Abstract
Aims/hypothesis
Inappropriate insulin secretion and biosynthesis are hallmarks of beta cell dysfunction and contribute to the progression from a prediabetic state to overt diabetes mellitus. During the prediabetic state, beta cells are exposed to elevated levels of proinflammatory cytokines. In the present study the effect of these cytokines and mitogen-activated protein kinase kinase kinase 1 (MEKK1), which is known to be activated by these cytokines, on human insulin gene (INS) transcription was investigated.
Methods
Biochemical methods and reporter gene assays were used in a beta cell line and in primary pancreatic islets from transgenic mice.
Results
IL-1β and MEKK1 specifically inhibited basal and membrane depolarisation and cAMP-induced INS transcription in the beta cell line. Also, in primary islets of reporter gene mice, IL-1β reduced glucose-stimulated INS transcription. A 5′- and 3′-deletion and internal mutation analysis revealed the rat insulin promoter element 3b (RIPE3b) to be a decisive MEKK1-responsive element of the INS. RIPE3b conferred strong transcriptional activity to a heterologous promoter, and this activity was markedly inhibited by MEKK1 and IL-1β. RIPE3b is also known to recruit the transcription factor MafA. We found here that MafA transcription activity is markedly inhibited by MEKK1 and IL-1β.
Conclusions/interpretation
These data suggest that IL-1β through MEKK1 inhibits INS transcription and does so, at least in part, by decreasing MafA transcriptional activity at the RIPE3b control element. Since inappropriate insulin biosynthesis contributes to beta cell dysfunction, inhibition of MEKK1 might decelerate or prevent progression from a prediabetic state to diabetes mellitus.
Similar content being viewed by others
Introduction
The decompensation of pancreatic islet beta cells with loss of beta cell function and finally beta cell mass marks the progression from a prediabetic state, characterised by insulin resistance with impaired glucose tolerance, to clinically overt type 2 diabetes mellitus. Besides insulin secretion, insulin gene transcription is a beta cell-specific function, and inappropriate insulin biosynthesis might contribute to the pathogenesis of diabetes mellitus [1–4].
Among several transcription factors binding to the promoter of the insulin gene, the pancreatic and duodenal homeobox factor 1 (PDX1), the basic helix-loop-helix protein BETA2 (also known as neurogenic differentiation factor 1 or NEUROD) and the basic region/leucine zipper proteins MafA and cyclic AMP response element (CRE) binding protein (CREB) have been shown to be important for the maintenance of insulin biosynthesis as well as beta cell mass [5–12]. Human insulin gene (INS) transcription is activated by PDX1 binding to several sites within this promoter [10, 13]. Mice homozygous for a targeted disruption of Pdx1 fail to develop a pancreas, and inactivation of Pdx1 specifically in beta cells of mice decreased beta cell mass and insulin expression [5, 14]. The overexpression of Beta2 in the hamster insulinoma cell line In1024 enhanced human insulin gene transcription [10], whereas mice deficient in BETA2 showed a reduction of insulin-producing beta cells and failed to develop mature islets [15]. MafA was identified as a glucose-regulated transcription factor binding to the rat insulin promoter element 3b (RIPE3b) within the insulin gene promoter [16, 17], containing the C1 and A2 elements [18]. This transcription factor appears to be responsible for the beta-cell-specific expression of insulin [19]. Furthermore, in MafA-deficient mice, severely impaired glucose-stimulated insulin secretion with a reduction of insulin transcripts and an abnormal islet architecture was observed [12]. In contrast to PDX1, BETA2 and MafA, CREB is an ubiquitously expressed transcription factor. Its recognition site, the CRE, is present in many genes [20]. However, the loss of CREB function in beta cells results in apoptosis [7]. In addition, CREB participates in the maintenance of beta cell function by binding to its binding sites within the rat Ins1 and human INS genes, and by mediating insulin gene transcription induced by cAMP, membrane depolarisation and glucose [8, 9, 21, 22]. Incubation of the beta cell line MIN6 with a combination of the proinflammatory cytokines TNFα, IFNγ and IL-1β was shown to inhibit the transcriptional activity of CREB in a time- and dose-dependent manner [23].
Proinflammatory cytokines have been implicated in the pathogenesis of diabetes mellitus, both type 1 and type 2 [24]. In type 1 diabetes, due to an (auto)immune reaction, mononuclear cells infiltrate the islets of Langerhans and secrete TNFα, IFNγ and IL-1β, resulting in insulitis with beta cell toxicity and, ultimately, beta cell death [24]. In the insulin-resistant state and in obesity, both major risk factors for the development of type 2 diabetes, elevated levels of TNFα and IL-6 secreted from adipocytes are observed [2, 24]. In addition, the proinflammatory cytokine IL-1β was shown to be produced and secreted from beta cells during hyperglycaemic periods [24, 25].
Based on these findings, the aim of the present study was to investigate the effect of proinflammatory cytokines on human insulin gene transcription as a hallmark of beta cell function in a beta cell line and in primary mature islets of transgenic mice.
