Abstract
Key message
ClLOX, is located on chromosome 2 and encodes a lipoxygenase gene, which induced watermelon powdery mildew resistance by inhibiting pathogen spread.
Abstract
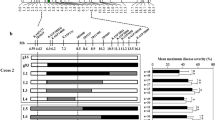
Powdery mildew is one of the most severe fungal diseases reducing yield and quality of watermelon (Citrullus lanatus L.) and other cucurbit crops. Genes responsible for powdery mildew resistance in watermelon are highly valuable. In this study, we first identified the QTL pm-lox for powdery mildew resistance in watermelon, located within a 0.93 Mb interval of chromosome 2, via XP-GWAS method using two F2 populations. The F2:3 families from one of the F2 populations were then used for fine-mapping the pm-lox locus into a 9,883 bp physical region between 29,581,906 and 29,591,789, containing only two annotated genes. Of these, only ClG42_02g0161300 showed a significant differential expression between the resistant and susceptible lines after powdery mildew inoculation based on RNA sequencing (RNA-seq) and qRT-PCR analysis, and is designated ClLOX. Derived Cleaved Amplified Polymorphic Sequence (dCAPs) markers were developed and validated. In addition, our tests showed that the resistance was anti-spread rather than anti-infection of the pathogen. This study identified a new resistance gene (ClLOX), provided insights into the mechanism of powdery mildew resistance, and developed a molecular marker for watermelon breeding.






Similar content being viewed by others
References
Ben-Naim Y, Cohen Y (2015) Inheritance of resistance to powdery mildew race 1W in watermelon. Phytopathology 105:1446–1457. https://doi.org/10.1094/PHYTO-02-15-0048-R
Christensen SA, Huffaker A, Kaplan F, Sims J, Ziemann S, Doehlemann G, Ji L, Schmitz RJ, Kolomiets MV, Alborn HT (2015) Maize death acids, 9-lipoxygenase–derived cyclopente (a) nones, display activity as cytotoxic phytoalexins and transcriptional mediators. Proc Natl Acad Sci USA 112:11407–11412. https://doi.org/10.1073/pnas.1511131112
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain W1118; iso-2; iso-3. Fly 6:80–92. https://doi.org/10.4161/fly.19695
Cohen R, Burger Y, Katzir N (2004) Monitoring physiological races of Podosphaera xanthii (syn. Sphaerotheca fuliginea), the causal agent of powdery mildew in cucurbits: factors affecting race identification and the importance for research and commerce. Phytoparasitica 32:174–183. https://doi.org/10.1007/BF02979784
Cui H, Zhu Z, Ding Z, Lv Y, Sun L, Luan F, Wang X (2021) First report of powdery mildew caused by Podosphaera xanthii race 1 on watermelon in China. J Plant Pathol 103:1029. https://doi.org/10.1007/s42161-021-00843-z
Cuthbert PA, Somers DJ, Brulé-Babel A (2007) Mapping of Fhb2 on chromosome 6BS: a gene controlling Fusarium head blight field resistance in bread wheat (Triticum aestivum L.). Theor Appl Genet 114:429–437. https://doi.org/10.1007/s00122-006-0439-3
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Davis AR, Levi A, Tetteh A, Wehner T, Russo V, Pitrat M (2007) Evaluation of watermelon and related species for resistance to race 1W powdery mildew. J Am Soc Hortic Sci 132:790–795. https://doi.org/10.21273/JASHS.132.6.790
De Waard M, Kipp E, Horn N, Van Nistelrooy J (1986) Variation in sensitivity to fungicides which inhibit ergosterol biosynthesis in wheat powdery mildew. Neth J Plant Pathol 92:21–32. https://doi.org/10.1007/BF01976373
Del Pino D, Olalla L, Pérez-García A, Rivera ME, García S, Moreno R, De Vicente A, Torés J (2002) Occurrence of races and pathotypes of cucurbit powdery mildew in southeastern Spain. Phytoparasitica 30:459–466. https://doi.org/10.1007/BF02979750
FAOSTAT (2021) http://www.faostat.com. Accessed 09 Oct 2023
Guo S, Zhao S, Sun H, Wang X, Wu S, Lin T, Ren Y, Gao L, Deng Y, Zhang J (2019) Resequencing of 414 cultivated and wild watermelon accessions identifies selection for fruit quality traits. Nat Genet 51:1616–1623. https://doi.org/10.1038/s41588-019-0518-4
