Abstract
Aims/hypothesis
Pancreatic beta-like cells generated from human induced pluripotent stem cells (hiPSCs) or human embryonic stem cells (hESCs) offer an appealing donor tissue source. However, differentiation protocols that mainly use growth factors are costly. Therefore, in this study, we aimed to establish efficient differentiation protocols to change hiPSCs/hESCs to insulin (INS)+ cells using novel small-molecule inducers.
Methods
We screened small molecules that increased the induction rate of INS+ cells from hESC-derived pancreatic and duodenal homeobox 1 (PDX1)+ pancreatic progenitor cells. The differentiation protocol to generate INS+ cells from hiPSCs/hESCs was optimised using hit compounds, and INS+ cells induced with the compounds were characterised for their in vitro and in vivo functions. The inducing activity of the hit compounds was also examined using mouse embryonic pancreatic tissues in an explant culture system. Finally, RNA sequencing analyses were performed on the INS+ cells to elucidate the mechanisms of action by which the hit compounds induced pancreatic endocrine differentiation.
Results
One hit compound, sodium cromoglicate (SCG), was identified out of approximately 1250 small molecules screened. When SCG was combined with a previously described protocol, the induction rate of INS+ cells increased from a mean ± SD of 5.9 ± 1.5% (n = 3) to 16.5 ± 2.1% (n = 3). SCG induced neurogenin 3-positive cells at a mean ± SD of 32.6 ± 4.6% (n = 3) compared with 14.2 ± 3.6% (n = 3) for control treatment without SCG, resulting in an increased generation of endocrine cells including insulin-producing cells. Similar induction by SCG was confirmed using mouse embryonic pancreatic explants. We also confirmed that the mechanisms of action by which SCG induced pancreatic endocrine differentiation included the inhibition of bone morphogenetic protein 4 signalling.
Conclusions/interpretation
SCG improves the generation of pancreatic endocrine cells from multiple hiPSC/hESC lines and mouse embryonic pancreatic explants by facilitating the differentiation of endocrine precursors. This discovery will contribute to elucidating the mechanisms of pancreatic endocrine development and facilitate cost-effective generation of INS+ cells from hiPSCs/hESCs.
Data availability
The RNA sequencing data generated during the current study are available in the Gene Expression Omnibus (www.ncbi.nlm.nih.gov/geo) with series accession number GSE89973.
Similar content being viewed by others
Introduction
Type 1 diabetes is an autoimmune disease that is characterised by beta cell destruction in the pancreas, deficient insulin production and persistently high blood glucose levels. Treatment with exogenous insulin, although life-saving, does not restore physiological control of blood glucose levels. Therefore, islet cell or entire pancreas transplantation is a potentially curative treatment for type 1 diabetes [1,2,3]. However, transplantation therapy is limited by donor shortages and the potential for tissue rejection. One solution to donor tissue shortages is to generate beta cells or islet tissues from human induced pluripotent stem cells (hiPSCs) [4, 5] or human embryonic stem cells (hESCs) [6].
Previous reports have described the generation of insulin-positive (INS+) cells from hiPSCs/hESCs in vitro [7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22]. In 2014, two landmark papers reported the efficient generation of glucose-responsive insulin-secreting cells from hiPSCs/hESCs in vitro, with an induction rate of more than 40% [19, 20]. However, previously reported methods are costly because they mainly use growth factors as differentiation inducers. The aim of this study was therefore to establish differentiation protocols that produce INS+ cells from hiPSCs/hESCs by identifying small molecules that act as potent inducers.
We used chemical screening to examine the effects of approximately 1250 compounds on pancreatic and duodenal homeobox 1 (PDX1)+ pancreatic progenitors generated from hESCs using a modified version of a previously described protocol [17]. We identified one small molecule, sodium cromoglicate (SCG), which facilitates the differentiation of PDX1+ cells into INS+ cells. Small molecules such as SCG, which have unique biological activities, have opened the doors to new biological studies and may further lead to the development of new treatments by significantly lowering cost [23].
Methods
For detailed methods, please see electronic supplementary material (ESM) Methods.
Cell culture
The use of hiPSCs and hESCs was approved by the Ethics Committee of Kyoto University, and informed consent was obtained from all donor individuals from which these cell lines were generated. Two peripheral blood-derived hiPSC lines, 648A1 and 692D2, three fibroblast-derived hiPSC lines, 409B2, 206B6 and 201B7 [4, 5, 24] and three hESC lines, H9, KhES1 and KhES3 [6, 25], were maintained on mitomycin C-treated SNL cell feeder layers, kindly provided by A. Bradley (Sanger Institute, Cambridge, UK), in Primate ES Cell Medium (ReproCELL, Kanagawa, Japan) containing 500 U/ml penicillin/streptomycin (PS; Thermo Fisher Scientific, Waltham, MA, USA) and 4 ng/ml recombinant human basic fibroblast growth factor (Wako, Osaka, Japan). hiPSCs and hESCs were routinely tested for mycoplasma contamination and were negative.