Materials and methods
Plasmid construction
The plasmids −336 hInsLuc, 4xhInsCRE2, Rous sarcoma virus (RSV)Luc [9], −711 c-fosLuc [26] and the expression vectors for MafA and MafAmt [27] have been described previously. The expression vector for catalytically active mitogen-activated protein kinase (MAPK) kinase kinase 1 (MEKK1) was purchased from Invitrogen (Karlsruhe, Germany). The plasmids encoding the 5′- and 3′-truncated human INS promoters were generated by PCR using oligonucleotides 1–6 as 5′ primers, and 5′-CGCGTCGACGAGCTGGGGCCTGGGGTCCA-3′ as the 3′ primer for the 5′ deletion constructs. The PCR fragments were cloned into the BamHI and SalI sites of pXP2 [28]. For the 3′ deletion constructs, 5′-GCGGGATCCGCCTCCAGCTCTCCTGGTCTA-3′ was used as the 5′ primer, and oligonucleotides 7–13 were used as 3′ primers. The PCR fragment generated by using the 3′ primer 7 was cloned into the BamHI and SalI sites of pXP2, and the PCR fragments for the other 3′ deleted INS promoter constructs were cloned into the BamHI/SalI sites of pT81 [28]. For the constructs 4xhInsC1 and 4xhInsA1, oligonucleotides 14 and 15, respectively, with 5′-GATC overhangs, were tetramerised and subcloned in the forward orientation into the BamHI site of pT81Luc [28]. For the internal mutation of the CRE2 element within the INS promoter, two PCR fragments were generated using oligonucleotides 16 and 17 as 5′ primers, and oligonucleotides 18 and 19 as 3′ primers, thereby changing the CRE2 element into a KpnI site. For the internal mutation of the RIPE3b element, two PCR fragments were amplified using oligonucleotides 20 and 21 as 5′ primers, and oligonucleotides 22 and 23 as 3′ primers, thereby mutating the core RIPE3b element into a SmaI site. For the internal mutation of the A1 element, two PCR fragments were generated using oligonucleotides 24 and 25 as 5′ primers, and oligonucleotides 26 and 27 as 3′ primers, changing the core A1 element into a HindIII site. The respective PCR fragments were cloned into the BamHI/SalI sites of pXP2 [28]. The sequences of the oligonucleotides used are given in Electronic supplementary material (ESM) Table 1 (mutated bases underlined). All constructs were verified by sequencing using the enzymatic method.
Cell culture and DNA transfection
HIT-T15 cells [29] were grown in RPMI 1640 medium supplemented with 10% FCS, 5% horse serum, penicillin (100 U/ml) and streptomycin (100 μg/ml). JEG-3 choriocarcinoma cells [30] were grown in DMEM supplemented with 10% FCS, penicillin (100 U/ml) and streptomycin (100 μg/ml). HIT cells were transfected by the diethylaminoethyl (DEAE)-dextran method [31] with 2 μg of the luciferase reporter gene and the expression vector as indicated. JEG-3 cells were transfected by calcium phosphate precipitation with 2 μg of the luciferase reporter gene and 2 μg of the expression plasmids. Co-transfections were carried out with a constant amount of DNA, which was maintained by addition of Bluescript (Stratagene, La Jolla, CA, USA). To check for transfection efficiency, 1 μg cytomegalovirus-green fluorescent protein (plasmid CMV-GFPtpz) per 6 cm dish was cotransfected. Cell extracts were prepared 48 h after transfection. Cells were stimulated by membrane depolarisation with KCl (40 mmol/l) and the adenylate cyclase activator forskolin (10 μmol/l) for 6 hours prior to collection. Cytokines (TNFα 10 ng/ml; IFNγ 10 ng/ml; IL-1β 2 ng/ml) were added 18 h prior to collection. The activities of luciferase and green fluorescent protein were determined as described previously [31, 32].
Generation and analysis of transgenic mice
The generation and analysis of transgenic mice carrying the luciferase reporter gene under the control of the INS promoter from −336 to +112 has been described before [8]. All animal studies were conducted according to the National Institutes of Health’s guidelines for care and use of experimental animals and were approved by the Committee on Animal Care and Use of the local institution and state.
Isolation and culture of islets
Pancreatic islets were isolated as described previously [8, 33]. Isolated islets were preincubated in a humidified atmosphere of 95% air/5% CO2 for 1 h in RPMI 1640 medium containing 5 mmol/l glucose supplemented with 10% fetal calf serum, 100 U/ml penicillin and 100 μg/ml streptomycin. Glucose (15 mmol/l) and IL-1β (10 ng/ml) were added 24 h and 23 h prior to collection. Islets were collected, washed once with PBS and resuspended in potassium phosphate buffer, pH 7.8, followed by three freeze–thaw cycles. Luciferase activity [30] and protein (Protein Assay; Bio-Rad Laboratories, Munich, Germany) content were determined in the supernatant.
Western blot analysis
JEG-3 cells were transfected with 3 μg of the expression vectors for MafA or its mutant per 6 cm dish by the metafectene method according to the manufacturer’s protocol. 48 h after transfection cells were collected into 150 μl hot 2× Laemmli gel loading buffer (Tris–HCl 62.5 mmol/l, pH 6.8, SDS 2% (wt/vol.), glycerol 10% (vol./vol.), β-mercaptoethanol 5% (vol./vol.), bromophenol blue 0.5% (wt/vol.) and scraped from the dish. The cell suspension was passed five times through a 26 gauge needle, centrifuged, boiled, subjected to SDS-PAGE (8% gel) and transferred to a nitrocellulose membrane. The membrane was incubated in 7.5% fat-free dried milk in TBST (Tris–HCl 25 mmol/l, pH 7.4; NaCl 137 mmol/l; KCl 5 mmol/l; CaCl2 0.7 mmol/l; MgCl2 0.1 mmol/l; Tween 20, 0.1%) for 2 h at room temperature and then overnight at 4°C with fresh TBST supplemented with an antibody against MafA (1:2,000; Bethyl Laboratories, Montgomery, TX, USA). Before and after the incubation with the secondary antibody the membrane was washed four times for 10 min, each time with TBST. The antibody–antigen complex was detected by electrochemiluminescence reagents (GE Healthcare, Little Chalfont, Bucks, UK).
Materials
Cytokines were purchased from Strathmann Biotec (Hamburg, Germany), luciferin was obtained from Promega (Mannheim, Germany), forskolin from Sigma (Taufkirchen, Germany). Forskolin was dissolved in DMSO. Controls received the solvent only.
Statistical analysis
All results are expressed as means±SEM. Statistical significance was calculated with ANOVA, followed by Student’s t test. A value of p < 0.05 was considered significant.
