Abstract
Aims/hypothesis
Skin fluorescence (SF) is a non-invasive marker of AGEs and is associated with the long-term complications of diabetes. SF increases with age and is also greater among individuals with diabetes. A familial correlation of SF suggests that genetics may play a role. We therefore performed parallel genome-wide association studies of SF in two cohorts.
Methods
Cohort 1 included 1,082 participants, 35–67 years of age with type 1 diabetes. Cohort 2 included 8,721 participants without diabetes, aged 18–90 years.
Results
rs1495741 was significantly associated with SF in Cohort 1 (p < 6 × 10−10), which is known to tag the NAT2 acetylator phenotype. The fast acetylator genotype was associated with lower SF, explaining up to 15% of the variance. In Cohort 2, the top signal associated with SF (p = 8.3 × 10−42) was rs4921914, also in NAT2, 440 bases upstream of rs1495741 (linkage disequilibrium r 2 = 1.0 for rs4921914 with rs1495741). We replicated these results in two additional cohorts, one with and one without type 1 diabetes. Finally, to understand which compounds are contributing to the NAT2–SF signal, we examined 11 compounds assayed from skin biopsies (n = 198): the fast acetylator genotype was associated with lower levels of the AGEs hydroimidazolones of glyoxal (p = 0.017).
Conclusions/interpretation
We identified a robust association between NAT2 and SF in people with and without diabetes. Our findings provide proof of principle that genetic variation contributes to interindividual SF and that NAT2 acetylation status plays a major role.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
AGEs accrue in the body during ageing, and their formation and accumulation are significantly accelerated in diabetes [1]. AGE formation is a multistep process in which the non-enzymatic glycation of proteins is followed by the formation of stable adducts and cross-links, leading to structural and functional tissue impairment [2]. Among people without diabetes, additive genetic effects explain 74% of the population variance of serum-determined AGEs, but the loci are unknown [3].
Skin collagen undergoes glycation and glycoxidation [1]. Given that skin collagen has a half-life of 10–15 years [4], skin AGEs capture decades-long glycaemia [5, 6]. AGEs in skin biopsies predict microvascular complications of type 1 diabetes, independent of HbA1c level [5–7]. However, skin biopsies are impracticable for large studies.
The measurement of skin fluorescence (SF) using optical spectroscopy, corrected for pigmentation, offers a non-invasive measurement of AGEs [8–10]. SF reflects glycaemia in type 1 and type 2 diabetes [8, 11–13] and can screen for abnormal glucose tolerance [14]. SF has been associated with macro- and microvascular complications in type 1 diabetes, independent of long-term HbA1c level [15–18]. In addition, SF is significantly associated with mortality from CHD in diabetes, independent of other risk factors [11, 19].
When adjusted for age and HbA1c level, SF is correlated (r = 0.43, p = 0.01) between siblings discordant for type 1 diabetes, suggesting genetic contributions [20]. A twin study has shown that lens fluorescence is significantly heritable [21]. However, the specific genetic determinants of SF are unknown. Given that SF is greater in people with than without diabetes [8], we performed genome-wide association studies (GWASs) in two parallel discovery cohorts to identify loci associated with SF in participants with type 1 diabetes and without diabetes.
Methods
Design
The first cohort included 1,082 participants with type 1 diabetes from the Diabetes Control and Complications Trial/Epidemiology of Diabetes Interventions and Complications (DCCT/EDIC) study (Table 1, electronic supplementary material [ESM] Table 1) [12]. The second cohort consisted of 8,721 participants without diabetes from the LifeLines Cohort Study (Table 2, ESM Table 2) [22]. Two additional cohorts contributed to replication of the results (ESM Tables 3–5). The first of these cohorts included participants with type 1 diabetes from the Wisconsin Epidemiologic Study of Diabetic Retinopathy (WESDR, n = 202) [23] and the second included an older cohort of participants from the LonGenity study (n = 515) [24], consisting primarily of participants without diabetes. All individuals were of European descent. These studies were approved by the institutional review boards of all the participating institutions, and informed consent was obtained from all the participants.
Discovery cohort 1: DCCT/EDIC
Participants
At a point 16–17 years after the close-out of the DCCT [25], SF was assessed in 1,082 active participants who had GWAS data [12].
Measurement of SF
Skin intrinsic fluorescence (SIF), corrected for factors that affect light scattering and absorption, was measured from the underside of the left forearm using the SCOUT DS SF spectrometer (VeraLight, Inc., Albuquerque, NM, USA) [12, 15]. SIF excited with a light-emitting diode (LED) centred at 375 nm, and emission detected over 435–655 nm (with the reflectance adjusted by the dimensionless exCitation and emission exponents, k x = 0.6, k m = 0.2, respectively), referred to as SIF1, was the primary outcome [15, 18]. In secondary analyses, we examined SIFs measured using excitation LEDs centred at 405 nm, 416 nm, 435 nm and 456 nm (SIF2–SIF15; ESM Table 6).
Discovery cohort 2: LifeLines
Participants
The LifeLines observational follow-up study includes a random sample of 165,000 inhabitants of three northern provinces of the Netherlands, who have been recruited since 2006 [22]. For this analysis, we included participants 18–90 years of age who had had both GWAS and AGE Reader (DiagnOptics Technologies BV, Groningen, the Netherlands) measurement of skin autofluorescence (SAF) collected from 2006 to 2012. For the GWAS, we excluded participants who were known to have type 1 (n = 12) or type 2 diabetes and/or had a fasting blood plasma glucose level >7.0 mmol/l (n = 318), leaving 8,721 individuals. We also examined participants with type 2 diabetes (Table 2), either previously known (n = 216) or newly diagnosed by fasting blood plasma glucose level (>7.0 mmol/l, n = 102) in secondary analyses.