Differentiation protocol
In order to generate pancreatic lineage cells, hiPSC/hESC colonies grown on SNL feeder cells at 70% confluence were first dissociated using CTK dissociation solution and subsequently incubated on gelatin-coated plates for 30 min to remove SNL cells. The cells were then dissociated into single cells via gentle pipetting after treatment with Accutase (Innovative Cell Technologies, San Diego, CA, USA) for 20 min and seeded on Matrigel (BD Biosciences, San Jose, CA, USA)-coated 24-well plates at a density of 2 × 105 cells/well. The induction protocol was based on a modified version of a previously described method [17]. In stage 1, the dissociated single hiPSCs/hESCs were treated with 1 μmol/l CHIR99021 (Axon Mechem, Groningen, the Netherlands), 100 ng/ml activin A (R&D Systems, Minneapolis, MN, USA) and 10 μmol/l ROCK inhibitor Y-27632 (Wako) in stage 1 medium containing RPMI 1640 (Nacalai Tesque, Kyoto, Japan) supplemented with 2× B27 supplements (Thermo Fisher Scientific) and 500 U/ml PS for 2 days, after which the medium was changed to stage 1 medium containing 1 μmol/l CHIR99021 and 100 ng/ml activin A every day for an additional 3 days. In stage 2, the cells were treated with 1 μmol/l dorsomorphin (Merck, Darmstadt, Germany), 2 μmol/l retinoic acid (Sigma-Aldrich, St Louis, MO, USA) and 10 μmol/l SB431542 (Cayman Chemical, Ann Arbor, MI, USA) in stage 2 medium containing Improved MEM Zinc Option medium (Thermo Fisher Scientific) supplemented with 2× B27 supplements and 500 U/ml PS, after which the medium was changed every 2 days for a total of 6 days. In stage 3, four chemical compounds (4Fs)—10 μmol/l forskolin (Wako), 10 μmol/l dexamethasone (Wako), 5 μmol/l ALK5 inhibitor II (Wako) and 10 mmol/l nicotinamide (STEMCELL Technologies, Vancouver, BC, Canada)—were used with or without SCG (Sigma) in stage 3 medium containing Improved MEM Zinc Option medium supplemented with 2× B27 supplements and 500 U/ml PS, and the medium was changed every 4 days for a total of 8 days.
Chemical screening
To screen chemical compounds that facilitate differentiation into INS+ cells, PDX1+ pancreatic progenitors generated from KhES3 cells were dissociated into single cells using Accutase treatment and reseeded on Matrigel-coated 384-well plates. The cells were then treated with 4Fs and individual compounds using a robotic dispenser (Biomek 3000; Beckman Coulter, Fullerton, CA, USA) and cultured for an additional 8 days, changing the medium with the compounds every 4 days. The cells were subsequently immunostained with anti-insulin antibodies, and the number and induction rate of INS+ cells were analysed using an imaging cytometer (Celigo; Brooks, Chelmsford, MA, USA). The data were normalised as the fold change compared with that observed in the control samples treated with 4Fs alone (see ESM Methods).
Immunostaining
Immunostaining was performed on hiPSC-derived differentiated cells and embryonic pancreatic cells from MIP-GFP (CD1-Tg(Ins1-GFP)1Hara) mice, kindly provided by M. Hara, University of Chicago, IL, USA [26], as previously described, using primary antibodies against SRY-box 17 (SOX17) (AF1924, 1:500; R&D Systems), forkhead box A2 (FOXA2) (AF2400, 1:500; R&D Systems), PDX1 (AF2419, 1:500; R&D Systems), insulin (A0564, 1:500; Dako, Glostrup, Denmark), C-peptide (MON5021, 1:1000; Cell Sciences, Newburyport, MA, USA), glucagon (GCG) (G2654, 1:500; Sigma), somatostatin (SST) (A0566, 1:500; Dako), ghrelin (GHRE) (sc10368, 1:500; Santa Cruz Biotechnology, Dallas, TX, USA), amylase (AMY) (A8273, 1:500; Sigma), CK19 (M0888, 1:500; Dako), NKX6.1 (F55A12, 1:50; University of Iowa, Iowa City, IA, USA), neurogenin 3 (NGN3) (AB5684, 1:500; Millipore, Darmstadt, Germany) and Ki67 (556003, 1:100; BD Biosciences) [7, 13, 17, 22]. Nuclei were stained with Hoechst 33342 (H3570, 1:1000; Thermo Fisher Scientific). For quantification, 15 images per well were obtained using Celigo.
RT-PCR and quantitative real-time RT-PCR
RT-PCR and quantitative real-time RT-PCR (qRT-PCR) were performed on hiPSC-derived differentiated cells and embryonic pancreatic cells from MIP-GFP mice, as previously described, to detect the mRNA expression of SOX17, PDX1, NGN3 (also known as Neurog3), ISL1, INS, GCG, SST, GHRE (also known as GHRL), MAFA, Ins1, Ngn3, BMP4, ACTB and GAPDH [27]. Total RNA was isolated using an RNeasy kit (Qiagen, Hilden, Germany), followed by cDNA synthesis using ReverTra Ace (Toyobo, Osaka, Japan). PCR was performed using a thermal cycler (Veriti 96-well Thermal Cycler; Thermo Fisher Scientific) and the Ex-Taq PCR kit (Takara, Shiga, Japan). qPCR was performed using the Step One Plus Real-Time PCR System (Thermo Fisher Scientific) and SYBR Green PCR Master Mix (Takara). PCR reactions were performed in triplicate for each sample. The primer sequences are shown in ESM Table 1.
Flow cytometry
Flow cytometric analyses were performed on dissociated embryonic pancreatic cells from MIP-GFP mice using FACSAria II (BD Biosciences) (see ESM Methods).
C-peptide release assay and insulin content analysis
These examinations were performed on hiPSC-derived differentiated cells using the Human C-peptide ELISA Kit (Mercodia, Uppsala, Sweden) and the Human Insulin ELISA Kit (Mercodia), respectively (see ESM Methods).
Cell transplantation into mouse models of diabetes
All animal experiments were approved by the Animal Care Committee of Kyoto University in accordance with the Guidelines for Animal Experiments of Kyoto University. Mice were maintained in a specific pathogen-free environment at 25°C on a 12 h light–dark cycle, with free access to a standard irradiated diet. A mouse model of type 1 diabetes was generated from male NOD-Scid (NOD/ShiJic-scid/Jcl) mice (CLEA Japan, Osaka, Japan) of 6 weeks of age or older using a single i.p. injection of 120 mg/kg streptozotocin (STZ) and stabilised for 10 days prior to transplantation. All mice used in this study (n = 3 per group) demonstrated hyperglycaemia (blood glucose level > 22.2 mmol/l) before transplantation. hiPSC-derived differentiated cells were dissociated with 0.25% trypsin and resuspended in DMEM with 10% FBS, and 2.0 × 106 or 4.0 × 106 cells were injected into the renal subcapsules of STZ mice using a 24 G catheter (NIPRO, Osaka, Japan). Blood glucose levels were observed for 6 months after transplantation.
