Abstract
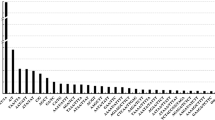
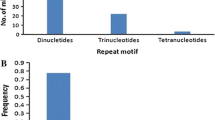
Ninety-four newly developed microsatellite markers were integrated into existing RFLP framework maps of four rice populations, including two doubled haploid, a recombinant inbred, and an interspecific backcross population. These simple sequence repeats (SSR) were predominantly poly(GA) motifs, targetted because of their abundance in rice. They were isolated from a previously described sheared library and a newly constructed enzyme-digested library. Differences in the average length of poly(GA) tracts were observed for clones isolated from the two libraries. The length of GA motifs averaged 21 repeat units for clones isolated from the Tsp-509-digested library, while motifs averaged 17 units for clones from the sheared library. There was no evidence of clustering of microsatellite markers near centromeres or telomeres. Mapping of the 94 newly developed markers as well as of 27 previously reported microsatellites provided genome-wide coverage of the 12 chromosomes, with an average distance of 1 SSLP (simple sequence repeat polymorphism) per 16–20 cM.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 13 February 1997/Accepted: 28 February 1997
Rights and permissions
About this article
Cite this article
Chen, X., Temnykh, S., Xu, Y. et al. Development of a microsatellite framework map providing genome-wide coverage in rice (Oryza sativa L.). Theor Appl Genet 95, 553–567 (1997). https://doi.org/10.1007/s001220050596
Issue Date:
DOI: https://doi.org/10.1007/s001220050596