Measurement of SF
SAF was measured with the AGE Reader [8, 13]. Similar to SIF1, the excitation light source used a peak at 370 nm. The spectrometer measures emitted and reflected light over 420–600 nm and 300–420 nm, respectively. AGE Reader software (v2.3) calculates SAF from the ratio between the emitted and the reflected light, multiplied by 100 (expressed as arbitrary units [AU]), taking skin colour into account [26]. For each SAF value, measurements were carried out at three different sites of the same forearm, and the mean was used for the analysis.
Skin collagen ancillary substudy
Skin biopsies were obtained on a subset of DCCT participants (n = 216) at close-out, and 11 AGEs and collagen cross-linking variables were determined [5, 6].
Genotyping
The Illumina 1 M beadchip assay (Illumina, San Diego, CA, USA), which underwent extensive quality control [27], was used in the DCCT/EDIC cohort for genome-wide genotyping. Quality control checks were applied to ensure that there were no sample mix-ups, and participants were excluded if they were determined to be admixed using population genetic approaches [28]. A total of 841,342 autosomal single nucleotide polymorphisms (SNPs) with a minor allele frequency (MAF) >1% were subsequently analysed. In addition, 1,609,583 (INFO ≥ 0.30) imputed autosomal SNPs using release 22 Phase II Centre d'Etude du Polymorphisme (Utah residents with northern and western European ancestry) (CEU) HapMap data (IMPUTE v2; https://mathgen.stats.ox.ac.uk/impute/impute.html) were used for analysis.
In the LifeLines cohort, Illumina CytoSNP 12v2 was used for genome-wide genotyping. Genotyped SNPs with an MAF <0.01, a call rate ≤95% and a Hardy–Weinberg equilibrium (HWE) p value <10−4 were excluded. A total of 837,184 genotyped or imputed SNPs (allelic R 2 ≥ 0.80) using HapMap (release 24) CEU (Beagle, v3.1.0) were used for GWAS. Sample relatedness was assessed by participant self-report and population genetic approaches, and the sample with the highest genotyping quality was included for first-degree relatives. Samples were further excluded due to discrepancy in sex, an average heterozygosity >4SD from the mean and non-European ancestry assessed using identical-by-state analysis and Eigenstrat (http://genetics.med.harvard.edu/reich/Reich_Lab/Software.html) [29].
Statistical analysis
GWAS
The primary outcome variable for GWAS in the DCCT/EDIC group was loge SIF1. Since genetic variants may be associated with SF through effects on factors associated with SF, such as smoking or HbA1c level, we first performed a univariate GWAS of SIF1 using genotyped SNPs by linear regression (Model 1 [M1]). To increase the power to detect loci associated with SIF1 by explaining additional variance in the trait, a second GWAS adjusted for age, sex, smoking status, skin tone, clinic latitude and ever having had an estimated GFR (eGFR) <60 ml/min/1.73 m2 (Model 2 [M2]). Finally, a third GWAS was conducted adjusting for the same covariates in M2 with the addition of HbA1c measured as follows: at the DCCT eligibility screening, as the mean during the DCCT and as the mean during the EDIC study (M3). DCCT/EDIC Model 3 [M3] was also analysed using imputed SNPs as dosages. To determine whether there were any independent signals, a GWAS conditioning on the top SNP observed from the GWAS was conducted using M1.
In the LifeLines cohort, the primary outcome variable for GWAS was SAF. We first performed an analysis without covariates (M1). Next, we corrected for age, sex and smoking status (M2). Finally, we added BMI, fasting plasma glucose level, HbA1c level, Cockcroft–Gault eGFR and principal components (PCs) to M2 (M3). An additive model was used in all models. A p-value <5 × 10−8 was required for genome-wide significance in each discovery cohort.
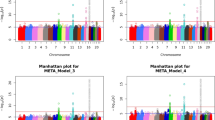
Using METAL (v2011-03-25; www.sph.umich.edu/csg/abecasis/Metal/download/), we also ran a meta-GWAS for M3 using genotyped or imputed SNPs from the DCCT/EDIC and LifeLines studies using weights proportional to √n on z scores [30].
Characterisation of rs1495741 in the DCCT/EDIC
Further analyses were conducted using DCCT/EDIC M1 including adjustment for PCs, and testing for deviation from an additive genetic model by including a heterozygous indicator to a model with rs1495741 coded additively. We also examined the association of rs1495741 with SIF2–SIF15. The heterogeneity of rs1495741 effects was assessed by including rs1495741–covariate interactions. rs1495741 was tested for association with glycaemia measured by HbA1c level during the DCCT and EDIC studies, and with capillary glucose during the DCCT, using linear regression adjusted for DCCT treatment group. The association with the presence of diabetes complications was tested using logistic regression adjusted for covariates in M2 and M3 (each with and without adjustment for any eGFR <60 ml/min/1.73 m2). Logistic regression was also used to test rs1495741 for an association with the risk of hypoglycaemia during the DCCT, comparing participants with one or more episodes of hypoglycaemia requiring assistance with those without, and similarly comparing participants with hypoglycaemia resulting in coma with those without. Linear mixed models were used to determine the association with lipids measured annually during the DCCT.
Skin biopsy ancillary substudy in the DCCT cohort
Multiple linear regression adjusting for age, duration of diabetes and randomised treatment group in the DCCT was used to determine whether rs1495741 was associated with AGEs measured from skin biopsies [5, 6].
Characterisation of rs4921914 in the LifeLines cohort
rs4921914 was tested for an association with HbA1c using linear regression adjusted for age, sex, smoking status, BMI, Cockcroft–Gault eGFR, fasting plasma glucose level and PCs. To test the association with fasting plasma glucose, the same analysis was used, but was adjusted for HbA1c instead of fasting plasma glucose level. The association of rs4921914 with lipid variables was adjusted for all the above covariates with lipid levels adjusted for statin use [31].
Testing rs4921914 among type 2 diabetes in the LifeLines cohort
Linear regression was used to test whether rs4921914 was associated with SAF in 318 participants with type 2 diabetes in the LifeLines study. To determine whether the association for rs4921914 differed between participants with type 2 diabetes (n = 318) and those without diabetes (n = 8,721), we added an rs4921914–diabetes interaction.