Explant culture
Embryonic day (E)12.5–16.5 pancreatic tissues derived from the MIP-GFP mice were dissected and isolated as previously described [28]. The tissues were then cultured at the air–fluid interface of 0.4 μm pore polycarbonate filters (Millipore) in stage 3 medium containing 4Fs with or without SCG for 4 days, as previously described [26, 27].
RNA sequencing
Libraries were prepared from hiPSC-derived differentiated cells with the Illumina TruSeq Stranded Total RNA Sample Prep kit (Illumina, San Diego, CA, USA) and sequenced using the HiSeq2500 in 100-cycle single-read mode (see ESM Methods).
Statistical analysis
Statistical analyses were performed using two-tailed paired Student’s t tests to compare two individual groups and one-way ANOVA followed by Bonferroni’s test to compare multiple groups. Data are expressed as means ± SD. No randomisation or investigator blinding was performed. Sample size was not predefined by power calculations. No experimental samples were excluded. p values of <0.05 were considered statistically significant.
Results
Screening for small molecules that improve the induction efficiency of INS+ cells
We designed a screening procedure based on a modified version of a previously published protocol that mimics the embryonic development of pancreatic beta cells (ESM Fig. 1a) [17]. Both SOX17+ definitive endoderm and PDX1+ pancreatic progenitor cells were induced at a high efficiency (83.3 ± 5.6% and 80.3 ± 4.6%, respectively) from an hESC line, KhES3, although INS+ cells were induced at a low rate (5.9 ± 1.5%; ESM Fig. 1b). Four factors/compounds (4Fs: nicotinamide, forskolin, dexamethasone and ALK5 inhibitor) were used in the rate-limiting step from PDX1+ to INS+ cells. Chemical screening was performed by adding individual compounds to PDX1+ cells on culture day 11 (ESM Fig. 1c). Approximately 1250 compounds were examined; however, no hits were found that induced INS+ cells from PDX1+ cells. The same compounds were then screened by adding them individually to 4Fs to induce INS+ cells with higher efficiency. Hit compounds were defined as those with which the induction rate of INS+ cells was more than twice as high as that achieved with 4Fs alone, without toxicity or autofluorescence (ESM Fig. 1d). Among the approximately 1250 compounds, only one hit chemical, SCG (Fig. 1a), was identified.
(a) Chemical structure of SCG. (b) Schematic of the protocol to differentiate hiPSCs/hESCs into INS+ cells. (c, d) Analysis of the induction rate of INS+ cells (c) and INS mRNA expression (d) after 4 (day 15), 8 (day 19), 12 (day 23), 16 (day 27) and 20 (day 31) days of SCG treatment. Each value was normalised to that without SCG treatment on day 15 in (d). White bars: 4Fs alone; black bars: 4Fs + SCG. (e) Concentration-dependent curve of the induction activity of SCG. (f) Induction of INS+ cells on day 19 by SCG alone. qPCR analyses of INS gene expression were normalised to samples with medium (as control) alone. (g) Effects of combination treatment with three factors and SCG on INS gene expression. Each value was normalised to samples with medium alone. FAD: forskolin, ALK5 inhibitor and dexamethasone; NAD: nicotinamide, ALK5 inhibitor and dexamethasone; NFD: nicotinamide, forskolin and dexamethasone; NFA: nicotinamide, forskolin and ALK5 inhibitor. (h) Differentiation of multiple hiPSC/hESC lines into INS+ cells. White bars: 4Fs alone; black bars: 4Fs + SCG. Data in (c–h) are from three independent experiments and are presented as means ± SD. Scale bars, 100 μm. *p < 0.05, **p < 0.01
Optimisation of differentiation protocol for INS+ cells using SCG
In order to determine the optimal conditions for SCG application, we examined the induction rate of INS+ cells from an hiPSC line, 201B7, at various time points using 10 mmol/l SCG (Fig. 1b). The induction rate of INS+ cells following treatment with 4Fs + SCG was highest on day 19 (16.5 ± 2.1%), and was considerably higher than that obtained with control treatment using 4Fs alone (5.9 ± 1.5%; Fig. 1c). The samples with or without SCG treatment on day 23 contained more than 10% INS+ cells. However, the induction rate of INS+ cells was higher with SCG treatment (Fig. 1c). Consistently, qRT-PCR analysis showed that the gene expression of INS reached its peak after 8 days (on day 19) of 4Fs + SCG treatment (Fig. 1d).
We then exposed the 201B7-derived PDX1+ cells to 4Fs and 0.001–20 mmol/l SCG for 8 days. The effect was dose-dependent, with the highest efficiency and no toxicity at a concentration of 10 mmol/l (Fig. 1e) [29]. Optimal results were obtained by treating PDX1+ cells with a combination of 4Fs and 10 mmol/l SCG for 8 days.
Treatment with SCG alone induced INS+ cells at a rate of 3.2 ± 0.4% compared with 0.6 ± 0.2% for control treatment without SCG, which is consistent with qRT-PCR results showing INS gene expression levels of around six times higher than those achieved with control treatment (Fig. 1f). We then confirmed that the addition of SCG to all combinations of three factors did not significantly increase INS expression, indicating that SCG requires the combination of all four factors for synergistic activity (Fig. 1g).
Different hiPSC/hESC lines vary in their differentiation potential [24, 30]. We examined the effects of 4Fs + SCG treatment using eight hiPSC/hESC lines. The addition of SCG to 4Fs increased the induction rate of INS+ cells in six out of the eight cell lines examined (648A1, 692D2, 409B2, 201B7, KhES3 and H9), although the induction rate of INS+ cells differed among the hiPSC/hESC lines (Fig. 1h). These results suggest that our differentiation protocol using 4Fs + SCG has broad application for multiple hiPSC/hESC lines.