Gyawali A, Shrestha V, Guill KE, Flint GS, Beissinger TM (2019) Single-plant GWAS coupled with bulk segregant analysis allows rapid identification and corroboration of plant-height candidate SNPs. BMC Plant Biol 19:1–15. https://doi.org/10.1186/s12870-019-2000-y
Han BK, Rhee SJ, Jang YJ, Sim TY, Kim YJ, Park TS, Lee GP (2016) Identification of a causal pathogen of watermelon powdery mildew in Korea and development of a genetic linkage marker for resistance in watermelon (Citrullus lanatus). Hortic Sci Technol 34:912–923. https://doi.org/10.12972/kjhst.20160095
Hendricks KEM, Roberts PD (2023) Evaluation of the sensitivity of Podosphaera xanthii to several fungicides for management of powdery mildew on squash in Florida. Crop Prot 172:106328. https://doi.org/10.1016/j.cropro.2023.106328
Hwang IS, Hwang BK (2010) The pepper 9-lipoxygenase gene CaLOX1 functions in defense and cell death responses to microbial pathogens. Plant Physiol 152:948–967. https://doi.org/10.1104/pp.109.147827
Iovieno P, Andolfo G, Schiavulli A, Catalano D, Ricciardi L, Frusciante L, Ercolano MR, Pavan S (2015) Structure, evolution and functional inference on the Mildew Locus O (MLO) gene family in three cultivated Cucurbitaceae spp.. BMC Genomics 16:1–13. https://doi.org/10.1186/s12864-015-2325-3
Jensen HP, Christensen E, Jørgensen JH (1992) Powdery mildew resistance genes in 127 Northwest European spring barley varieties. Plant Breed 108:210–228. https://doi.org/10.1111/j.1439-0523.1992.tb00122.x
Keinath AP (2000) Effect of protectant fungicide application schedules on gummy stem blight epidemics and marketable yield of watermelon. Plant Dis 84:254–260. https://doi.org/10.1094/PDIS.2000.84.3.254
Keinath AP (2015) Efficacy of fungicides against powdery mildew on watermelon caused by Podosphaera xanthii. Crop Prot 75:70–76. https://doi.org/10.1016/j.cropro.2015.05.013
Keinath AP, DuBose B (2004) Evaluation of fungicides for prevention and management of powdery mildew on watermelon. Crop Prot 23:35–42. https://doi.org/10.1016/S0261-2194(03)00165-0
Kim SH, Shin JE, Lee KJ, Xu SJ, Kim BS (2012) Evaluation of disease resistance of cucurbit cultivars to powdery mildew and root-knot nematode. Res Plant Dis 18:29–34. https://doi.org/10.5423/RPD.2012.18.1.029
Kim KH, Ahn SG, Hwang JH, Choi YM, Moon HS, Park YH (2013) Inheritance of resistance to powdery mildew in the watermelon and development of a molecular marker for selecting resistant plants. Hortic Environ Biotechnol 54:134–140. https://doi.org/10.1007/s13580-013-0156-1
Kim KH, Hwang JH, Han DY, Park M, Kim S, Choi D, Kim Y, Lee GP, Kim ST, Park YH (2015) Major quantitative trait loci and putative candidate genes for powdery mildew resistance and fruit-related traits revealed by an intraspecific genetic map for watermelon (Citrullus lanatus var. lanatus). PLoS One 10:e0145665. https://doi.org/10.1371/journal.pone.0145665
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL (2019) Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37:907–915. https://doi.org/10.1038/s41587-019-0201-4
Knight N, Sutherland M (2011) A rapid differential staining technique for Fusarium pseudograminearum in cereal tissues during crown rot infections. Plant Pathol 60:1140–1143. https://doi.org/10.1111/j.1365-3059.2011.02462.x
Kontaxis DG (1977) Powdery mildew on watermelon fruit in imperial valley, California. Plant Dis Rep 61:397
Kourelis J, Van Der Hoorn RA (2018) Defended to the nines: 25 years of resistance gene cloning identifies nine mechanisms for R protein function. Plant Cell 30:285–299. https://doi.org/10.1105/tpc.17.00579
Kousik CS, Donahoo R, Webster C, Turechek W, Adkins S, Roberts P (2011) Outbreak of cucurbit powdery mildew on watermelon fruit caused by Podosphaera xanthii in southwest Florida. Plant Dis 95:1586–1586. https://doi.org/10.1094/PDIS-06-11-0521
Kousik CS, Ikerd J, Mandal M, Adkins S, Turechek WW (2018) Watermelon germplasm lines USVL608-PMR, USVL255-PMR, USVL313-PMR, and USVL585-PMR with broad resistance to powdery mildew. HortScience 53:1212–1217. https://doi.org/10.21273/HORTSCI12979-18