PLINK (v1.07) (http://pngu.mgh.harvard.edu/~purcell/plink/) [32] was used for GWASs of genotyped SNPs in DCCT/EDIC, PLINK (v1.90alpha) in LifeLines, R (v2.15.2; www.r-project.org/) for imputed SNPs and for the generation of plots and calculation of genomic control lambda (GenABEL; www.genabel.org/). SAS (v9.2; Cary, NC, USA) and STATA (v.11; College Station, TX, USA) were used for all analyses at the top SNP in the DCCT/EDIC and LifeLines cohorts, respectively.
Results
Discovery GWAS 1: DCCT/EDIC
The GWAS of SIF1 from the DCCT/EDIC cohort identified a significant association with rs1495741 (p = 6.1 × 10−10; Table 3), 14 kb downstream of NAT2 (Fig. 1a, ESM Fig. 1a). rs1495741 is associated with the in vitro measured NAT2 acetylator phenotype and tags haplotypes that define the acetylator status with 99% sensitivity and 95% specificity in white individuals [33–35]. Examination of the rs1495741 allele intensity cluster plot showed a clear separation of genotypes (ESM Fig. 2, G-allele MAF = 0.22, chromosome 8p22, build 36 position 18,317,161), with no significant deviation from HWE (p = 0.75). The association with SIF1 was not materially different after adjusting for the first three PCs (p for M1 = 5.5 × 10−10). Upon adjusting for covariates in M2 and M3, no additional genome-wide significant loci were identified (ESM Fig. 1b, c, ESM Fig. 3, Table 3, GWAS results available from dbGaP: ftp://ftp.ncbi.nlm.nih.gov/dbgap/studies/phs000086/analyses/). Results from an analysis conditional on rs1495741 did not identify any independent signals that were significant genome-wide (data not shown). SIF1 was lower with each copy of the fast acetylator G-allele (Fig. 1b) and there was no deviation from an additive model (p = 0.69). Results from M1 show that rs1495741 explains 3.5% of the variance in SIF1, and explains an additional 3% of the variance when added to the covariates in M3, which explains 33% of the variance in SIF1 [12].
(a) Regional plot of a 300 kb region surrounding rs1495741 (p = 1.7 × 10−12) showing genotyped and imputed SNPs plotted with their (–log10) p values from the DCCT/EDIC cohort (M3) on the left y-axis and their genomic position (NCBI Build 35; www.ncbi.nlm.nih.gov/mapview/stats/BuildStats.cgi?taxid=9606&build=3) on the x-axis. Gene annotations (Genome Browser; http://genome.ucsc.edu/) are shown above the x-axis. Estimated recombination rates (HapMap II release 22; http://hapmap.ncbi.nlm.nih.gov/) are plotted on the right y-axis. For genotyped SNPs, the LD values shown were calculated based on pairwise r 2 values for rs1495741 from the DCCT/EDIC cohort, and for imputed SNPs are based on r 2 values from HapMap phase II (Nov08, release 24, on NCBI B36 assembly, dbSNP b126). The blue diamond indicates rs1495741 and the SNPs are coloured based on their LD with it (red, r 2 ≥ 0.8; orange, 0.5 ≤ r 2 < 0.8; yellow, 0.2 ≤ r 2 < 0.5; white, r 2 < 0.2); (www.broadinstitute.org/diabetes/scandinavs/figures.html). (b) Beeswarm plot showing level of unadjusted log e SIF1 for each participant in the DCCT/EDIC cohort according to their rs1495741 genotype, with the mean ± SD shown above the x-axis
We also examined the association for rs1495741 with SIF2–SIF15 and observed the strongest relationship for SIF12 (p for M3 = 2.9 × 10−49), explaining 15.0% of the variance (Table 3, ESM Table 6). Importantly, rs1495741 was still significantly associated with SIF1 (p = 4.04 × 10−19) after adjusting for SIF12, suggesting that the effect on SIF1 is independent of the effect on SIF12.
Discovery GWAS 2: LifeLines
The top SAF association signals observed in the LifeLines cohort were 47 SNPs that were also in the NAT2 region (M3; p < 5 × 10−8) (Table 4, ESM Figs 4 and 5). The top signal (LifeLines M3, p = 1.0 × 10−60, R 2 = 2.1%) was rs4921914, 440 bases upstream of rs1495741 (ESM Fig. 6). According to HapMap phase 2 CEU (release 22), rs4921914 is in perfect linkage disequilibrium (LD; r 2 = 1.0) with rs1495741 (C- and G-alleles, respectively). rs4921914 was imputed with allelic R 2 = 0.81 (MAF = 0.19 for the C-allele) and showed no deviation from HWE (p = 0.36). The association was in the same direction as observed in the DCCT/EDIC group.
We did not identify any additional genome-wide significant loci after meta-GWAS (ESM Figs 7–8).
Heterogeneity of rs1495741 effect on SF
In the DCCT/EDIC cohort, rs1495741 was not significantly associated with any of the covariates in M2 (p > 0.05, data not shown). There was no heterogeneity of rs1495741 on SIF1 (ESM Table 7), with the exception of the updated weighted mean HbA1c levels for the DCCT/EDIC, which approached nominal significance (p for the SNP*HbA1c interaction = 0.058). When analysing the effect of rs1495741 separately by the median updated weighted mean HbA1c level for the DCCT/EDIC (8.07%), rs1495741 was stronger among participants with HbA1c levels greater than median (β±SE = −0.075 ± 0.01, p = 4.7 × 10−7) compared with those below the median (β±SE = −0.054 ± 0.01, p = 1.2 × 10−4). In the LifeLines group, no significant SNP × HbA1c interaction was observed among the non-diabetic participants (p = 0.28). However, there was a significant heterogeneity of rs4921914 by age (p = 0.03).