Characteristics of INS+ cells induced by SCG
We next examined the gene expression characteristics at various stages of pancreatic development in culture using 4Fs + SCG treatment. The examined marker genes included SOX17 for definitive endoderm, PDX1, NGN3 and ISL1 for pancreatic progenitor, and INS, GCG, SST, GHRE and MAFA for endocrine cells, and were analysed from days 0 to 19. The results confirmed the gene expression of all markers except MAFA (Fig. 2a).
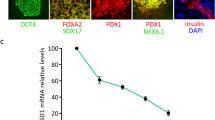
(a) Expression of pancreatic lineage markers in cells induced by 4Fs + SCG. PC (positive control): adult human pancreas. Representative data from at least three independent experiments are shown. (b) C-peptide secretion by INS+ cells induced by 4Fs alone or 4Fs + SCG on day 19 in response to two concentrations of d-glucose and KCl. (c) Insulin contents of INS+ cells induced by 4Fs alone or 4Fs + SCG on day 19. (d–f) Fasting blood glucose levels in diabetic mouse groups that received a sham operation (red line) or were transplanted with 400 mouse islets (purple line) or cells generated using treatment with 4Fs (yellow line; percentage of INS+ cells: 7.4 ± 1.6%) or 4Fs + SCG (blue line; percentage of INS+ cells: 14.6 ± 1.2%) for the first 4 weeks (d) and from 1 to 6 months (e) after transplantation. Control mice did not receive STZ injection (green line; n = 3 for each group). The arrow in (d) indicates the date of transplantation. (f) The effects of transplanting 2.0 × 106 cells (blue line) and 4.0 × 106 cells (red line) generated with 4Fs + SCG on blood glucose levels (n = 3; percentage of INS+ cells: 15.9 ± 1.8%). (g) Plasma human C-peptide levels in host mice measured 7 days after implantation. (h) Images of anti-insulin immunostaining (red) and TUNEL assay (green) on grafts from 4Fs + SCG-treated cells at 1 day after transplantation. Data in (b–g) are from three independent experiments and are presented as the means ± SD. Scale bar, 100 μm. *p < 0.05 for samples treated with 4Fs vs 4Fs + SCG in (d) and for samples transplanted with 2.0 × 106 vs 4.0 × 106 cells in (f). Statistical analyses in (b) were performed using ANOVA
We then examined the synthesis and release of C-peptide in order to evaluate the functionality of INS+ cells induced by 4Fs + SCG. The cells induced by 4Fs alone and those induced by 4Fs + SCG both secreted C-peptide in response to KCl, although they did not differentiate into fully mature beta cells with glucose-responsive insulin secretion (Fig. 2b). Insulin content was not significantly different between the two cell populations (Fig. 2c). We also evaluated the physiological function of INS+ cells induced by 4Fs + SCG in vivo by transplanting a total of 2.0 × 106 cells into one renal subcapsule of NOD-Scid mice with type 1 diabetes induced by STZ administration. The transplanted INS+ cells induced by 4Fs + SCG mildly ameliorated hyperglycaemia in these mice, but only for the first 10 days (Fig. 2d, e; blue line). The therapeutic effects were significantly higher than with the transplantation of cells induced with 4Fs alone on days 5, 6, 7 and 9 (Fig. 2d; yellow line). We examined the therapeutic effects up to 6 months after transplantation, but found no improvement in blood glucose levels after the first 10 days (Fig. 2d, e).
We increased the cell number to 4.0 × 106 cells (2.0 × 106 cells per subcapsule of each kidney), which approximates the capacity of the kidney subcapsular space (Fig. 2f; red line). However, the transplantation experiments produced similar results, with mild therapeutic effects obtained around the first 10 days, although the transplantation of a larger number of cells lowered the blood glucose levels of the host mice to a larger extent (Fig. 2f).
A slight increase in plasma human C-peptide levels in host mice was detected 7 days after transplantation (Fig. 2g), with the values falling below detection levels by day 14 (data not shown). We then performed a detailed histological examination of the grafts from 1 to 10 days after transplantation and found that cells in the grafts were apoptotic, as evidenced by positive staining for TUNEL (Fig. 2h). The size and cell number of the grafts gradually decreased, and the grafts eventually disappeared by day 7 (data not shown). These results suggest that although the INS+ cells induced by treatment with 4Fs + SCG were not fully mature, they developed the ability to secrete functional insulin protein.
Addition of SCG facilitates differentiation into multiple endocrine cell types
In order to further characterise the cells that differentiated in the presence of SCG, we examined the expression of pancreatic endocrine lineage markers. After 8 days of treatment, the cells induced by 4Fs + SCG were positively stained with multiple endocrine markers, including INS, GCG, SST and GHRE (Fig. 3a). We should note that cells positive for NKX6.1 or pancreatic polypeptide were not found in these cultures.
(a) Immunostaining images of cells induced by 4Fs + SCG on day 19 for pancreatic lineage markers. Representative data from at least three independent experiments are shown. (b) Quantification of the induction rate of INS+, GCG+, SST+, GHRE+, AMY+ and CK19+ cells on day 19. White bars: 4Fs alone; black bars: 4Fs + SCG. (c–f) SCG treatment increased the gene expression levels of endocrine markers. Each value was normalised to adult human pancreas (positive control, PC) samples. Data are from three independent experiments and are presented as means ± SD in (b–f). Scale bars, 100 μm. *p < 0.05, **p < 0.01
After 8 days of treatment with 4Fs + SCG, the induction rate of the cells positively stained with the endocrine lineage markers and the mRNA expression levels of the markers were significantly higher in cells induced with 4Fs + SCG than in the 4Fs-induced cells (Fig. 3b–f). Most INS+ cells induced with 4Fs + SCG were GCG− (Fig. 3a). The percentage of double-positive cells for INS and GCG among the total INS+ cells induced by 4Fs + SCG (INS+GCG+/INS+) was low (4.2 ± 1.9%, n = 3) and not significantly different from that induced by 4Fs alone (2.8 ± 1.2%, n = 3).
