Abstract
Key message
A single nucleotide (G) deletion in the third exon of BraA02.PES2-2 (Bra032957) leads to the conversion of flower color from yellow to white in B. rapa, and knockout mutants of its orthologous genes in B. napus showed white or pale yellow flowers.
Abstract
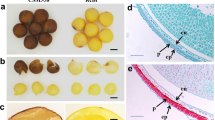
Brassica rapa (2n = 20, AA) is grown worldwide as an important crop for edible oil and vegetables. The bright yellow flower color and long-lasting flowering period give it aesthetic qualities appealing to countryside tourists. However, the mechanism controlling the accumulation of yellow pigments in B. rapa has not yet been completely revealed. In this study, we characterized the mechanism of white flower formation using a white-flowered natural B. rapa mutant W01. Compared to the petals of yellow-flowered P3246, the petals of W01 have significantly reduced content of yellowish carotenoids. Furthermore, the chromoplasts in white petals of W01 are abnormal with irregularly structured plastoglobules. Genetic analysis indicated that the white flower was controlled by a single recessive gene. By combining BSA-seq with fine mapping, we identified the target gene BraA02.PES2-2 (Bra032957) homologous to AtPES2, which has a single nucleotide (G) deletion in the third exon. Seven homologous PES2 genes including BnaA02.PES2-2 (BnaA02g28340D) and BnaC02.PES2-2 (BnaC02g36410D) were identified in B. napus (2n = 38, AACC), an allotetraploid derived from B. rapa and B. oleracea (2n = 18, CC). Knockout mutants of either one or two of BnaA02.PES2-2 and BnaC02.PES2-2 in the yellow-flowered B. napus cv. Westar by the CRISPR/Cas9 system showed pale-yellow or white flowers. The knock-out mutants of BnaA02.PES2-2 and BnaC02.PES2-2 had fewer esterified carotenoids. These results demonstrated that BraA02.PES2-2 in B. rapa, and BnaA02.PES2-2 and BnaC02.PES2-2 in B. napus play important roles in carotenoids esterification in chromoplasts that contributes to the accumulation of carotenoids in flower petals.




Similar content being viewed by others
Data availability
All data generated or analyzed in this study are included in this published article and its supplementary information files.
References
Ariizumi T, Kishimoto S, Kakami R, Maoka T, Hirakawa H, Suzuki Y, Ozeki Y, Shirasawa K, Bernillon S, Okabe Y, Moing A, Asamizu E, Rothan C, Ohmiya A, Ezura H (2014) Identification of the carotenoid modifying gene PALE YELLOW PETAL 1 as an essential factor in xanthophyll esterification and yellow flower pigmentation in tomato (Solanum lycopersicum). Plant J 79:453–465. https://doi.org/10.1111/tpj.12570
Belser C, Istace B, Denis E, Dubarry M, Baurens FC, Falentin C, Genete M, Berrabah W, Chèvre AM, Delourme R, Deniot G, Denoeud F, Duffé P, Engelen S, Lemainque A, Manzanares-Dauleux M, Martin G, Morice J, Noel B, Vekemans X, D’Hont A, Rousseau-Gueutin M, Barbe V, Cruaud C, Wincker P, Aury JM (2018) Chromosome-scale assemblies of plant genomes using nanopore long reads and optical maps. Nat Plants 4:879–887. https://doi.org/10.1038/s41477-018-0289-4
Brandi G, Béchir M, Sailer S, Haberthür C, Stocker R, Stover JF (2010) Transcranial color-coded duplex sonography allows to assess cerebral perfusion pressure noninvasively following severe traumatic brain injury. Acta Neurochir (wien) 152:965–972. https://doi.org/10.1007/s00701-010-0643-4