Results
Effect of IL-1β, TNFα and IFNγ on human insulin gene transcription
Treatment with the adenylate cyclase activator forskolin plus high potassium-induced membrane depolarisation enhanced insulin gene transcription 2.5-fold (Fig. 1). Addition of IFNγ or TNFα or a combination thereof had no effect on basal or stimulated insulin gene transcription (Fig. 1). In contrast, IL-1β decreased basal and stimulated insulin gene transcription by 80% and 74%, respectively. The treatment of HIT cells with a combination containing IL-1β resulted in a similar decrease of insulin gene transcription as with IL-1β alone. To investigate the effect of IL-1β in primary beta cells and on glucose-stimulated human insulin gene transcription, the islets of adult mice carrying a luciferase reporter transgene under the control of the INS promoter were used (Fig. 2a). Human insulin-driven reporter gene transcription within the islets of these transgenic mice has been shown to be stimulated by glucose in physiological concentrations [8]. The treatment of the isolated islets with IL-1β for 23 h inhibited glucose-induced human insulin gene transcription by 47% (Fig. 2b). Thus, of the proinflammatory cytokines tested, only IL-1β decreased basal and stimulated human insulin gene transcription in HIT cells. In addition, IL-1β decreased glucose-stimulated human insulin gene transcription in primary islets. IL-1β inhibited human INS transcription, whereas it somewhat enhanced c-fos (also known as FOS) gene transcription and had no effect on RSV promoter activity in HIT cells (Fig. 3), indicating that the effect of IL-1β on human insulin gene transcription was specific.
Effect of the proinflammatory cytokines IFNγ, IL-1β and TNFα on basal and stimulated human INS transcription in a beta cell line. The plasmid −336 hInsLuc (2 μg) was transiently transfected into HIT cells. Cells were treated as indicated with IFNγ (10 ng/ml), IL-1β (2 ng/ml) and TNFα (10 ng/ml) for 18 h and with the adenylate cyclase activator forskolin (10 μmol/l) plus KCl (40 mmol/l; white bars) for 6 h prior to collection. Luciferase activity is expressed relative to the mean value of the control in each experiment (no treatment). Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the respective control in the absence of cytokine treatment
Effect of IL-1β on glucose-induced INS transcription in primary islets of transgenic mice. a Scheme of the microinjected DNA fragment containing the human INS gene promoter from −336 to +114 bp fused to the firefly luciferase gene. The known sequence elements are depicted according to Ohneda et al. [6] and Inagaki et al. [50]. b Islets of transgenic mice were isolated and treated with glucose (15 mmol/l) for 24 h with and without IL-1β (10 ng/ml) for 23 h prior to collection. Luciferase activity is expressed relative to the value in the control treated with glucose only. Values are means±SEM of five independent experiments. *p < 0.05 vs control
Specific inhibition of INS transcription by IL-1β. The plasmids −336 hInsLuc (2 μg), −711 c-fosLuc (2 μg) and RSVLuc (0.5 μg) were transiently transfected into HIT cells. Cells were treated with IL-1β (2 ng/ml) (black bars) for 18 h prior to collection. Luciferase activity is expressed relative to the mean value measured in the respective controls (no IL-1β, white bars) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the respective control
Cytokines like IL-1β have been reported to increase the concentration of reactive oxygen species within the beta cell [34]. In vitro, reactive oxygen species were shown to inhibit the activity of the calcium/calmodulin-dependent serine/threonine phosphatase calcineurin [35], while recent data demonstrate the pivotal role of calcineurin for beta cell proliferation and human insulin gene transcription [8, 9, 36]. However, IL-1β did not change calcineurin phosphatase activity in HIT cells (data not shown).
Inhibition of human insulin gene transcription by MEKK1
IL-1β has been shown to enhance the activity of the mitogen-activated protein kinase kinase kinase [37–39]. The effect of IL-1β on the transcriptional activity of the INS promoter, the c-fos gene promoter and the RSV promoter was thus compared with the effect of MEKK1 on the activity of these promoters. Like IL-1β, overexpression of the catalytic domain of MEKK1 decreased INS transcription, increased the transcriptional activity of the c-fos promoter and had only a slight effect on the promoter of the RSV (Figs. 3 and 4a). As shown in Fig. 4b, increasing amounts of the expression vector for MEKK1 resulted in a growing inhibition of basal insulin gene transcription and insulin gene transcription induced by forskolin plus membrane depolarisation. These data suggest that MEKK1 mimics the effect of IL-1β.
Effect of MEKK1 on human insulin gene transcription. a The plasmids −336 hInsLuc (2 μg), −711 c-fosLuc (2 μg) and RSVLuc (0.5 μg) were transiently transfected into HIT cells. Bluescript (2 μg) (white bars) or an expression vector for the catalytic domain of MEKK1 (2 μg) (black bars) was cotransfected as indicated. Luciferase activity is expressed relative to the mean value measured in the respective control (no MEKK1) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the respective control. b The plasmid −336 hInsLuc (2 μg) was transiently cotransfected with 0.02, 0.06, 0.2, 0.6 and 2 μg of an expression vector for MEKK1. Cells were stimulated with the adenylate cyclase activator forskolin (10 μmol/l) plus KCl (40 mmol/l) (open circles) for 6 h prior to collection or left untreated (black circles). Luciferase activity is expressed relative to the mean value measured in the control (no forskolin plus KCl, no MEKK1) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the respective control in the absence of MEKK1
Effect of inhibitors of mitogen-activated kinases downstream of MEKK1
In various tissues, MEKK1 has been shown to phosphorylate and activate the dual-specificity kinases mitogen-activated extracellular-regulated kinase 1 (MEK1), mitogen-activated kinase kinase 4/7 (MKK4/7) and MKK3/6 [40], leading to the activation of the MAPKs extracellular signal-regulated kinase (ERK)1/2, c-Jun-N-terminal kinase (JNK) and stress-activated protein kinase/p38 (SAPK/p38), respectively [40]. To investigate whether one of these downstream kinases mediates the inhibitory effect of MEKK1 on insulin gene transcription, SP600125, SB203580 and U0126 were used as pharmacological inhibitors of JNK, SAPK/p38 and ERK activity, respectively [40]. The transcriptional activity of the human insulin gene itself was slightly decreased by U0126 and the combination of these inhibitors; however, no decrease of the transcriptional activity of the cytomegalovirus promoter directing GFP expression was observed (data not shown). None of these inhibitors or their combination relieved the inhibitory effect of MEKK1 (ESM Fig. 1a), although they effectively decreased phosphorylation of JNK, SAPK/p38 or ERK, respectively (ESM Fig. 1b–g). This suggests that MEKK1 directly impacts on activation of an insulin gene transcription factor(s).
