield GE (2003) DNA microarray analyses of circadian timing: the genomic basis of biological time. J Neuroendocrinol 15:991–1002
Panda S, Antoch MP, Miller BH et al (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109:307–320
Mehta N, Cheng HY (2013) Micro-managing the circadian clock: the role of microRNAs in biological timekeeping. J Mol Biol 425:3609–3624
Asher G, Sassone-Corsi P (2015) Time for food: the intimate interplay between nutrition, metabolism, and the circadian clock. Cell 161:84–92
Peek CB, Ramsey KM, Marcheva B, Bass J (2012) Nutrient sensing and the circadian clock. Trends Endocrinol Metab 23:312–318
Ribas-Latre A, Eckel-Mahan K (2016) Interdependence of nutrient metabolism and the circadian clock system: importance for metabolic health. Mol Metab 5:133–152
Perrin L, Loizides-Mangold U, Skarupelova S et al (2015) Human skeletal myotubes display a cell-autonomous circadian clock implicated in basal myokine secretion. Mol Metab 4:834–845
Ansari N, Agathagelidis M, Lee C, Korf HW, von Gall C (2009) Differential maturation of circadian rhythms in clock gene proteins in the suprachiasmatic nucleus and the pars tuberalis during mouse ontogeny. Eur J Neurosci 29:477–489
Yamazaki S, Yoshikawa T, Biscoe EW et al (2009) Ontogeny of circadian organization in the rat. J Biol Rhythm 24:55–63
Sumova A, Bendova Z, Sladek M et al (2006) Setting the biological time in central and peripheral clocks during ontogenesis. FEBS Lett 580:2836–2842
Rivkees SA (2007) The development of circadian rhythms: from animals to humans. Sleep Med Clin 2:331–341
Price DA, Close GC, Fielding BA (1983) Age of appearance of circadian rhythm in salivary cortisol values in infancy. Arch Dis Child 58:454–456
Balsalobre A, Brown SA, Marcacci L et al (2000) Resetting of circadian time in peripheral tissues by glucocorticoid signaling. Science 289:2344–2347
Sladek M, Jindrakova Z, Bendova Z, Sumova A (2007) Postnatal ontogenesis of the circadian clock within the rat liver. Am J Phys Regul Integr Comp Phys 292:R1224–R1229
Gotoh M, Maki T, Satomi S et al (1987) Reproducible high yield of rat islets by stationary in vitro digestion following pancreatic ductal or portal venous collagenase injection. Transplantation 43:725–730
Hohmeier HE, Mulder H, Chen G, Henkel-Rieger R, Prentki M, Newgard CB (2000) Isolation of INS-1-derived cell lines with robust ATP-sensitive K+ channel-dependent and -independent glucose-stimulated insulin secretion. Diabetes 49:424–430
Jacovetti C, Abderrahmani A, Parnaud G et al (2012) MicroRNAs contribute to compensatory beta cell expansion during pregnancy and obesity. J Clin Invest 122:3541–3551
Liu AC, Tran HG, Zhang EE, Priest AA, Welsh DK, Kay SA (2008) Redundant function of REV-ERBα and beta and non-essential role for Bmal1 cycling in transcriptional regulation of intracellular circadian rhythms. PLoS Genet 4:e1000023
Petrenko V, Saini C, Perrin L, Dibner C (2016) Parallel measurement of circadian clock gene expression and hormone secretion in human primary cell cultures. J Vis Exp. doi:10.3791/54673/
Qian J, Block GD, Colwell CS, Matveyenko AV (2013) Consequences of exposure to light at night on the pancreatic islet circadian clock and function in rats. Diabetes 62:3469–3478
Ru Y, Kechris KJ, Tabakoff B et al (2014) The multiMiR R package and database: integration of microRNA-target interactions along with their disease and drug associations. Nucleic Acids Res 42:e133
Su G, Morris JH, Demchak B, Bader GD (2014) Biological network exploration with Cytoscape 3. Curr Protoc Bioinformatics 47:8.13.1–8.1324
Lee J, Kim MS, Li R et al (2011) Loss of Bmal1 leads to uncoupling and impaired glucose-stimulated insulin secretion in beta-cells. Islets 3:381–388
Sadacca LA, Lamia KA, deLemos AS, Blum B, Weitz CJ (2011) An intrinsic circadian clock of the pancreas is required for normal insulin release and glucose homeostasis in mice. Diabetologia 54:120–124
Rakshit K, Hsu TW, Matveyenko AV (2016) Bmal1 is required for beta cell compensatory expansion, survival and metabolic adaptation to diet-induced obesity in mice. Diabetologia 59:734–743