Biochemical measures and complications of type 1 diabetes
In the DCCT/EDIC cohort, rs1495741 was not associated with updated weighted mean HbA1c up to the time of SIF measurement (p = 0.95; Table 3). In participants without diabetes in the LifeLines cohort, rs4921914 was also not associated with HbA1c (p = 0.36) or fasting plasma glucose (p = 0.36) level (Table 4). Furthermore, rs1495741 was not associated with the mean of 7-point capillary blood glucose profiles measured during the DCCT (p = 0.23). In the DCCT/EDIC cohort, rs1495741 was not associated with any microvascular complications or markers of macrovascular complications in models adjusted for M2 and/or M3 covariates (either with or without adjusting for any eGFR<60 ml/min/1.73 m2), or with hypoglycaemia (Table 3).
Since rs1495741 has been associated with lipids [36], we examined its association with repeated measures of lipids during the DCCT. rs1495741 was nominally associated with triacylglycerol (p = 0.01) and HDL-cholesterol (p = 0.008) values (Table 3). The effect of rs1495741 on SIF1 was unchanged after adjusting for time-weighted mean triacylglycerol (p = 1.8 × 10−12) or HDL-cholesterol (p = 9.1 × 10−13) on M3. In the LifeLines cohort, no association could be demonstrated (Table 4), although the power to detect an effect on triacylglycerol and total cholesterol [36] was 61% and 31%, respectively.
Skin biopsy ancillary substudy in the DCCT cohort
Of the 11 AGEs and collagen cross-linking variables measured in the skin biopsies, rs1495741 was associated only with hydroimidazolones of glyoxal (G-H1; p = 0.017; Table 3), in the same direction as the association with SF.
SAF among individuals with type 2 diabetes in the LifeLines cohort
rs4921914 was significantly associated (β±SE = −0.20 ± 0.06, p = 5.2 × 10−4) with SAF in participants with type 2 diabetes and explained 3.4% of the variance in SAF. The difference in the SNP effect between the participants with and without type 2 diabetes was p = 0.07, with a stronger association observed in those with type 2 diabetes.
Replication in the WESDR and LonGenity cohorts
In a second cohort of type 1 diabetes participants, rs1495741 (G-allele MAF = 25%, HWE p = 0.51) was associated with SIF1 (β±SE = −0.06 ± 0.02, p for the multivariate model = 0.002, r 2 for the univariate model = 5.3%), in the same direction of effect as in the discovery cohorts (Table 5). The effect of rs1495741 on SIF1 did not differ according to smoking status (p = 0.54). rs1495741 was also associated with SIF14 (β±SE = 0.10 ± 0.02, p for the multivariate model = 6.0 × 10−5, r 2 for the univariate model = 9.2%). Finally, rs1495741 was not associated with repeated measures of HbA1c in linear mixed models (p = 0.53), or time to mild (p = 0.28) or severe diabetic retinopathy (p = 0.18).
rs1495741 (G-allele MAF = 18%, HWE p = 0.92) was not significantly associated with SIF1 (β±SE = −0.01 ± 0.007, p = 0.09; Table 5) in LonGenity, an older cohort (mean ± SD age 75 ± 6 years) of participants predominantly without diabetes (89%). However, consistent with the DCCT/EDIC and WESDR cohorts, rs1495741 was associated with SIF14 (β±SE = −0.02 ± 0.007, p = 0.0004) in the same direction.
Discussion
Most GWASs of quantitative traits use blood-based measures, for which levels may vary widely within a person over time due to clearance predominantly by the kidneys and liver. SF, on the other hand, provides a measure of long-term tissue damage associated with ageing [8, 9] as well as with decades-long blood glucose levels in people with diabetes [12], given the long half-life of skin collagen [4]. Using a GWAS, we identified rs1495741 located 14 kb downstream of NAT2 to be associated with SF in individuals with type 1 diabetes, and the same signal (rs4921914, r 2 = 1.0) was observed in a separate discovery cohort of individuals without diabetes. We further replicated the association for SF in individuals with type 1 and 2 diabetes. In addition, the signal observed for rs1495741 with SIF14 was also replicated in the WESDR and LonGenity populations. In the DCCT/EDIC cohort, rs1495741 explains 3.5% of the variance in SIF1 and appears to be tagging NAT2, with no signals observed in the neighbouring NAT1. In all four studies, each copy of the fast acetylator allele was associated with less SF.
The association of NAT2 with SF was robust across different populations and different measures of SF both within and across studies. First, we observed the association in individuals both with and without type 2 diabetes in the LifeLines group, as well as in those with type 1 diabetes in the DCCT/EDIC and WESDR cohorts. There was suggestion that a stronger effect of NAT2 on SF might exist in individuals with type 2 diabetes than in those without diabetes. Similarly, the effect of NAT2 on SIF was stronger in participants who had higher HbA1c levels in comparison to those with lower HbA1c levels in the DCCT/EDIC population. However, no significant SNP*HbA1c interaction was observed among non-diabetic participants in the LifeLines group. Second, NAT2 was strongly associated with SF excited across the range of 375–456 nm in the DCCT/EDIC group, with the strongest effect observed for excitation at 435 nm (SIF12). Finally, NAT2 was the top signal observed in both discovery GWAS cohorts, despite using different SF devices. Indeed, both the SCOUT DS and the AGE Reader, used in the LifeLines study, have been shown to correlate with skin biopsy-determined AGEs such as pentosidine [8, 9].
Although SF has been associated with HbA1c level in people with diabetes [8, 11, 12], rs1495741 was not associated with glycaemia in people either with or without diabetes in our cohorts. However, in the Meta-Analyses of Glucose and Insulin-related traits Consortium (MAGIC; http://www.magicinvestigators.org), a large meta-GWAS of more than 46,000 non-diabetic adults of European descent, the G-allele of rs1495741 was associated with higher HbA1c (p = 0.003) [37] and fasting plasma glucose (p = 0.03) values [38]. Both associations were in the opposite direction to the effect of NAT2 on SF (ESM Table 8). Thus, a small effect for rs1495741 on glycaemic traits in non-diabetic individuals may exist and may be observed only in very large analyses.