Identification of MEKK1/IL-1β response elements within the INS promoter
The INS promoter contains several enhancer-like elements among them the CRE bound by CREB, the A1, A3 and A5 elements bound by PDX1, the E1 element bound by BETA2/NEUROD, and the C1 element bound by MafA [6, 9, 16, 18, 19]. To identify a region within the INS promoter that confers MEKK1 responsiveness, a 5′- and 3′-deletion analysis was undertaken. As shown in Fig. 5a, MEKK1 decreased human INS transcription (−336 hInsLuc) by 70%. 5′-deletions up to −193 bp neither decreased the transcriptional activity of the promoter fragments nor attenuated the inhibitory effect of MEKK1 (Fig. 5a). Truncation to −140 bp diminished the transcriptional activity of the INS promoter fragment by 85%. However, MEKK1 still inhibited insulin gene transcription, although the inhibitory effect of MEKK1 was reduced (inhibition by 50%) (Fig. 5a). Deletion to −93 bp resulted in a further reduction of INS promoter activity and a complete loss of MEKK1 responsiveness (Fig. 5a). Additionally, 3′-deleted fragments of the INS promoter were fused to the minimal thymidine kinase promoter (−81 to +52) of the herpes simplex virus (Fig. 5b). Truncation up to −57 bp did not impair basal activity or the inhibition by MEKK1 of these INS fragments (Fig. 5b). Deletion from −57 bp to −94 resulted in a dramatic decrease of basal transcriptional activity. However, MEKK1 still inhibited gene transcription (Fig. 5b), although the response to MEKK1 was somewhat diminished. Further 3′-deleted INS promoter fragments exhibited a further reduced transcriptional activity, albeit still higher than that conferred by the minimal herpes simplex virus thymidine kinase promoter alone (Fig. 5b). In addition, the transcriptional activities of these 3′-deletion INS fragments were no longer inhibited by MEKK1 (Fig. 5b). These data suggest that the C1 element and, to some degree, also the CRE2 and A1 elements confer the inhibitory effect of MEKK1 upon the insulin gene promoter.
Effect of MEKK1 on 5′- or 3′-deleted INS promoter fragments. a 5′-deletion analysis. The plasmids (2 μg) depicted on the left were transiently cotransfected with Bluescript (2 μg; minus signs, x-axis) or the expression vector for the catalytically active domain of MEKK1 (2 μg; plus signs, x-axis). Luciferase activity is expressed relative to mean value measured in the −336 control (no MEKK1) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the control in the absence of MEKK1. b 3′-deletion analysis. The plasmids (2 μg) depicted on the left were cotransfected with Bluescript (2 μg; minus signs, x-axis) or the expression vector for the catalytically active domain of MEKK1 (2 μg; plus signs, x-axis). Luciferase activity is expressed relative to the mean value measured in the −336 control (no MEKK1) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the control in the absence of MEKK1
Mutation of the CRE2, the RIPE3b and the A1 element lowered the transcriptional activity of the INS promoter to 40 ± 12%, 18 ± 1% and 37 ± 4%, respectively, when compared with wild-type (100 ± 2%) (ESM Fig. 2). Whereas MEKK1 decreased the activity of the wild-type, the CRE2-mutated and the A1-mutated INS promoters to 18, 31 and 11% of their non-inhibited values, respectively, the transcriptional activity of the RIPE3b-mutated promoter was decreased to a lesser extent (to 46% of original activity) (ESM Fig. 2). The effect of MEKK1 on the isolated promoter elements was investigated using luciferase reporter genes under the control of the minimal thymidine kinase promoter fused to four copies of CRE2, RIPE3b and A1, respectively. Compared with the minimal thymidine kinase promoter (100 ± 1.3%; n = 6), the basal transcriptional activities after fusion to CRE2, C1 and A1 were 194 ± 16%, 18,190 ± 4349% and 89 ± 3.2%, respectively. Membrane depolarisation plus forskolin enhanced CRE2-directed transcription, confirming previous data [9] (Fig. 6a). Stimulated CRE2-directed transcription was completely abolished by MEKK1 (Fig. 6a). In addition, MEKK1 markedly diminished the basal transcriptional activity conferred by C1, but enhanced basal CRE2- and A1-directed transcription (Fig. 6a–c). Like MEKK1, IL-1β decreased C1 transcriptional activity (Fig. 6d).
Effect of MEKK1 on isolated INS promoter fragments. a–c Isolated tetramerised promoter elements CRE2 (a), C1 (b) and A1 (c). The plasmids 4xhInsCRE2 (2 μg), 4xhInsC1 (2 μg) and 4xhInsA1 (2 μg) were transiently cotransfected with Bluescript (2 μg) (white and hatched bars) or with MEKK1 (2 μg) (black and crosshatched bars). Cells (hatched and crosshatched bars) were treated with the adenylate cyclase activator forskolin (10 μmol/l) plus KCl (40 mmol/l) for 6 h prior to collection. Luciferase activity is expressed relative to the mean value of the respective controls (no MEKK1, no forskolin plus KCl) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs control in the absence of MEKK1; **p < 0.05 vs the treatment with forskolin and KCl. d Effect of IL-1β on C1-dependent transcription. The plasmid 4xhInsC1 (2 μg) was transiently transfected into HIT cells. Cells were treated with IL-1β (2 ng/ml; black bar) for 18 h prior to collection. Luciferase activity is expressed relative to the mean value measured in the control (no IL-1β; open bar) in each experiment. Values are means±SEM of three independent experiments, each performed in triplicate. *p < 0.05 vs the control in the absence of IL-1β
Effect of MEKK1 and IL-1β on the RIPE3b-binding transcriptional activator MafA
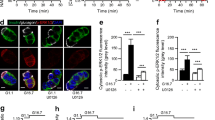
The insulin C1 element within RIPE3b [18] has been shown to recruit the transcription factor MafA to the insulin promoter, which is produced in pancreatic islet beta cells and all beta cell lines, including HIT cells [27]. To investigate the effect of MEKK1 and IL-1β on MafA transcriptional activity, the human insulin gene promoter construct was transfected into the human choriocarcinoma cell line JEG, together with expression vectors for MEKK1, MafA or MafA mutant carrying a dysfunctional mutation within the DNA binding domain [11]. MafA exhibited marked transcriptional activity whereas MafA-R265A only slightly enhanced transcription (Fig. 7a). Western blot analysis demonstrated similar levels of production of wild-type and mutant MafA (inset Fig. 7a). Co-transfection of the expression vector for MEKK1 nearly abolished MafA-dependent transcription (Fig. 7a). Like MEKK1, IL-1β also decreased MafA transcriptional activity (Fig. 7b). Taken together, these findings are consistent with the notion that IL-1β, through activation of MEKK1, inhibits MafA transcriptional activity and thus human insulin gene transcription.