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363. https://doi.org/10.1126/science.1166453
Křístková E, Lebeda A, Sedláková B (2009) Species spectra, distribution and host range of cucurbit powdery mildews in the Czech Republic, and in some other European and Middle Eastern countries. Phytoparasitica 37:337–350. https://doi.org/10.1007/s12600-009-0045-4
Lebeda A, McGrath MT, Sedláková B (2010) Fungicide resistance in cucurbit powdery mildew fungi. In: Carisse O (ed) Fungicides. InTech, Rijeka, pp 221–246
Lebeda A, Křístková E, Sedláková B, McCreight JD, Coffey MD (2016) Cucurbit powdery mildews: methodology for objective determination and denomination of races. Eur J Plant Pathol 144:399–410. https://doi.org/10.1007/s10658-015-0776-7
Liu T, Zhou Q, Wu Q, Xiong X (2016) Fluorescent staining with solophenyl flavine for infection observation in Phytophthora infestans. Plant Sci J 34:316–324. https://doi.org/10.11913/PSJ.2095-0837.2016.20316
Liu X, Meng G, Wang M, Qian Z, Zhang Y, Yang W (2021) Tomato SlPUB24 enhances resistance to Xanthomonas euvesicatoria pv. perforans race T3. Hortic Res. https://doi.org/10.1038/s41438-021-00468-4
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21. https://doi.org/10.1186/s13059-014-0550-8
Mandal MK, Suren H, Kousik C (2020) Elucidation of resistance signaling and identification of powdery mildew resistant mapping loci (ClaPMR2) during watermelon-Podosphaera xanthii interaction using RNA-Seq and whole-genome resequencing approach. Sci Rep 10:14038. https://doi.org/10.1038/s41598-020-70932-z
McGrath MT (2001) Fungicide resistance in cucurbit powdery mildew: experiences and challenges. Plant Dis 85:236–245. https://doi.org/10.1094/PDIS.2001.85.3.236
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Nascimento FA, Duarte HdSS, Souza FF, Ishikawa FH, Capucho AS (2020) Development and validation of a standard area diagram set to assess powdery mildew severity on watermelon leaves. Cienc Rural 50(10):e20200281. https://doi.org/10.1590/0103-8478cr20200281
Niks RE, Qi X, Marcel TC (2015) Quantitative resistance to biotrophic filamentous plant pathogens: concepts, misconceptions, and mechanisms. Annu Rev Phytopathol 53:445–470. https://doi.org/10.1146/annurev-phyto-080614-115928
Porta H, Rocha-Sosa M (2002) Plant lipoxygenases. Physiological and molecular features. Plant Physiol 130:15–21. https://doi.org/10.1104/pp.010787
Reis A, Buso JA (2004) Preliminary survey of Sphaerotheca fuliginea races occurring in cucurbits in Brazil. Hortic Bras 22(3):628–631. https://doi.org/10.1590/S0102-05362004000300026
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76. https://doi.org/10.1007/BF00020088
Shi Q, Fan J, Zhou Y, Zou Y, Duan X (2015) Triadimefon sensitivity and its correlation with the virulence population of Blumeria graminis f.sp. tritici in some wheat growing areas in 2012. Acta Phytopathol Sin 45(2):181–187. https://doi.org/10.11913/PSJ.2095-0837.2016.20316
Sun H, Wei J, Zhang J, Yang W (2014) A comparison of disease severity measurements using image analysis and visual estimates using a category scale for genetic analysis of resistance to bacterial spot in tomato. Eur J Plant Pathol 139:125–136. https://doi.org/10.1007/s10658-013-0371-8
Tetteh AY, Wehner TC, Davis AR (2010) Identifying resistance to powdery mildew race 2W in the USDA-ARS watermelon germplasm collection. Crop Sci 50:933–939. https://doi.org/10.2135/cropsci2009.03.0135
Tetteh AY, Wehner TC, Davis AR (2013) Inheritance of resistance to the new race of powdery mildew in watermelon. Crop Sci 53:880–887. https://doi.org/10.2135/cropsci2012.07.0453
Vasimuddin M, Misra S, Li H, Aluru S (2019) Efficient architecture-aware acceleration of BWA-MEM for multicore systems. In: 2019 IEEE international parallel and distributed processing symposium (IPDPS), Rio de Janeiro, pp 314–324. https://doi.org/10.1109/IPDPS.2019.00041.