Despite the association of SF with the complications of type 1 diabetes [15–18, 39], rs1495741 was not associated with coronary artery calcium, nephropathy, neuropathy or retinopathy in the DCCT/EDIC group, or with time from baseline to either mild or severe retinopathy in the WESDR cohort. The non-significant association with these outcomes is due to the low power to detect small effects, since we have good power for detecting an OR >1.50 for a sustained AER >30 mg/24 h with α = 0.05 [15]. In line with this, the rs1495741 G-allele has previously been associated (OR 1.06, p = 2 × 10−5) with an increased risk of coronary artery disease (CAD) [36].
Of the 11 AGEs and collagen cross-linking variables assayed, rs1495741 was nominally associated with G-H1 in the same direction as with SF. G-H1 is a hydroimidazolone derived from arginine residues modified by glyoxal, a potent glycating agent [40]. G-H1 itself is not fluorescent, but its reactive AGE precursor, glyoxal, can participate in the formation of fluorescent structures such as vesperlysine A, which is detected at 370 nm/440 nm excitation/emission [41]. In the DCCT, G-H1 was positively associated with age, but not with duration of diabetes or with recent or long-term HbA1c levels [6]. Finally, on its own, G-H1 was not significantly associated with risk of any of the microvascular complications in the DCCT [6]. Although G-H1 was not correlated with SIF1 measured 15 years later (ESM Table 9), this may be due to its half-life of 2–6 weeks [40]. Replication of the association of NAT2 with G-H1 is needed in individuals with diabetes, as is a determination of the association in those without diabetes.
We used SIF1 for our GWAS in the DCCT/EDIC cohort based on previous observations that SIF1 had the strongest association with complications [15, 18] and was closest to the excitation wavelength used in the LifeLines study. However, SF measures can be obtained from a wide range of excitation/emission spectra, which reflect different fluorophores [8, 42]. For example, SF measured with 370 nm/440 nm excitation/emission has been shown to capture AGEs in general and has been shown to correlate with pentosidine levels determined by skin biopsy [42]. The less studied 440 nm/520 nm excitation/emission fluorescence, which closely corresponds with SIF12, correlated with pentosidine as well as with N ε-carboxymethyl-lysine, carboxyethyl-lysine and methionine sulphoxide [42]. However, NAT2 was not associated with these AGEs in the DCCT skin biopsy substudy. Thus, our observation of a stronger association for NAT2 with SIF12 in the DCCT/EDIC cohort may also reflect other AGEs that are possibly associated with elastin–collagen cross-links detected in response to excitation at higher wavelengths [42]. SIF12, however, also coincides with non-AGE fluorophores including flavin adenine dinucleotide and oxidised flavin mononucleotide [10], as well as phospholipids, and therefore the NAT2 signal for SIF12 may occur through non-AGE mechanisms [43]. Follow-up studies are needed to determine the compounds responsible for the association between NAT2 and SF.
N-Acetyltransferase 2 (NAT2) is known to metabolise drugs and carcinogens, but no known endogenous substrates have been identified [44]. A previous GWAS identified an association of rs1495741 with bladder cancer risk, with significant evidence for a gene-by-smoking interaction [45]. However, we observed no such heterogeneity of effect for rs1495741 on SIF1 by smoking status. NAT2 has also been identified in several other GWASs, including those involving plasma [46] and urine metabolites [47] as well as lipids [36]. In both the previous meta-GWAS [36] and the DCCT, the fast acetylator G-allele was associated with higher triacylglycerol levels. Unlike the previous study of primarily individuals without diabetes [36], rs1495741 was associated with lower HDL-cholesterol values among fast acetylators in the DCCT cohort (Table 3, ESM Table 10). Importantly, the effect of rs1495741 on SIF1 was unchanged after adding time-weighted mean triacylglycerol or HDL-cholesterol to M3, arguing that the genetic effect on SF is not mediated by triacylglycerol or HDL-cholesterol. The mechanism(s) linking the fast NAT2 acetylator genotype with both elevated triacylglycerol levels and CAD is unknown [36]. However, the opposing direction of effect of rs1495741 on SF in comparison to triacylglycerol, CAD and possibly glycaemic traits [37, 38] suggests that multiple underlying pathways may exist.
The LifeLines study used the AGE Reader to measure SAF with a peak excitation of 370 nm, compared with SIF1 measured in the DCCT/EDIC study using the SCOUT DS with an excitation peak at 375 nm, and different corrections were made for skin pigmentation. Despite this limitation, the NAT2 region was robustly the top signal associated with both SIF1 in the DCCT/EDIC and SAF in the LifeLines cohort. Accordingly, in the DCCT/EDIC cohort, a correlation of r pearson = 0.69 (p 10−15) was observed for SIF1 with a proxy for SAF (LED 375 nm, k x = 1.0, k m = 0.0), showing that SIF and SAF are comparable measurements. NAT2 was not significantly associated with SIF1 in the LonGenity study (p = 0.09), possibly due to the age of this cohort, given that collagen production is altered with older age [48]. However, it was associated with SIF14. Although NAT2 activity and protein levels have not been detected in normal human dermal fibroblasts [49], the metabolism of substrates by NAT2 in the liver or colon may contribute to the differences in SF between slow and fast acetylators [44, 50]. Finally, since not all AGEs fluoresce, a GWAS of SF may not identify loci specific to the production or detoxification of non-fluorescent AGEs, unless they are highly correlated with a fluorophore in the skin.
In conclusion, using a GWAS, our study demonstrates that genetic variation contributes to SF, and that NAT2 is a major locus observed across four independent studies. Although NAT2 was not significantly associated with the risk of complications of type 1 diabetes, larger studies are needed. NAT2 may improve the screening properties of SF in predicting the risk of complications or impaired glucose tolerance [14–18]. The robust association observed for NAT2 with SF in multiple cohorts provides proof of principle that genetic variation contributes to the variance in SF.