We also examined the induction rate of cells positive for a pancreatic ductal marker, cytokeratin (CK)19, and those positive for an acinar cell marker, AMY. After 8 days of treatment with 4Fs alone, 51.9 ± 3.6% of cells were CK19+ and 21.6 ± 2.3% were AMY+. There were no significant differences in the induction rate of CK19+ or AMY+ cells (53.8 ± 3.2% and 22.7 ± 1.9%, respectively) after the 8-day treatment with 4Fs + SCG (Fig. 3a, b). These results indicate that the addition of SCG to 4Fs facilitates differentiation into endocrine lineages, but not into ductal or acinar lineages.
Addition of SCG facilitates the induction of NGN3+ endocrine progenitor cells
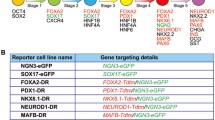
The addition of SCG to 4Fs increased the induction rate of multiple endocrine cell types. We therefore evaluated the possibility that NGN3+ endocrine precursor cells are induced by adding SCG to 4Fs. We evaluated the gene expression of PDX1 and NGN3 under the conditions of our differentiation protocol. While there were no significant increases in PDX1 expression, the expression level of NGN3 increased after 4 days (on day 15) with 4Fs + SCG treatment (Fig. 4a, b). Consistently, the induction rate of NGN3+ cells during 4Fs + SCG treatment was highest on day 15 (32.6 ± 4.6%) and higher than that obtained with 4Fs alone (14.2 ± 3.6%; Fig. 4c). These results suggest that SCG enhanced the differentiation into NGN3+ cells, thereby resulting in the increased generation of multiple endocrine but not non-endocrine cell types.
(a, b) The addition of SCG to 4Fs significantly increased the gene expression level of NGN3 (b), but not that of PDX1 (a). Each value was normalised to adult human pancreas (positive control, PC) samples. (c) Immunostaining images of cells induced by 4Fs or 4Fs + SCG on day 15 for NGN3 and nuclei, with graphs showing the induction rate of NGN3+ endocrine precursors on days 15 and 19. White bars: 4Fs alone; black bars: 4Fs + SCG. (d, e) Immunostaining images of cells induced by 4Fs + SCG on day 19 for INS, Ki67 and nuclei in (d) or NGN3, Ki67 and nuclei in (e). SCG treatment did not expand the INS+ (d) or NGN3+ (e) cell populations on day 19. Data are from three independent experiments and are presented as means ± SD. Scale bars, 100 μm. **p < 0.01
We next examined the percentage of cells double-positive for INS and Ki67 (INS+Ki67+ cells) among the INS+ cells induced by 4Fs + SCG (INS+Ki67+/INS+). The percentage of cells induced by 4Fs + SCG or 4Fs was similar (approximately 1–2%; Fig. 4d). Similar results were obtained for the percentage of NGN3+Ki67+/NGN3+ cells (Fig. 4e). These results indicate that SCG treatment does not expand the populations of NGN3+ endocrine precursors or INS+ cells.
SCG induces NGN3+ cells in mouse embryonic pancreatic explants
In order to examine the effects of SCG on NGN3+ and INS+ cells in mouse embryos, we treated E12.5, 14.5 and 16.5 pancreatic tissues removed from MIP-GFP mice with 4Fs + SCG in an explant culture system for 4 days (Fig. 5a). In comparison with 4Fs treatment, the addition of SCG increased the number of INS1 (green fluorescent protein)-positive cells (Fig. 5b). The expression levels of Ngn3 and Ins1 were also higher in the samples treated with 4Fs + SCG compared with 4Fs alone (Fig. 5c, d). Similar to the results of the in vitro hiPSC differentiation cultures, there were no significant changes in the percentages of NGN3+Ki67+/NGN3+ or INS+Ki67+/INS+, which were each approximately 1–2% (Fig. 5e). These results suggest that SCG increases the generation of INS+ cells in mice by inducing NGN3+ cell differentiation and not by expanding the populations of NGN3+ or INS+ cells.
(a) Schematic of explant cultures of embryonic mouse pancreatic tissues. (b) The addition of SCG to 4Fs increased the percentage of green fluorescent protein (GFP)-positive cells in the embryonic pancreatic tissues of MIP-GFP mice. White bars: 4Fs alone; black bars: 4Fs + SCG. (c, d) The addition of SCG to 4Fs increased the gene expression levels of Ngn3 (c) and Ins1 (d) in E14.5 mouse pancreatic tissues. Each value was normalised to samples of E14.5 pancreas (positive control, PC). (e, f) Immunostaining of E14.5 mouse pancreatic cells after 4 days of treatment with 4Fs + SCG for NGN3, Ki67 and nuclei (e) and INS, Ki67 and nuclei (f). Treatment with 4Fs + SCG did not facilitate the proliferation of NGN3+ (e) or INS+ cells (f). Data are from three independent experiments and are presented as means ± SD in (b–e). Scale bars, 100 μm. *p < 0.05, **p < 0.01. Statistical analyses were performed using ANOVA in (c, d)
SCG promotes differentiation into endocrine progenitors by inhibiting BMP4 signalling
In an attempt to elucidate the mechanisms of action by which SCG induces pancreatic endocrine differentiation, we performed RNA sequencing analyses to compare the gene expression profiles of cells after 8 days of treatment with SCG and those treated with H2O (controls). We also compared the gene expression profiles of cells treated with 4Fs + SCG and those treated with 4Fs alone. In both comparisons, differentially expressed genes with more than twofold change were selected. Genes common in the two comparisons were chosen to identify molecules whose expressions were specifically changed by SCG treatment. We found a small number of candidates (19 and 17 transcripts, respectively) that were significantly upregulated or downregulated, according to these criteria (ESM Tables 2, 3). Next, we performed two pathway analyses, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses, both of which showed the bone morphogenetic protein (BMP)4 pathway to be significantly altered (ESM Tables 4, 5), and we therefore focused on BMP4 out of the genes significantly downregulated in the SCG-treated group.