Campbell DR, Bischoff M, Lord JM, Robertson AW (2010) Flower color influences insect visitation in alpine New Zealand. Ecology 91:2638–2649. https://doi.org/10.1890/09-0941.1
Campbell NR, Harmon SA, Narum SR (2015) Genotyping-in-thousands by sequencing (GT-seq): a cost effective SNP genotyping method based on custom amplicon sequencing. Mol Ecol Resour 15:855–867. https://doi.org/10.1111/1755-0998.12357
Cao H, Zhang J, Xu J, Ye J, Yun Z, Xu Q, Xu J, Deng X (2012) Comprehending crystalline β-carotene accumulation by comparing engineered cell models and the natural carotenoid-rich system of citrus. J Exp Bot 63:4403–4417. https://doi.org/10.1093/jxb/ers115
Chalhoub B, Denoeud F, Liu S et al (2014) Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345:950–953. https://doi.org/10.1126/science.1253435
Chao H, Li T, Luo C, Huang H, Ruan Y, Li X, Niu Y, Fan Y, Sun W, Zhang K, Li J, Qu C, Lu K (2020) BrassicaEDB: a gene expression database for Brassica crops. Int J Mol Sci 21:5831. https://doi.org/10.3390/ijms21165831
Chen B, Heneen W, Jönsson R (1988) Independent inheritance of erucic acid content and flower colour in the C-genome of Brassica napus L. Plant Breed 100:147–149. https://doi.org/10.1111/j.1439-0523.1988.tb00230.x
Egea I, Barsan C, Bian W, Purgatto E, Latche A, Chervin C, Bouzayen M, Pech J (2010) Chromoplast differentiation:current status and perspectives. Plant Cell Physiol 51:1601–1611. https://doi.org/10.1093/pcp/pcq136
Fu W, Chen D, Pan Q, Li F, Zhao Z, Ge X, Li Z (2018) Production of red-flowered oilseed rape via the ectopic expression of Orychophragmus violaceus OvPAP2. Plant Biotechnol J 16:367–380. https://doi.org/10.1111/pbi.12777
Fu H, Chao H, Zhao X, Wang H, Li H, Zhao W, Sun T, Li M, Huang J (2022) Anthocyanins identification and transcriptional regulation of anthocyanin biosynthesis in purple Brassica napus. Plant Mol Biol 110:53–68. https://doi.org/10.1007/s11103-022-01285-6
Geyer R, Peacock AD, White DC, Lytle C, Van GJ (2014) Atmospheric pressure chemicalionization and atmospheric pressure photoionization forsimultaneous mass spectrometric analysis of microbial respiratory ubiquinones and menaquinones. J Mass Spectrom 39:922–929. https://doi.org/10.1002/jms.670
Grotewold E (2006) The genetics and biochemistry of floral pigments. Annu Rev Plant Biol 57:761–780. https://doi.org/10.1146/annurev.arplant.57.032905.105248
Han F, Cui H, Zhang B, Liu X, Yang L, Zhuang M, Lv H, Li Z, Wang Y, Fang Z, Song J, Zhang Y (2019) Map-based cloning and characterization of BoCCD4, a gene responsible for white/yellow petal color in B. oleracea. BMC Genomics 20:242. https://doi.org/10.1186/s12864-019-5596-2
Hansmann P, Sitte P (1982) Composition and molecular structure of chromoplast globules of Viola tricolor. Plant Cell Rep 1:111–114. https://doi.org/10.1007/BF00272366
Hao P, Liu H, Lin B, Ren Y, Huang L, Jiang L, Hua S (2022) BnaA0.3 ANS identified by metabolomics and RNA-seq partly played irreplaceable role in pigmentation of red rapeseed (Brassica napus) petal. Front Plant Sci 13:940765. https://doi.org/10.3389/fpls.2022.940765