Effect of MEKK1 and IL-1β on MafA transcriptional activity. a Effect of MEKK1. The plasmid −336 hInsLuc (2 μg) was transiently cotransfected with Bluescript (2 μg) (white bars), an expression for MafA (2 μg) (black bars) or an expression for MafA R265A (2 μg) (crosshatched bars) into JEG cells with and without MEKK1. Luciferase activity is expressed relative to the mean value in the respective controls (no MafA, no MafA R265A) in each experiment. Values are means±SEM of three independent experiments, each performed in triplicate. *p < 0.05 vs the control in the absence of MEKK1. The inset shows a typical Western blot using an antibody against MafA to check for equal production of MafA and its mutant. b Effect of IL-1β. The plasmids −336 hInsLuc and MafA (2 μg each) were transiently cotransfected into JEG cells. When indicated, MEKK1 (2 μg) was cotransfected or the cells were treated with IL-1β (2 ng/ml) for 18 h prior to collection. Luciferase activity is expressed relative to the mean value in the control (no IL-1β, no MEKK1) in each experiment. Values are means±SEM of three independent experiments, each performed in duplicate. *p < 0.05 vs the control in the absence of MEKK1 or IL-1β
Discussion
Besides insulin secretion, the appropriate transcription of the insulin gene is an essential function of beta cells. Progressive beta cell failure with inadequate insulin secretion and biosynthesis, as well as loss of beta cells is characteristic for the pathogenesis of type 1 and type 2 diabetes mellitus [1–4]. Proinflammatory cytokines have been implicated in the development of both types [24, 41]. In obesity, a major risk factor for peripheral insulin resistance and diabetes mellitus type 2, elevated levels of proinflammatory cytokines are found, with TNFα and IL-6 secreted by the adipocytes [2]. IL-1β might be produced and secreted in human beta cells during hyperglycaemic periods [25], although an elevation of IL-1β levels has not been demonstrated in the islets of type 2 diabetic patients or in the islets of hyperglycaemic Psammomys obesus, an animal model of type 2 diabetes [41, 42]. However, for the pathogenesis of type 1 diabetes, the involvement of IL-1β secreted by infiltrating mononuclear cells seems undisputed [41 and references therein]. The present study shows that IL-1β interferes with beta cell function by inhibiting insulin gene transcription. This inhibition was specific since neither the transcriptional activity of the c-fos gene promoter nor that of the RSV promoter was decreased by IL-1β. The combination of IL-1β with either TNFα or IFNγ did not add to IL-1β’s inhibitory effect. In addition, this cytokine also inhibited INS transcription after stimulation by high potassium-induced membrane depolarisation plus forskolin in a beta cell line and by glucose in primary mature islets of transgenic luciferase reporter mice. Since the regulation of insulin gene transcription is highly conserved throughout the species [6], these results might be easily transferable. Binding of IL-1β to its type 1 receptor (IL-1R1) leads to the recruitment of the IL-1 receptor accessory protein, which is followed by the recruitment of the IL-1R-associated kinase (IRAK) via the adaptor protein MyD88. IRAK then interacts with the TNF receptor-associated factor (TRAF)6 [24] and references therein], and oligomerisation of TRAF6 and TRAF2 results in the activation of MEKK1 [24, 37, 38]. Indeed, in mouse embryonic stem cells deficient in MEKK1, the requirement of this kinase for IL-1 action has been demonstrated [43]. MEKK1 belongs to the group of MAPKs. As a triple kinase, MEKK1 activates through phosphorylation of the dual specificity kinases MKK4, MKK7 and MEK1, which in turn phosphorylate and activate the MAPKs JNK and ERK, respectively [40]. Like IL-1β, MEKK1 inhibited basal and membrane depolarisation plus forskolin-induced insulin gene transcription, enhanced the transcriptional activity of the c-fos promoter with only a slight effect on the RSV promoter activity, decreased insulin C1-directed transcription and blocked MafA transcriptional activity. MEKK1 thus mimics the effect of IL-1β. These findings are consistent with the notion that IL-1β exerts its inhibitory effect on human insulin gene transcription through the activation of MEKK1. In the present study the MEKK1-induced inhibition of the human insulin gene was not relieved by inhibitors of MAPKs downstream of MEKK1. In U2OS cells the activated form of MEKK1 was detected within the nucleus and the transcriptional coactivator p300 was directly phosphorylated by MEKK1 [44]. In response to IL-1β signalling, MEKK1 was shown to phosphorylate the nuclear TAK binding protein 2 (also known as TAB2), resulting in its conformational change and binding to the nuclear receptor corepressor, thereby leading to a disinhibition of androgen receptor activity [39]. Thus, MEKK1 might exert its effects on insulin gene transcription by acting directly on a transcription factor or accessory factor, without involvement of its downstream kinases.
A 5′- and 3′-deletion and internal mutation analysis, together with experiments using isolated promoter elements, revealed the C1 element to be a cis-acting element conferring the inhibitory effect of MEKK1 and IL-1β in the insulin gene. With the 5′- or 3′-deletion of RIPE3b, MEKK1 responsiveness of the human insulin gene was reduced profoundly. Furthermore, RIPE3b/C1 was sufficient to confer MEKK1 and IL-1β responsiveness. The C1 element has been shown to mediate glucose-responsive and beta-cell-specific insulin gene transcription, demonstrating its importance for beta cell function [17, 45]. In addition to C1, other control elements may also be involved, including CRE2 and A1. Hence, 5′-deletion of CRE2 and 3′-deletion of A1 attenuated the inhibitory effect of MEKK1 on human insulin gene transcription, although neither CRE2 nor A1 was sufficient to confer MEKK1 responsiveness to a heterologous promoter, suggesting that these elements are only MEKK1-responsive within the context of the INS promoter. Different transcription factors bind to these elements: CRE2 is bound by CREB [9], the C1-binding protein was identified as MafA [16, 17] and PDX1 binds to the A1 element [10, 13].