Marcheva B, Ramsey KM, Buhr ED et al (2010) Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature 466:627–631
Kowalska E, Moriggi E, Bauer C, Dibner C, Brown SA (2010) The circadian clock starts ticking at a developmentally early stage. J Biol Rhythm 25:442–449
Brooks E, Canal MM (2013) Development of circadian rhythms: role of postnatal light environment. Neurosci Biobehav Rev 37:551–560
Petrenko V, Saini C, Giovannoni L et al (2017) Pancreatic alpha- and beta-cellular clocks have distinct molecular properties and impact on islet hormone secretion and gene expression. Genes Dev 31:383–398
Jermendy A, Toschi E, Aye T et al (2011) Rat neonatal beta cells lack the specialised metabolic phenotype of mature beta cells. Diabetologia 54:594–604
Rey G, Valekunja UK, Feeney KA et al (2016) The pentose phosphate pathway regulates the circadian clock. Cell Metab 24:462–473
Froy O (2007) The relationship between nutrition and circadian rhythms in mammals. Front Neuroendocrinol 28:61–71
Huang J, Lu C, Chen S, Hua L, Qian R (2010) Postnatal ontogenesis of clock genes in mouse suprachiasmatic nucleus and heart. Lipids Health Dis 9:22
Brown SA, Schibler U (1999) The ins and outs of circadian timekeeping. Curr Opin Genet Dev 9:588–594
Saito M, Suda M, Matzuda H (1978) Postnatal development of circadian rhythms in disaccharidase activities in rat small intestine. Am J Phys 234:E500–E503
Stephan FK (2002) The “other” circadian system: food as a Zeitgeber. J Biol Rhythm 17:284–292
Johnson J, Wu V, Donovan M et al (2010) Melanopsin-dependent light avoidance in neonatal mice. Proc Natl Acad Sci U S A 107:17374–17378
Luby MD, Hsu CT, Shuster SA et al (2012) Food anticipatory activity behavior of mice across a wide range of circadian and non-circadian intervals. PLoS One 7:e37992
Carneiro BT, Araujo JF (2009) The food-entrainable oscillator: a network of interconnected brain structures entrained by humoral signals? Chronobiol Int 26:1273–1289
Delezie J, Dumont S, Sandu C, Reibel S, Pevet P, Challet E (2016) Rev-erbα in the brain is essential for circadian food entrainment. Sci Rep 6:29386
Acknowledgements
We thank F. Gachon for helpful scientific discussions and guidance. Part of the data were presented in abstract form at the 2016 meeting of the Swiss Society for Endocrinology and Diabetology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Data availability
All data generated or analysed during this study are included in this published article (and its supplementary information files).
Funding
This work was supported by the Swiss National Foundation grants 310030-146138, 310030-169480 (RR), 31003A-166700 and CRSII3-154405 (CD), the Fondation Francophone pour la Recherche sur le Diabète (RR) sponsored by Fédération Française des Diabétiques, AstraZeneca, Eli Lilly, Merck Sharp & Dohme, Novo Nordisk and Sanofi, the Société Francophone du Diabète (CJ), Société Académique de Genève and Olga Mayenfisch Stiftung (CD).
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Contribution statement
CJ and ART generated and analysed the data, wrote the manuscript and approved its final version. CG, JS, SG, VP and CS contributed to the generation and analysis of the data, critically revised the manuscript and approved its final version. CD contributed to the analysis and interpretation of the data, critically reviewed the manuscript and approved its final version. RR designed the experiments, interpreted the data, wrote the manuscript and approved its final version. RR is the guarantor of this work.
Electronic supplementary material
ESM
(PDF 120 kb)
Rights and permissions
About this article
Cite this article
Jacovetti, C., Rodriguez-Trejo, A., Guay, C. et al. MicroRNAs modulate core-clock gene expression in pancreatic islets during early postnatal life in rats. Diabetologia 60, 2011–2020 (2017). https://doi.org/10.1007/s00125-017-4348-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-017-4348-6