Heneen W, Chen B, Cheng B, Jonsson A, Simonsen V, Jørgensen R, Davik J (1995) Characterization of the A and C genomes of Brassica Campestris and B. Alboglabra Hereditas 123:251–267. https://doi.org/10.1111/j.1601-5223.1995.00251.x
Hornero-Méndez D, Mínguez-Mosquera MI (2000) Carotenoid pigments in Rosa mosqueta hips, an alternative carotenoid source for foods. J Agric Food Chem 48:825–828. https://doi.org/10.1021/jf991136n
Huang Z, Ban Y, Bao R, Zhang X, Xu A, Ding J (2014) Inheritance and gene mapping of the white flower in Brassica napus L. New Zeal J Crop Hort 42:111–117. https://doi.org/10.1080/01140671.2013.863211
Jia L, Wang J, Wang R, Duan M, Qiao C, Chen X, Ma G, Zhou X, Zhu M, Jing F, Zhang S, Qu C, Li J (2021) Comparative transcriptomic and metabolomic analyses of carotenoid biosynthesis reveal the basis of white petal color in Brassica napus. Planta 253:8. https://doi.org/10.1007/s00425-020-03536-6
Jones KN, Reithel JS (2001) Pollinator-mediated selection on a flower color polymorphism in experimental populations of Antirrhinum (Scrophulariaceae). Am J Bot 88:447–454. https://doi.org/10.2307/2657109
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li L, Yuan H (2013) Chromoplast biogenesis and carotenoid accumulation. Arch Biochem Biophys 539:102–109. https://doi.org/10.1016/j.abb.2013.07.002
Lichtenthaler HK, Buschmann C (2001) Chlorophylls and carotenoids: measurement and characterization by UV–VIS spectroscopy. Curr Protoc Food Analyt Chem 1:431–438. https://doi.org/10.1002/0471142913.faf0403s01
Lippold F, vom Dorp K, Abraham M, Hölzl G, Wewer V, Yilmaz JL, Lager I, Montandon C, Besagni C, Kessler F, Stymne S, Dörmann P (2012) Fatty acid phytyl ester synthesis in chloroplasts of Arabidopsis. Plant Cell 24:2001–2014. https://doi.org/10.1105/tpc.112.095588
Liu XP, Tu JX, Chen BY, Fu TD (2004) Identification of the linkage relationship between the flower colour and the content of erucic acid in the resynthesized Brassica napus L. Yi Chuan Xue Bao 31:357–362
Liu Y, Ye S, Yuan G, Ma X, Heng S, Yi B, Ma C, Shen J, Tu J, Fu T, Wen J (2020) Gene silencing of BnaA09.ZEP and BnaC09.ZEP confers orange color in Brassica napus flowers. Plant J 104:932–949. https://doi.org/10.1111/tpj.14970
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Ljubesić N, Wrischer M, Devidé Z (1991) Chromoplasts–the last stages in plastid development. Int J Dev Biol 35:251–258
Mahmood S, Li Z, Yue X, Wang B, Chen J, Liu K (2016) Development of INDELs markers in oilseed rape (Brassica napus L.) using re-sequencing data. Mol Breed 36:79. https://doi.org/10.1007/s11032-016-0501-z
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Mínguez-Mosquera MI, Hornero-Méndez D (1994) Formation and transformation of pigments during the fruit ripening of Capsicum annuum cv. Bola and Agridulce. J Agric Food Chem 42:38–44. https://doi.org/10.1021/jf00037a005
Mohammad A, Sikka SM, Aziz MA (1942) Inheritance of seed color in some oleiferous Brassiceae. Indian J Genet 2:112–127
Nagaharu U (1935) Genomic analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Ohmiya A, Kishimoto S, Aida R, Yoshioka S, Sumitomo K (2006) Carotenoid cleavage dioxygenase (CmCCD4a) contributes to white color formation in chrysanthemum petals. Plant Physiol 142:1193–1201. https://doi.org/10.1104/pp.106.087130