The finding of the present study that transcription directed by four copies of the isolated MafA-binding site C1 is inhibited by MEKK1 and by IL-1β suggests that MafA confers the cytokine’s inhibitory effect upon the insulin gene. Indeed, in the non-beta cell line JEG, the stimulating effect of MafA on insulin gene transcription was nearly completely abolished by MEKK1 and IL-1β. Thus, MafA transcriptional activity is regulated by MEKK1 and by IL-1β. MafA belongs to the class of basic region/leucine zipper transcription factors and has been implicated in the development and differentiation of the lens [46]. Moreover, identification of MafA as the transcription factor binding to the C1 element within the rat Ins2 and the INS promoter [16, 17] fuelled studies on the role of MafA in beta cell function. During development, MafA was shown to be exclusively produced in insulin-positive cells and essential for insulin gene transcription [19]. In addition, MafA was shown to be a master regulator of genes implicated in maintaining beta cell function [47]. In mice lacking MafA, reduced insulin 1 and 2 gene transcription, glucose intolerance and development of diabetes mellitus was observed [12]. At least at the beta cell level the transcriptional activity of MafA seems to be regulated by either changes in MafA synthesis or by an enhanced degradation of this protein [11, 16, 26, 48, 49]. However, in the present study overproduction of MEKK1 did not decrease the amount of MafA produced in JEG cells (data not shown), arguing against a MEKK1-induced degradation of MafA. In addition to protein synthesis or degradation, the transcriptional activity of MafA is regulated by phosphorylation [46]. The present study suggests that IL-1β through activation of MEKK1 phosphorylates MafA or an associated protein, thereby resulting in a decrease of human insulin gene transcription. In addition, the transcription of other MafA-dependent genes maintaining beta cell function might be disturbed by IL-1β and MEKK1. Given (1) that beta cells are exposed to elevated levels of IL-1β; and (2) the importance of MafA for beta cell function, IL-1β, through activation of MEKK1, might contribute to the pathogenesis of diabetes mellitus. Thus, beta-cell-specific inhibition of MEKK1 might provide a useful approach for pharmacological intervention to decelerate or prevent the progression from a prediabetic state to overt diabetes mellitus.
Abbreviations
- CRE:
-
cyclic AMP response element
- CREB:
-
cyclic AMP response element binding protein
- ERK:
-
extracellular signal-regulated kinase
- IL-1R1:
-
IL-1β type 1 receptor
- IRAK:
-
IL-1R-associated kinase
- JNK:
-
c-Jun-N-terminal kinase
- MAPK:
-
mitogen-activated protein kinase
- MEKK1:
-
mitogen-activated protein kinase kinase kinase 1
- MEK1:
-
mitogen-activated extracellular-regulated protein kinase kinase 1
- MKK4/7:
-
mitogen-activated protein kinase kinase 4/7
- PDX1:
-
pancreatic and duodenal homeobox factor 1
- RIPE3b:
-
rat insulin promoter element 3b
- RSV:
-
rous sarcoma virus
- SAPK/p38:
-
stress-activated protein kinase/p38
- TRAF:
-
TNF receptor-associated factor
References
Stumvoll M, Goldstein BJ, van Haeften TW (2005) Type 2 diabetes: principles of pathogenesis and therapy. Lancet 365:1333–1346
Lazar MA (2005) How obesity causes diabetes: not a tall tale. Science 307:373–375
Rhodes CJ (2005) Type 2 diabetes—a matter of beta-cell life and death? Science 307:380–384
Sherry NA, Tsai EB, Herold KC (2005) Natural history of β-cell function in type 1 diabetes. Diabetes 54 (Suppl 2):S32–S39
Ahlgren U, Jonsson J, Jonsson L, Simu K, Edlund H (1998) β-Cell-specific inactivation of the mouse Ipf1/Pdx1 gene results in loss of β-cell phenotype and maturity onset diabetes. Genes Dev 12:1763–1768
Ohneda K, Ee H, German M (2000) Regulation of insulin gene transcription. Semin Cell Dev Biol 11:227–233
Jhala US, Canettieri G, Screaton RA et al (2003) cAMP promotes pancreatic beta-cell survival via CREB-mediated induction of IRS2. Genes Dev 17:1575–1580
Oetjen E, Baun D, Beimesche S et al (2003) Inhibition of human insulin gene transcription by the immunosuppressive drugs cyclosporin A and tacrolimus in primary, mature islets of transgenic mice. Mol Pharmacol 63:1289–1295
Oetjen E, Grapentin D, Blume R et al (2003) Regulation of human insulin gene transcription by the immunosuppressive drugs cyclosporin A and tacrolimus at concentrations that inhibit calcineurin activity and involving the transcription factor CREB. Naunyn Schmiedebergs Arch Pharmacol 367:227–236
Aramata S, Han S, Yasuda K, Kataoka K (2005) Synergistic activation of the insulin gene promoter by the β-cell enriched transcription factors MafA, Beta2, and Pdx1. Biochim Biophys Acta 1730:41–46
Zhao L, Guo M, Matsuoka TA et al (2005) The islet β cell-enriched MafA activator is a key regulator of insulin gene transcription. J Biol Chem 280:11887–11894
Zhang C, Moriguchi T, Kajihara M et al (2005) MafA is a key regulator of glucose-stimulated insulin secretion. Mol Cell Biol 25:4969–4976
Le Lay J, Stein R (2006) Involvement of PDX-1 in activation of human insulin gene transcription. J Endocrinol 188:287–294
Jonsson J, Carlsson L, Edlund T, Edlund H (1994) Insulin-promoter-factor 1 is required for pancreas development in mice. Nature 371:606–609
Naya FJ, Huang H, Qiu Y et al (1997) Diabetes, defective pancreatic morphogenesis, and abnormal differentiation in BETA2/neuroD-deficient mice. Genes Dev 11:2323–2334
Kataoka K, Han S, Shioda S, Hiria M, Nishizawa M, Handa H (2002) MafA is a glucose-regulated and pancreatic β-cell-specific transcriptional activator for the insulin gene. J Biol Chem 277:49903–49910
Olbrot M, Rud J, Moss LG, Sharma A (2002) Identification of β-cell-specific insulin gene transcription factor RIPE3b1 as mammalian MafA. Proc Natl Acad Sci USA 99:6737–6742
Harrington RH, Sharma A (2001) Transcription factors recognizing overlapping C1–A2 binding sites positively regulate insulin gene expression. J Biol Chem 276:104–113