Abbreviations
- AU:
-
Arbitrary units
- CAD:
-
Coronary artery disease
- EDIC:
-
Epidemiology of Diabetes Interventions and Complications
- eGFR:
-
Estimated GFR
- G-H1:
-
Hydroimidazolones of glyoxal
- GWAS:
-
Genome-wide association study
- HWE:
-
Hardy–Weinberg equilibrium
- LD:
-
Linkage disequilibrium
- LED:
-
Light-emitting diode
- M1:
-
Model 1
- M2:
-
Model 2
- M3:
-
Model 3
- MAF:
-
Minor allele frequency
- NAT2:
-
N-Acetyltransferase 2
- PC:
-
Principal component
- SAF:
-
Skin autofluorescence
- SF:
-
Skin fluorescence
- SIF:
-
Skin intrinsic fluorescence
- SNP:
-
Single nucleotide polymorphism
- UKPDS:
-
UK Prospective Diabetes Study
- WESDR:
-
Wisconsin Epidemiologic Study of Diabetic Retinopathy
References
Dyer DG, Dunn JA, Thorpe SR et al (1993) Accumulation of Maillard reaction products in skin collagen in diabetes and aging. J Clin Invest 91:2463–2469
Brownlee M, Cerami A, Vlassara H (1988) Advanced glycosylation end products in tissue and the biochemical basis of diabetic complications. N Engl J Med 318:1315–1321
Leslie RD, Beyan H, Sawtell P, Boehm BO, Spector TD, Snieder H (2003) Level of an advanced glycated end product is genetically determined: a study of normal twins. Diabetes 52:2441–2444
Verzijl N, DeGroot J, Thorpe SR et al (2000) Effect of collagen turnover on the accumulation of advanced glycation end products. J Biol Chem 275:39027–39031
Monnier VM, Bautista O, Kenny D et al (1999) Skin collagen glycation, glycoxidation, and crosslinking are lower in subjects with long-term intensive versus conventional therapy of type 1 diabetes: relevance of glycated collagen products versus HbA1c as markers of diabetic complications. DCCT Skin Collagen Ancillary Study Group, Diabetes Control and Complications Trial. Diabetes 48:870–880
Monnier VM, Sell DR, Strauch C et al (2013) The association between skin collagen glucosepane and past progression of microvascular and neuropathic complications in type 1 diabetes. J Diabetes Complications 27:141–149
Genuth S, Sun W, Cleary P et al (2005) Glycation and carboxymethyllysine levels in skin collagen predict the risk of future 10-year progression of diabetic retinopathy and nephropathy in the Diabetes Control and Complications Trial and Epidemiology of Diabetes Interventions and Complications participants with type 1 diabetes. Diabetes 54:3103–3111
Meerwaldt R, Graaff R, Oomen PH et al (2004) Simple non-invasive assessment of advanced glycation endproduct accumulation. Diabetologia 47:1324–1330
Hull E, Ediger M, Unione A, Deemer E, Stroman M, Baynes J (2004) Noninvasive, optical detection of diabetes: model studies with porcine skin. Opt Express 12:4496–4510
Richards-Kortum R, Sevick-Muraca E (1996) Quantitative optical spectroscopy for tissue diagnosis. Annu Rev Phys Chem 47:555–606
Meerwaldt R, Lutgers HL, Links TP et al (2007) Skin autofluorescence is a strong predictor of cardiac mortality in diabetes. Diabetes Care 30:107–112
Cleary PA, Braffett BH, Orchard T et al (2013) Clinical and technical factors associated with skin intrinsic fluorescence in subjects with type 1 diabetes from the Diabetes Control and Complications Trial/Epidemiology of Diabetes Interventions and Complications Study. Diabetes Technol Ther 15:466–474
Koetsier M, Lutgers HL, de Jonge C, Links TP, Smit AJ, Graaff R (2010) Reference values of skin autofluorescence. Diabetes Technol Ther 12:399–403
Maynard JD, Rohrscheib M, Way JF, Nguyen CM, Ediger MN (2007) Noninvasive type 2 diabetes screening: superior sensitivity to fasting plasma glucose and A1C. Diabetes Care 30:1120–1124
Orchard TJ, Lyons TJ, Cleary PA et al (2013) The association of skin-intrinsic fluorescence with type 1 diabetes complications in the DCCT/EDIC Study. Diabetes Care 36:3146–3153
Conway BN, Aroda VR, Maynard JD et al (2011) Skin intrinsic fluorescence correlates with autonomic and distal symmetrical polyneuropathy in individuals with type 1 diabetes. Diabetes Care 34:1000–1005
Conway BN, Aroda VR, Maynard JD et al (2012) Skin intrinsic fluorescence is associated with coronary artery disease in individuals with long duration of type 1 diabetes. Diabetes Care 35:2331–2336
Conway B, Edmundowicz D, Matter N, Maynard J, Orchard T (2010) Skin fluorescence correlates strongly with coronary artery calcification severity in type 1 diabetes. Diabetes Technol Ther 12:339–345
Lutgers HL, Gerrits EG, Graaff R et al (2009) Skin autofluorescence provides additional information to the UK Prospective Diabetes Study (UKPDS) risk score for the estimation of cardiovascular prognosis in type 2 diabetes mellitus. Diabetologia 52:789–797
Barat P, Cammas B, Lacoste A et al (2012) Advanced glycation end products in children with type 1 diabetes: family matters? Diabetes Care 35:e1
Kessel L, Hougaard JL, Sander B, Kyvik KO, Sorensen TI, Larsen M (2002) Lens ageing as an indicator of tissue damage associated with smoking and non-enzymatic glycation—a twin study. Diabetologia 45:1457–1462
Stolk RP, Rosmalen JG, Postma DS et al (2008) Universal risk factors for multifactorial diseases: LifeLines: a three-generation population-based study. Eur J Epidemiol 23:67–74
Klein R, Knudtson MD, Lee KE, Gangnon R, Klein BE (2009) The Wisconsin Epidemiologic Study of Diabetic Retinopathy XXIII: the twenty-five-year incidence of macular edema in persons with type 1 diabetes. Ophthalmology 116:497–503