Further analysis by qRT-PCR confirmed that the expression of BMP4 was significantly downregulated in cells after 8 days of treatment with 4Fs + SCG relative to cells treated with 4Fs alone (Fig. 6a). Then, in order to examine whether BMP4 signals are involved in the differentiation into NGN3+ endocrine progenitors and the subsequent generation of INS+ cells, we added BMP4 (100 ng/ml) and a BMP signal antagonist, LDN-193189 (0.5 μg/ml), to the differentiation culture with 4Fs + SCG. The results showed that the addition of BMP4 significantly reduced the expression levels of NGN3 and INS as well as the induction rate of INS+ cells, while the addition of LDN-193189 significantly increased the expression of NGN3 and INS, although it did not increase the induction rate of INS+ cells (Fig. 6b–d). These results suggest that the mechanisms of SCG-induced endocrine differentiation include the inhibition of BMP4 signalling.
(a) BMP4 expression was significantly downregulated in cells treated with 4Fs + SCG in comparison with 4Fs alone. NGN3 (b) and INS (c) expression was compared among cells after 8 days of four types of treatment. (d) Induction rate of INS+ cells after 8 days of treatment. The samples treated with 4Fs alone were used to normalise the data in (a–c). Data are from three independent experiments and are presented as means ± SD. *p < 0.05, **p < 0.01. Statistical analyses were performed using ANOVA in (b–d). LDN, LDN–193189
Discussion
In this study, we used unbiased chemical screening to identify a small molecule, SCG, that facilitates the differentiation of hiPSCs/hESCs into INS+ cells. The advantages of using small molecules for hiPSC/hESC differentiation include low cost, stability, ready supply, mechanistic implications and ease of use [23]. Previously reported protocols for the generation of pancreatic lineage cells mainly used expensive growth factors, which limits their clinical application. Modification of the protocols by using small molecules such as SCG may significantly lower the cost.
SCG is a small molecule that is used to treat allergies and asthma. It exerts its anti-allergic effects by inhibiting the release of chemical inflammatory mediators, including histamine [31,32,33]. It has also been shown that chloride channels, heat-shock protein 90 and G-protein-coupled receptor 35 are target molecules of SCG [34,35,36,37,38,39]. Although no reports have described the effects of SCG on pancreatic cells, based on those previous works, we suggest that SCG operates through signals related to chloride channels, heat-shock protein 90 or G-protein-coupled receptor 35 in pancreatic endocrine progenitors or on the membranes of these cells, since the mechanisms of the SCG anti-allergic effects include membrane stabilisation. In contrast to SCG, other anti-allergic drugs in the examined compound libraries did not increase the induction efficiency of INS+ cells (data not shown). These drugs included: (1) second-generation histamine H1 receptor antagonists (antihistamines) such as cetirizine dihydrochloride, levocetirizine dihydrochloride, fexofenadine hydrochloride, epinastine hydrochloride, olopatadine, loratadine, ebastine and ketotifen; (2) leukotriene receptor antagonists such as pranlukast and montelukast; and (3) chemical mediator release inhibitors such as tranilast and ibudilast.
SCG facilitates the generation of INS+ cells, but not their maturation into functional beta cells, as shown by the absence of MAFA expression. In addition, INS+ cells induced with SCG were negative for NKX6.1 and did not show potential for glucose-responsive insulin secretion, suggesting that they may be embryonic or immature beta-like cells [40]. INS+ cells induced with 4Fs + SCG showed slightly higher KCl-stimulated C-peptide secretion and therapeutic effects on blood glucose levels after transplantation than 4Fs-induced cells. However, considering the similar insulin content per cell in the two cell populations, the effects of SCG treatment were most likely due to an increased cell number and not functional improvement of the INS+ cells. Future studies combining exogenous maturating signals, such as those produced by BMPs, TGF-β and EGF, are likely required to enable the generation of functionally mature beta cells [41,42,43].
SCG induced the differentiation of PDX1+ pancreatic progenitor cells into NGN3+ endocrine precursors in both cultures of hiPSCs/hESCs in vitro and mouse embryonic pancreas tissues ex vivo. In contrast, SCG did not act on adult islet cells (data not shown), among which NGN3+ cells may not exist or are at low numbers [44,45,46,47]. The results of our RNA sequencing analyses led us to focus on the effects of BMP4 signalling. Pancreatic progenitor cells were generated under the continuous suppression of BMP signalling in hESC differentiation culture [16, 48, 49]. The present study also showed that SCG inhibits BMP4 signalling at the transcription level to induce the differentiation of pancreatic endocrine progenitors. A previous report showed that Sox2 was a downstream molecule of Bmp4 in mouse lens development [50]. We found, through the RNA sequencing results, that SCG treatment downregulated SOX2 (fold change: 0.537 for SCG/control and 0.391 for 4Fs + SCG/4Fs). This finding hints that a SOX2-dependent mechanism might regulate the differentiation of pancreatic endocrine progenitors upon SCG treatment and is worthy of future study.
In conclusion, we identified a small molecule, SCG, that improves INS+ cell generation from multiple hiPSC/hESC lines by facilitating the differentiation of endocrine precursors. SCG may contribute to elucidating the mechanisms of pancreatic endocrine development and provide a cost-effective supply of new cell sources for regenerative therapy against type 1 diabetes.