Matsuoka T, Artner I, Henderson E, Means A, Sander M, Stein R (2004) The MafA transcription factor appears to be responsible for tissue-specific expression of insulin. Proc Natl Acad Sci USA 101:2930–2933
Mayr B, Montminy M (2001) Transcriptional regulation by the phosphorylation-dependent factor CREB. Nat Rev Mol Cell Biol 2:599–609
Eggers A, Siemann G, Blume R, Knepel W (1998) Gene-specific transcriptional activity of the insulin cAMP-responsive element is conferred by NF-Y in combination with cAMP response element-binding protein. J Biol Chem 273:18499–18508
Oetjen E, Diedrich T, Eggers A, Eckert B, Knepel W (1994) Distinct properties of the cAMP-responsive element of the rat insulin I gene. J Biol Chem 269:27036–27044
Jambal P, Masterson S, Nesterova A et al (2003) Cytokine-mediated down-regulation of the transcription factor cAMP-response element-binding protein in pancreatic beta-cells. J Biol Chem 278:23055–23065
Donath MY, Storling J, Maedler K, Mandrup-Poulsen T (2003) Inflammatory mediators and islet β-cell failure: a link between type 1 and type 2 diabetes. J Mol Med 81:455–470
Maedler K, Sergeev P, Ris F et al (2002) Glucose-induced β cell production of IL-1β contributes to glucotoxicity in human pancreatic islets. J Clin Invest 110:851–860
Siemann G, Blume R, Grapentin D, Oetjen E, Schwaninger M, Knepel W (1999) Inhibition of cyclic AMP response element-binding protein/cyclic AMP response element-mediated transcription by the immunosuppressive drugs cyclosporin A and FK506 depends on the promoter context. Mol Pharmacol 55:1094–1100
Harmon JS, Stein R, Robertson RP (2005) Oxidative stress-mediated, post-translational loss of MafA protein as a contributing mechanism to loss of insulin gene expression in glucotoxic beta cells. J Biol Chem 280:11107–11113
Nordeen SK (1988) Luciferase reporter gene vectors for analysis of promoters and enhancers. BioTechniques 6:454–458
Santerre RF, Cook RA, Crisel RM et al (1981) Insulin synthesis in a clonal cell line of simian virus 40-transformed hamster pancreatic beta cells. Proc Natl Acad Sci USA 78:4339–4343
Kohler PO, Bridson WE (1971) Isolation of hormone-producing clonal lines of human choriocarcinoma. J Clin Endocrinol Metab 32:683–687
Schwaninger M, Lux G, Blume R, Oetjen E, Hidaka H, Knepel W (1993) Membrane depolarization and calcium influx induce glucagon gene transcription in pancreatic islet cells through the cyclic AMP-responsive element. J Biol Chem 268:5168–5177
Oetjen E, Thoms KM, Laufer Y et al (2005) The immunosuppressive drugs cyclosporin A and tacrolimus inhibit membrane depolarization-induced CREB transcription activity at the coactivator level. Br J Pharmacol 144:982–993
Lacy PE, Kostianovsky M (1967) Method for the isolation of intact islets of Langerhans from the rat pancreas. Diabetes 16:335–339
Rabinovitch A, Suarez-Pinzon WL, Strynadka K, Lakey JR, Rajotte RV (1996) Human pancreatic islet beta-cell destruction by cytokines involves oxygen free radicals and aldehyde production. J Clin Endocrinol Metab 81:3197–3202
Namgaladze D, Hofer HW, Ullrich V (2002) Redox control of calcineurin by targeting the binuclear Fe(2+)–Zn(2+) center at the enzyme active site. J Biol Chem 277:5962–5969
Heit JJ, Apelqvist AA, Gu X et al (2006) Calcineurin/NFAT signalling regulates pancreatic beta-cell growth and function. Nature 443:345–349
Baud V, Liu ZG, Bennett B, Suzuki N, Xia Y, Karin M (1999) Signaling by proinflammatory cytokines: oligomerization of TRAF2 and TRAF6 is sufficient for JNK and IKK activation and target gene induction via an amino-terminal effector domain. Genes Dev 13:1297–1308
Chadee DN, Yuasa T, Kyriakis JM (2002) Direct activation of mitogen-activated protein kinase kinase kinase MEKK1 by the Ste20p homologue GCK and the adapter protein TRAF2. Mol Cell Biol 22:737–749
Zhu P, Baek SH, Bourk EM et al (2006) Macrophage/cancer cell interactions mediate hormone resistance by a nuclear receptor derepression pathway. Cell 124:615–629
Schlesinger TK, Fanger GR, Yujiri T, Johnson GL (1998) The TAO of MEKK. Front Biosci 3:D1181–D1186
Cnop M, Welsh N, Jonas JC, Jörns A, Lenzen S, Eizirik DC (2005) Mechanisms of pancreatic β-cell death in type 1 and type 2 diabetes. Diabetes 54(Suppl 2):S97–S107
Jörns A, Rath KJ, Bock O, Lenzen S (2006) Beta cell death in hyperglycaemic Psammomys obesus is not cytokine-mediated. Diabetologia 49:2704–2712
Xia Y, Makris C, Su B et al (2000) MEK kinase 1 is critically required for c-Jun N-terminal kinase activation by proinflammatory stimuli and growth factor-induced cell migration. Proc Natl Acad Sci USA 97:5243–5248
See RH, Calvo D, Shi Y et al (2001) Stimulation of p300-mediated transcription by the kinase MEKK1. J Biol Chem 276:16310–16317
Stellrecht CM, DeMayo FJ, Finegold MJ, Tsai MJ (1997) Tissue-specific and developmental regulation of the rat insulin II gene enhancer, RIPE3b, in transgenic mice. J Biol Chem 272:3567–3572
Benkhelifa S, Provot S, Nabais E, Eychene A, Calothy G, Felder-Schmittbuhl MP (2001) Phosphorylation of MafA is essential for its transcriptional and biological properties. Mol Cell Biol 21:4441–4452
Wang H, Brun T, Kataoka K, Sharma A, Wollheim CB (2007) MAFA controls genes implicated in insulin biosynthesis and secretion. Diabetologia 50:348–358
Vanderford NL, Andrali SS, Özcan S (2007) Glucose induces MafA expression in pancreatic beta cell lines via the hexosamine biosynthetic pathway. J Biol Chem 282:1577–1584
Hagman DK, Hays LB, Parazzoli SD, Poitout V (2005) Palmitate inhibits insulin gene expression by altering PDX-1 nuclear localization and reducing MafA expression in isolated rat islets of Langerhans. J Biol Chem 280:32413–32418
Inagaki N, Maekawa T, Sudo T, Ishii S, Seino Y, Imura H (1992) c-Jun represses the human insulin promoter activity that depends on multiple cAMP response elements. Proc Natl Acad Sci USA 89:1045–1049
Acknowledgements
This study was supported by grants from the Deutsche Diabetes Gesellschaft (German Diabetes Society; to E. Oetjen), the Deutsche Forschungsgemeinschaft SFB 402 (A3) (German Research Foundation; to W. Knepel) and the National Institutes of Health (grant 42502) (to R. Stein).