Han J, Ryu S, Moskowitz DM et al (2013) Discovery of novel non-synonymous SNP variants in 988 candidate genes from 6 centenarians by target capture and next-generation sequencing. Mech Ageing Dev 134:478–485
Epidemiology of Diabetes Interventions and Complications (EDIC) Research Group (1999) Design, implementation, and preliminary results of a long-term follow-up of the Diabetes Control and Complications Trial cohort. Diabetes Care 22:99–111
Koetsier M, Nur E, Chunmao H et al (2010) Skin color independent assessment of aging using skin autofluorescence. Opt Express 18:14416–14429
Paterson AD, Waggott D, Boright AP et al (2010) A genome-wide association study identifies a novel major locus for glycemic control in type 1 diabetes, as measured by both A1C and glucose. Diabetes 59:539–549
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. PLoS Genet 2:e190
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Willer CJ, Li Y, Abecasis GR (2010) METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26:2190–2191
Tobin MD, Sheehan NA, Scurrah KJ, Burton PR (2005) Adjusting for treatment effects in studies of quantitative traits: antihypertensive therapy and systolic blood pressure. Stat Med 24:2911–2935
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Garcia-Closas M, Hein DW, Silverman D et al (2011) A single nucleotide polymorphism tags variation in the arylamine N-acetyltransferase 2 phenotype in populations of European background. Pharmacogenet Genomics 21:231–236
He YJ, Shapero MH, McLeod HL (2012) Novel tagging SNP rs1495741 and 2-SNPs (rs1041983 and rs1801280) yield a high prediction of the NAT2 genotype in HapMap samples. Pharmacogenet Genomics 22:322–324
Selinski S, Blaszkewicz M, Lehmann ML et al (2011) Genotyping NAT2 with only two SNPs (rs1041983 and rs1801280) outperforms the tagging SNP rs1495741 and is equivalent to the conventional 7-SNP NAT2 genotype. Pharmacogenet Genomics 21:673–678
Teslovich TM, Musunuru K, Smith AV et al (2010) Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466:707–713
Soranzo N, Sanna S, Wheeler E et al (2010) Common variants at 10 genomic loci influence hemoglobin A(1)(C) levels via glycemic and nonglycemic pathways. Diabetes 59:3229–3239
Manning AK, Hivert MF, Scott RA et al (2012) A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet 44:659–669
Chabroux S, Canoui-Poitrine F, Reffet S et al (2010) Advanced glycation end products assessed by skin autofluorescence in type 1 diabetics are associated with nephropathy, but not retinopathy. Diabetes Metab 36:152–157
Thornalley PJ (2008) Protein and nucleotide damage by glyoxal and methylglyoxal in physiological systems—role in ageing and disease. Drug Metabol Drug Interact 23:125–150
Tessier F, Obrenovich M, Monnier VM (1999) Structure and mechanism of formation of human lens fluorophore LM-1. Relationship to vesperlysine A and the advanced Maillard reaction in aging, diabetes, and cataractogenesis. J Biol Chem 274:20796–20804
Beisswenger PJ, Howell S, Mackenzie T, Corstjens H, Muizzuddin N, Matsui MS (2012) Two fluorescent wavelengths, 440(ex)/520(em) nm and 370(ex)/440(em) nm, reflect advanced glycation and oxidation end products in human skin without diabetes. Diabetes Technol Ther 14:285–292
Ramanujam N (2000) Fluorescence spectroscopy in vivo. In: Meyers RA (ed) Encyclopedia of analytical chemistry. Wiley, Chichester, pp 20–56
Sabbagh A, Marin J, Veyssiere C et al (2013) Rapid birth-and-death evolution of the xenobiotic metabolizing NAT gene family in vertebrates with evidence of adaptive selection. BMC Evol Biol 13:62
Rothman N, Garcia-Closas M, Chatterjee N et al (2010) A multi-stage genome-wide association study of bladder cancer identifies multiple susceptibility loci. Nat Genet 42:978–984
Suhre K, Shin SY, Petersen AK et al (2011) Human metabolic individuality in biomedical and pharmaceutical research. Nature 477:54–60
Suhre K, Wallaschofski H, Raffler J et al (2011) A genome-wide association study of metabolic traits in human urine. Nat Genet 43:565–569
Varani J, Dame MK, Rittie L et al (2006) Decreased collagen production in chronologically aged skin: roles of age-dependent alteration in fibroblast function and defective mechanical stimulation. Am J Pathol 168:1861–1868
Bhaiya P, Roychowdhury S, Vyas PM, Doll MA, Hein DW, Svensson CK (2006) Bioactivation, protein haptenation, and toxicity of sulfamethoxazole and dapsone in normal human dermal fibroblasts. Toxicol Appl Pharmacol 215:158–167
Hickman D, Pope J, Patil SD et al (1998) Expression of arylamine N-acetyltransferase in human intestine. Gut 42:402–409
Acknowledgements
Industry contributors have had no role in the DCCT/EDIC study but the following have provided free or discounted supplies or equipment to support participants’ adherence to the study: Abbott Diabetes Care (Alameda, CA, USA), Animas (Westchester, PA, USA), Bayer Diabetes Care (North America Headquarters, Tarrytown, NY, USA), Becton Dickinson (Franklin Lakes, NJ, USA), CanAm (Atlanta, GA, USA), Eli Lilly (Indianapolis, IN, USA), Lifescan (Milpitas, CA, USA), Medtronic Diabetes (Minneapolis, MI, USA), Nova Diabetes Care (Billerica, MA, USA), OmniPod® Insulin Management System (Bedford, MA, USA), Omron (Shelton, CT, USA), Roche Diabetes Care (Indianapolis, IN, USA) and Sanofi-Aventis (Bridgewater, NJ, USA).