Abbreviations
- 4Fs:
-
Nicotinamide forskolin dexamethasone and ALK5 inhibitor
- AMY:
-
Amylase
- BMP:
-
Bone morphogenetic protein
- CK:
-
Cytokeratin
- E:
-
Embryonic day
- ESC:
-
Embryonic stem cell
- GCG:
-
Glucagon
- hESC:
-
Human embryonic stem cell
- hiPSC:
-
Human induced pluripotent stem cell
- GHRE:
-
Ghrelin
- INS+ :
-
Insulin-positive
- iPSC:
-
Induced pluripotent stem cell
- NGN3:
-
Neurogenin 3
- PDX1:
-
Pancreatic and duodenal homeobox 1
- PS:
-
Penicillin/streptomycin
- qRT-PCR:
-
Quantitative real-time RT-PCR
- SCG:
-
Sodium cromoglicate
- SOX17:
-
SRY-box 17
- SST:
-
Somatostatin
- STZ:
-
Streptozotocin
References
Ryan EA, Paty BW, Senior PA et al (2005) Five-year follow-up after clinical islet transplantation. Diabetes 54:2060–2069
Speight J, Reaney MD, Woodcock AJ, Smith RM, Shaw JW (2010) Patient-reported outcomes following islet cell or pancreas transplantation (alone or after kidney) in type 1 diabetes: a systematic review. Diabet Med 27:812–822
Shapiro AM, Ricordi C, Hering BJ et al (2006) International trial of the Edmonton protocol for islet transplantation. N Engl J Med 355:1318–1330
Takahashi K, Tanabe K, Ohnuki M et al (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872
Okita K, Matsumura Y, Sato Y et al (2011) A more efficient method to generate integration-free human iPS cells. Nat Methods 8:409–412
Thomson JA, Itskovitz-Eldor J, Shapiro SS et al (1998) Embryonic stem cell lines derived from human blastocysts. Science 282:1145–1147
D’Amour KA, Bang AG, Eliazer S et al (2006) Production of pancreatic hormone-expressing endocrine cells from human embryonic stem cells. Nat Biotechnol 24:1392–1401
Jiang J, Au M, Lu K et al (2007) Generation of insulin-producing islet-like clusters from human embryonic stem cells. Stem Cells 25:1940–1953
Kroon E, Martinson LA, Kadoya K et al (2008) Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo. Nat Biotechnol 26:443–452
Tateishi K, He J, Taranova O, Liang G, D’Alessio AC, Zhang Y (2008) Generation of insulin-secreting islet-like clusters from human skin fibroblasts. J Biol Chem 283:31601–31607
Maehr R, Chen S, Snitow M et al (2009) Generation of pluripotent stem cells from patients with type 1 diabetes. Proc Natl Acad Sci U S A 106:15768–15773
Chen S, Borowiak M, Fox JL et al (2009) A small molecule that directs differentiation of human ESCs into the pancreatic lineage. Nat Chem Biol 5:258–265
Thatava T, Nelson TJ, Edukulla R et al (2011) Indolactam V/GLP-1-mediated differentiation of human iPS cells into glucose-responsive insulin-secreting progeny. Gene Ther 18:283–293
Kelly OG, Chan MY, Martinson LA et al (2011) Cell-surface markers for the isolation of pancreatic cell types derived from human embryonic stem cells. Nat Biotechnol 29:750–756
Nostro MC, Sarangi F, Ogawa S et al (2011) Stage-specific signalling through TGFβ family members and WNT regulates patterning and pancreatic specification of human pluripotent stem cells. Development 138:861–871
Schulz TC, Young HY, Agulnick AD et al (2012) A scalable system for production of functional pancreatic progenitors from human embryonic stem cells. PLoS One 7:e37004
Kunisada Y, Tsubooka-Yamazoe N, Shoji M, Hosoya M (2012) Small molecules induce efficient differentiation into insulin-producing cells from human induced pluripotent stem cells. Stem Cell Res 8:274–284
Rezania A, Bruin JE, Riedel MJ et al (2012) Maturation of human embryonic stem cell-derived pancreatic progenitors into functional islets capable of treating pre-existing diabetes in mice. Diabetes 61:2016–2029
Pagliuca FW, Millman JR, Gürtler M et al (2014) Generation of functional human pancreatic β cells in vitro. Cell 159:428–439
Rezania A, Bruin JE, Arora P et al (2014) Reversal of diabetes with insulin-producing cells derived in vitro from human pluripotent stem cells. Nat Biotechnol 32:1121–1133
Bruin JE, Erener S, Vela J, Hu X (2014) Characterization of polyhormonal insulin-producing cells derived in vitro from human embryonic stem cells. Stem Cell Res 12:194–208
Toyoda T, Mae S, Tanaka H et al (2015) Cell aggregation optimizes the differentiation of human ESCs and iPSCs into pancreatic bud-like progenitor cells. Stem Cell Res 14:185–197
Araoka T, Mae S, Kurose Y et al (2014) Efficient and rapid induction of human iPSCs/ESCs into nephrogenic intermediate mesoderm using small molecule-based differentiation methods. PLoS One 9:e84881
Kajiwara M, Aoi T, Okita K et al (2012) Donor-dependent variations in hepatic differentiation from human-induced pluripotent stem cells. Proc Natl Acad Sci U S A 109:12538–12543
Suemori H, Yasuchika K, Hasegawa K, Fujioka T, Tsuneyoshi N, Nakatsuji N (2006) Efficient establishment of human embryonic stem cell lines and long-term maintenance with stable karyotype by enzymatic bulk passage. Biochem Biophys Res Commun 345:926–932
Hara M, Wang X, Kawamura T et al (2003) Transgenic mice with green fluorescent protein-labeled pancreatic β-cells. Am J Physiol Endocrinol Metab 284:e177–e183
Mae S, Shono A, Shiota F et al (2013) Monitoring and robust induction of nephrogenic intermediate mesoderm from human pluripotent stem cells. Nat Commun 4:1367
Gotoh M, Maki T, Satomi S et al (1987) Reproducible high yield of rat islets by stationary in vitro digestion following pancreatic ductal or portal venous collagenase injection. Transplantation 43:725–730
Yuengsrigul A, Chin TW, Nussbaum E (1999) Immunosuppressive and cytotoxic effects of furosemide on human peripheral blood mononuclear cells. Ann Allergy Asthma Immunol 83:559–566
Osafune K, Caron L, Borowiak M et al (2008) Marked differences in differentiation propensity among human embryonic stem cell lines. Nat Biotechnol 26:313–315
Cox JS (1967) Disodium cromoglycate (FPL 670) (‘Intal’): a specific inhibitor of reaginic antibody-antigen mechanisms. Nature 216:1328–1329