Duality of interest
The authors declare that there is no duality of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to electronic supplementary material.
ESM Fig. 1
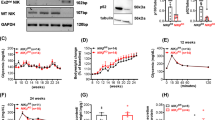
Effect of inhibitors of MEKK1-downstream kinases. a Effect of inhibitors on MEKK1-induced inhibition of human INS transcription. The plasmid -336 hInsLuc (2μg) was transiently cotransfected either with Bluescript (2μg) (white bars) or the expression vector for the catalytically active domain of MEKK1 (2μg) (black bars). Cells were treated as indicated with SP600125 (SP; 25 μmol/l), SB203580 (SB; 20 μmol/l), U0126 (U; 20μmol/l) or a combination of the three for 7 h prior to collection. Luciferase activity is expressed (in %) relative to the mean value of the respective control in each experiment. Values are means±SEM of two independent experiments, each performed in duplicate. b, c Effect of SP600125 on the phosphorylation (phosphor) of JNK1 and 2 in HIT cells. Cells were treated with anisomycin (0.5μg/ml) and/or SP600125 (25 μmol/l) 15 min and 30 min prior to collection respectively, and a western blot was performed. Typical western blots (b) are shown, using antibodies against phosphorylated JNK1 and 2, and against JNK1 and 2, where an equal amount of unphosphorylated protein is seen in each lane. Arrows indicate JNK1 (54 K) and JNK2 (46 K). Densitometric evaluation (c) shows values expressed relative to the optical density of the band representing phosphorylated JNK1 or 2 in the presence of anisomycin. Values are means±SEM of four independent experiments. *p < 0.05 vs anisomycin-treated control. d, e Effect of SB203580 on the phosphorylation of SAPK/p38 in HIT cells. Cells were treated with anisomycin (0.5 μg/ml) and SB203580 (20 μmol/l) 15 min and 30 min prior to harvest respectively, and a western blot was performed. Typical western blots (d) are shown using antibodies against phosphorylated SAPK/p38 and against SAPK/p38, demonstrating an equal amount of unphosphorylated protein in each lane. The arrow indicates SAPK/p38 (38 K). Densitometric evaluation (e) shows values expressed relative to the optical density of the band representing phosphorylated SAPK/p38 in the presence of anisomycin. Values are means±SEM of four independent experiments. *p < 0.05 vs the anisomycin-treated control. f, g Effect of U0126 on the phosphorylation of ERK1 and 2 in HIT cells. Cells were treated with the phorbolester TPA (300 nmol/l) and KCl (40 mmol/l), and with U0126 (20 μmol/l) 15 min and 30 min prior to harvest respectively. Western blot was performed. Tpical western blots (f) are shown using antibodies against phosphorylated ERK1 or 2 and against ERK1 and 2, demonstrating an equal amount of unphosphorylated protein in each lane. The arrows indicate ERK1 (44 K) and ERK2 (42 K). Densitometric evaluation (g) shows values expressed relative to the optical density of the band representing phosphorylated ERK1 or 2 in the presence of TPA/KCl. Values are means±SEM of four independent experiments. *p < 0.05 vs the anisomycin-treated control (a JPG 57 kb) (b-c JPG 64 kb) (d-e JPG 59 kb) (f-g JPG 66 kb)
ESM Fig. 2
Internal mutation analysis. The plasmids −336 hInsLuc (wt) (2 μg), −336 hInsCRE2mutLuc (CRE2mt) (2μg), −336 hInsRIPE3bmutLuc (RIPE3bmt) (2μg) or −336hInsA1mutLuc (A1mt) (2μg) were transiently cotransfected with Bluescript (2μg; denoted by minus sign) or MEKK1 (2μg; denoted by plus sign). Luciferase activity is expressed relative to the mean value in each experiment measured in the wild type (wt) control (no MEKK1). Values are means±SEM of three independent experiments, each done in duplicate. *p < 0.05 vs the control in the absence of MEKK1 (JPG 42 kb)
ESM Table 1
Sequences of the oligonucleotides used for PCR (PDF 66 kb)
Rights and permissions
About this article
Cite this article
Oetjen, E., Blume, R., Cierny, I. et al. Inhibition of MafA transcriptional activity and human insulin gene transcription by interleukin-1β and mitogen-activated protein kinase kinase kinase in pancreatic islet beta cells. Diabetologia 50, 1678–1687 (2007). https://doi.org/10.1007/s00125-007-0712-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-007-0712-2