We would like to acknowledge the DCCT/EDIC and the LifeLines Cohort Study and Biobank study participants, staff and participating general practitioners and pharmacists. We would like to thank C. Strauch (Department of Pathology, Case Western Reserve University, Cleveland, OH, USA) for his assistance with AGE determination.
Funding
The DCCT/EDIC has been supported by U01 Cooperative Agreement grants (1982–93, 2011–2016) and contracts (1982–2011) with the Division of Diabetes Endocrinology and Metabolic Diseases of the National Institute of Diabetes and Digestive and Kidney Disease (current grant numbers U01 DK094176 and U01 DK094157), and through support from the National Eye Institute, the National Institute of Neurologic Disorders and Stroke, the General Clinical Research Centers Program (1993–2007) and Clinical Translational Science Center Program (2006–present), Bethesda, MD, USA. Trial Registration: clinicaltrials.gov NCT00360815 and NCT00360893.
Additional support for this DCCT/EDIC collaborative study was provided by grants from the National Institute of Diabetes and Digestive and Kidney Diseases contract N01-DK-6-2204, National Institute of Diabetes and Digestive and Kidney Diseases Grants R01-DK-077510, R01-DK077489 and P60-DK20595, and support from Genome Canada through the Ontario Genomics Institute. ADP holds a Canada Research Chair in the Genetics of Complex Diseases. KME is a recipient of the Gibney Family Scholar Award of the Heart and Stroke Foundation. The content is solely the responsibility of the authors and does not necessarily reflect the official views of the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK).
The skin biopsy ancillary substudy in DCCT/EDIC was funded by the Juvenile Diabetes Research Foundation International (grant no. 17-2010-318 to VMM) and the NIDDK (DK-79432 to DRS).
The LifeLines Cohort Study is supported by the: Netherlands Organization for Scientific Research (NWO) (grant 175.010.2007.006); Economic Structure Enhancing Fund (FES) of the Dutch government; Ministry of Economic Affairs; Ministry of Education, Culture and Science; Ministry for Health, Welfare and Sports; Northern Netherlands Collaboration of Provinces (SNN); Province of Groningen; University Medical Center Groningen; University of Groningen; Dutch Kidney Foundation; and Dutch Diabetes Research Foundation.
Participation in this work was in part also supported by the National Consortium for Healthy Ageing and the BioSHaRE-EU consortium (KP7, project reference 261433). LifeLines (BRIF4568) is engaged in a Bioresource research impact factor (BRIF) policy pilot study, details of which can be found at www.bioshare.eu/content/bioresource-impact-factor.
WESDR is supported by the National Institutes of Health, Bethesda, MD, USA (no. EY016379, to RK and BEKK) and by an unrestricted grant from Research to Prevent Blindness, New York, NY, USA. The content is solely the responsibility of the authors and does not necessarily reflect the official views of the National Eye Institute or the National Institutes of Health.
LonGenity is supported by the: National Institutes of Health (NIH) (P01 AG027734 Barzilai); Glenn Foundation for the Biology of Aging; Nathan Shock Center (P30 AG038072); Einstein Institute for Clinical and Translational Research (CTSA Grant 8UL1 TR000086 from the National Center for Advancing Translational Sciences); Diabetes Research Center (NIH-5P60 DK20541); and NIH (RO1 1R01AG042188 Atzmon). Results for association of rs1495741 with lipids were made available by Teslovich et al [36] and were downloaded from: http://www.sph.umich.edu/csg/abecasis/public/lipids2010/. Data on glycaemic traits have been contributed by MAGIC investigators [37, 38] and have been downloaded from www.magicinvestigators.org.
Duality of interest
JM is a former employee of VeraLight, Inc., the manufacturer of SCOUT, which was used in the present study. TJO has in the past received grant support from VeraLight, Inc and serves as a consultant to Lilly Inc. RG is founder and stockholder of the spin-off DiagnOptics B.V. (Groningen, the Netherlands), the producer of the AGE Reader, which has been used in the present study.
Contribution statement
All authors contributed to either the conception and design of the study, the acquisition of data or their analysis and the interpretation of data, as well as drafting the article or revising it critically, and have provided final approval of the version to be published.
ADP is the guarantor of this work related to the DCCT/EDIC, BHRW is the guarantor of the work related to LifeLines, RK and BEKK are the guarantors of this work related to the WESDR, and NB is the guarantor of this work related to LonGenity. The guarantors listed above had full access to all the data in the study and take full responsibility for the integrity of the data and the accuracy of the data analysis, and controlled the decision to publish.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Additional information
K. M. Eny and H. L. Lutgers contributed equally to this study.
See the electronic supplementary material (ESM) study group list for a full list of study group members.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Study writing groups
(PDF 27 kb)
ESM Table 1
(PDF 179 kb)
ESM Table 2
(PDF 167 kb)
ESM Table 3
(PDF 174 kb)
ESM Table 4
(PDF 102 kb)
ESM Table 5
(PDF 87 kb)
ESM Table 6
(PDF 219 kb)
ESM Table 7
(PDF 210 kb)
ESM Table 8
(PDF 100 kb)
ESM Table 9
(PDF 155 kb)
ESM Table 10
(PDF 96.7 kb)
ESM Fig. 1
(PDF 384 kb)
ESM Fig. 2
(PDF 26 kb)
ESM Fig. 3
(PDF 258 kb)
ESM Fig. 4
(PDF 785 kb)
ESM Fig. 5
(PDF 175 kb)
ESM Fig. 6
(PDF 120 kb)
ESM Fig. 7
(PDF 861 kb)
ESM Fig. 8
(PDF 284 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Eny, K.M., Lutgers, H.L., Maynard, J. et al. GWAS identifies an NAT2 acetylator status tag single nucleotide polymorphism to be a major locus for skin fluorescence. Diabetologia 57, 1623–1634 (2014). https://doi.org/10.1007/s00125-014-3286-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-014-3286-9