Vellosillo T, Aguilera V, Marcos R, Bartsch M, Vicente J, Cascón T, Hamberg M, Castresana C (2013) Defense activated by 9-lipoxygenase-derived oxylipins requires specific mitochondrial proteins. Plant Physiol 161:617–627. https://doi.org/10.1104/pp.112.207514
Viswanath KK, Varakumar P, Pamuru RR, Basha SJ, Mehta S, Rao AD (2020) Plant lipoxygenases and their role in plant physiology. J Plant Biol 63:83–95. https://doi.org/10.1007/s12374-020-09241-x
Wang X, Wang L, Song J, Liu Y, Gao Y, Li J, Zhang H, Xu Y (2021) Identification of powdery mildew pathogen of watermelons and muskmelons in Ningxia and screening of resistant germplasm resources. J Gansu Agric Univ 56:83–91. https://doi.org/10.13432/j.cnki.jgsau.2021.05.012
Welling MT, Liu L, Kretzschmar T, Mauleon R, Ansari O, King GJ (2020) An extreme-phenotype genome-wide association study identifies candidate cannabinoid pathway genes in Cannabis. Sci Rep 10:18643. https://doi.org/10.1038/s41598-020-75271-7
Xiao N, Wu Y, Pan C, Yu L, Chen Y, Liu G, Li Y, Zhang X, Wang Z, Dai Z (2017) Improving of rice blast resistances in Japonica by pyramiding major R genes. Front Plant Sci 7:1918. https://doi.org/10.3389/fpls.2016.01918
Yadav V, Wang Z, Guo Y, Zhang X (2022) Comparative transcriptome profiling reveals the role of phytohormones and phenylpropanoid pathway in early-stage resistance against powdery mildew in watermelon (Citrullus lanatus L.). Front Plant Sci 13:1016822. https://doi.org/10.3389/fpls.2022.1016822
Yan L, Zhai Q, Wei J, Li S, Wang B, Huang T, Du M, Sun J, Kang L, Li C-B (2013) Role of tomato lipoxygenase D in wound-induced jasmonate biosynthesis and plant immunity to insect herbivores. PLoS Genet 9:e1003964. https://doi.org/10.1371/journal.pgen.1003964
Yang J, Jiang H, Yeh CT, Yu J, Jeddeloh JA, Nettleton D, Schnable PS (2015) Extreme-phenotype genome-wide association study (XP-GWAS): a method for identifying trait-associated variants by sequencing pools of individuals selected from a diversity panel. Plant J 84:587–596. https://doi.org/10.1111/tpj.13029
Zhang H, Guo S, Gong G, Ren Y, Davis AR, Xu Y (2011) Sources of resistance to race 2WF powdery mildew in U.S. watermelon plant introductions. HortScience 46:1349–1352. https://doi.org/10.21273/HORTSCI.46.10.1349
Zhang Y, Huang C, van Peer AF, Sonnenberg AS, Zhao M, Gao W (2022) Fine mapping and functional analysis of the gene PcTYR, involved in control of cap color of Pleurotus cornucopiae. Appl Environ Microbiol 88:e02173-e2221. https://doi.org/10.1128/aem.02173-21
Zhou X, Stephens M (2012) Genome-wide efficient mixed-model analysis for association studies. Nat Genet 44:821–824. https://doi.org/10.1038/ng.2310
Acknowledgements
This work was supported by the Provincial Technology Innovation Program of Shandong, Ningxia Hui Autonomous Region Agricultural Breeding Special Project (NXNYYZ202001), Ningbo Science and Technology Innovation Project (2021Z132), Weifang Seed Innovation Group, Modern Agro–industry Technology Research System (CARS–25), National Natural Science Foundation of China (32302558), Natural Science Foundation of Jiangsu Province (BK20220305), Jiangsu Seed Industry Revitalization Competitive Project (JBGS (2021) 072), and Jiangsu Agricultural Innovation of New Cultivars (PZCZ201717).
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
YD and XL performed most of the experiments, analyzed the data and drafted the manuscript. SL performed bioinformatics analysis. XNL, XL, LX and TB performed disease evaluation and phenotyping. BX and YS conducted the field experiments. XZ, YS and GL modified the manuscript. YD, XZ, YS and GL designed and directed the entire study. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Amit Gur.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Deng, Y., Liu, X., Liu, S. et al. Fine mapping of ClLOX, a QTL for powdery mildew resistance in watermelon (Citrullus lanatus L.). Theor Appl Genet 137, 51 (2024). https://doi.org/10.1007/s00122-023-04520-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04520-w