Theoharides TC, Sieghart W, Greengard P, Douglas WW (1980) Antiallergic drug cromolyn may inhibit histamine secretion by regulating phosphorylation of a mast cell protein. Science 207:80–82
Mazurek N, Berger G, Pecht I (1980) A binding site on mast cells and basophils for the anti-allergic drug cromolyn. Nature 286:722–723
Okada M, Itoh H, Hatakeyama T, Tokumitsu H, Kobayashi R (2003) Hsp90 is a direct target of the anti-allergic drugs disodium cromoglycate and amlexanox. Biochem J 374:433–441
Romanin C, Reinsprecht M, Pecht I, Schindler H (1991) Immunologically activated chloride channels involved in degranulation of rat mucosal mast cells. EMBO J 10:3603–3608
Reinsprecht M, Pecht I, Schindler H, Romanin C (1992) Potent block of Cl- channels by antiallergic drugs. Biochem Biophys Res Commun 188:957–963
Heinke S, Szücs G, Norris A, Droogmans G, Nilius B (1995) Inhibition of volume-activated chloride currents in endothelial cells by chromones. Br J Pharmacol 115:1393–1398
Jenkins L, Brea J, Smith NJ et al (2010) Identification of novel species-selective agonists of the G-protein-coupled receptor GPR35 that promote recruitment of β-arrestin-2 and activate Gα13. Biochem J 432:451–459
Yang Y, Lu JY, Wu X et al (2010) G-protein-coupled receptor 35 is a target of the asthma drugs cromolyn disodium and nedocromil sodium. Pharmacology 86:1–5
Hrvatin S, O’Donnell CW, Deng F et al (2014) Differentiated human stem cells resemble fetal, not adult, β cells. Proc Natl Acad Sci U S A 111:3038–3043
Goulley J, Dahl U, Baeza N, Mishina Y, Edlund H (2007) BMP4-BMPR1A signaling in β cells is required for and augments glucose-stimulated insulin secretion. Cell Metab 5:207–219
Szabat M, Johnson JD, Piret JM (2010) Reciprocal modulation of adult beta cell maturity by activin A and follistatin. Diabetologia 53:1680–1689
Saito H, Takeuchi M, Chida K, Miyajima A (2011) Generation of glucose-responsive functional islets with a three-dimensional structure from mouse fetal pancreatic cells and iPS cells in vitro. PLoS One 6:e28209
Logothetopoulos J (1972) Islet cell regeneration and neogenesis. In: Geiger SR (ed) Handbook of physiology. American Physiological Society, Washington, p 67
Bonner-Weir S, Weir GC (2005) New sources of pancreatic β-cells. Nat Biotechnol 23:857–861
Dor Y, Brown J, Martinez OI, Melton DA (2004) Adult pancreatic β-cells are formed by self-duplication rather than stem-cell differentiation. Nature 429:41–46
Wang S, Jensen JN, Seymour PA et al (2009) Sustained Neurog3 expression in hormone-expressing islet cells is required for endocrine maturation and function. Proc Natl Acad Sci U S A 16:9715–9720
Rezania A, Bruin JE, Xu J et al (2013) Enrichment of human embryonic stem cell-derived NKX6.1-expressing pancreatic progenitor cells accelerates the maturation of insulin-secreting cells in vivo. Stem Cells 31:2432–2442
Bruin JE, Rezania A, Xu J et al (2013) Maturation and function of human embryonic stem cell-derived pancreatic progenitors in macroencapsulation devices following transplant into mice. Diabetologia 56:1987–1998
Furuta Y, Hogan BL (1998) BMP4 is essential for lens induction in the mouse embryo. Genes Dev 12:3764–3775
Acknowledgements
The authors are grateful to S. Yamanaka (CiRA, Kyoto University, Kyoto, Japan), M. Uesugi (Institute for Integrated Cell-Material Sciences/Institute for Chemical Research, Kyoto University), T. Araoka (Salk Institute for Biological Studies, La Jolla, CA, USA), F. Shiota (CiRA, Kyoto University), M. Kotaka (CiRA, Kyoto University), K. Yasuda (CiRA, Kyoto University) and T. Kasahara (CiRA, Kyoto University) for technical support and helpful suggestions. Some of these data were presented as an abstract at the 51st Annual Meeting of the EASD (Stockholm, 2015) and the 7th Scientific Meeting of the Asian Association for the Study of Diabetes and the Annual Scientific Meeting of the Hong Kong Society of Endocrinology, Metabolism and Reproduction, in 2015.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Data availability
The RNA sequencing data generated during the current study are available in the Gene Expression Omnibus (www.ncbi.nlm.nih.gov/geo) with series accession number GSE89973. All other datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
Funding
This research was supported by the Japan Society for the Promotion of Science (JSPS) through its ‘Funding Program for World-Leading Innovative R&D on Science and Technology (FIRST Program)’; by the Japan Agency for Medical Research and Development (AMED) through its research grant ‘Core Center for iPS Cell Research, Research Center Network for Realization of Regenerative Medicine’; and by the Japan Diabetes Foundation.
Duality of interest
KO is a founder and member without salary of the scientific advisory boards of iPS Portal, Japan. All other authors declare that there is no duality of interest associated with this manuscript.
Contribution statement
All authors contributed to the study conception and design, acquisition of data or analysis and interpretation of data. AO, NI and KO supervised all aspects of the study. YK and KO drafted the manuscript. All authors participated in critical revision of the manuscript and approved the final version. KO is the guarantor of this work.
Electronic supplementary material
ESM 1
(PDF 377 kb)
Rights and permissions
About this article
Cite this article
Kondo, Y., Toyoda, T., Ito, R. et al. Identification of a small molecule that facilitates the differentiation of human iPSCs/ESCs and mouse embryonic pancreatic explants into pancreatic endocrine cells. Diabetologia 60, 1454–1466 (2017). https://doi.org/10.1007/s00125-017-4302-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-017-4302-7