Pearson OH (1929) A dominant white flower color in Brassica oleracea L. Am Nat 63:561–565. https://doi.org/10.1086/280291
Rahman MH (2001) Inheritance of petal color and its independent segregation from seed color in Brassica rapa. Plant Breed 120:197–200. https://doi.org/10.1046/j.1439-0523.2001.00607.x
Schweiggert RM, Carle R (2017) Carotenoid deposition in plant and animal foods and its impact on bioavailability. Crit Rev Food Sci Nutr 9:1807–1830. https://doi.org/10.1080/10408398.2015.1012756
Schweiggert R, Mezger D, Schimpf F, Steingass C, Carle R (2012) Influence of chromoplast morphology on carotenoid bioaccessibility of carrot, mango, papaya, and tomato. Food Chem 135:2736–2742. https://doi.org/10.1016/j.foodchem.2012.07.035
Shumskaya M, Wurtzel ET (2013) The carotenoid biosynthetic pathway: thinking in all dimensions. Plant Sci 208:58–63. https://doi.org/10.1016/j.plantsci.2013.03.012
Song JM, Guan Z, Hu J, Guo C, Yang Z, Wang S, Liu D, Wang B, Lu S, Zhou R, Xie WZ, Cheng Y, Zhang Y, Liu K, Yang QY, Chen LL, Guo L (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6:34–45. https://doi.org/10.1038/s41477-019-0577-7
Sriboon S, Li H, Guo C, Senkhamwong T, Dai C, Liu K (2020) Knock-out of TERMINAL FLOWER 1 genes altered flowering time and plant architecture in Brassica napus. BMC Genet 21:52. https://doi.org/10.1186/s12863-020-00857-z
Steinmüller D, Tevini M (1985) Composition and function of plastoglobuli :I. Isolation and purification from chloroplasts and chromoplasts. Planta 163:201–207. https://doi.org/10.1007/BF00393507
Tan K, Stupack DG, Wilkinson MF (2022) Nonsense-mediated RNA decay: an emerging modulator of malignancy. Nat Rev Cancer 22:437–451. https://doi.org/10.1038/s41568-022-00481-2
Tang T, Yu X, Yang H, Gao Q, Ji H, Wang Y, Yan G, Peng Y, Luo H, Liu K, Li X, Ma C, Kang C, Dai C (2018) Development and validation of an effective CRISPR/Cas9 vector for efficiently isolating positive transformants and transgene-free mutants in a wide range of plant species. Front Plant Sci 9:1533. https://doi.org/10.3389/fpls.2018.01533
Wang X, Wang H, Wang J et al (2011) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1039. https://doi.org/10.1038/ng.919
Yamamizo C, Kishimoto S, Ohmiya A (2010) Carotenoid composition and carotenogenic gene expression during Ipomoea petal development. J Exp Bot 61:709–719. https://doi.org/10.1093/jxb/erp335
Yang H, Wu JJ, Tang T, Liu KD, Dai C (2017) CRISPR/Cas9-mediated genome editing efficiently creates specific mutations at multiple loci using one sgRNA in Brassica napus. Sci Rep 7:7489. https://doi.org/10.1038/s41598-018-23161-4
Yang S, Tian X, Wang Z, Wei X, Zhao Y, Su H, Zhao X, Tian B, Yuan Y, Zhang XW (2021) Fine mapping and candidate gene identification of a white flower gene BrWF3 in Chinese cabbage (Brassica rapa L. ssp. pekinensis). Front Plant Sci 12:646222. https://doi.org/10.3389/fpls.2021.646222
Yang S, Liu H, Zhao Y, Su H, Wei X, Wang Z, Zhao X, Zhang XW, Yuan Y (2022) Map-based cloning and characterization of Br-dyp1, a gene conferring dark yellow petal color trait in Chinese cabbage (Brassica rapa L. ssp pekinensis). Front Plant Sci 13:841328. https://doi.org/10.3389/fpls.2022.841328
Ye S, Hua S, Ma T, Ma X, Chen Y, Wu L, Zhao L, Yi B, Ma C, Tu J, Shen J, Fu T, Wen J (2022) Genetic and multi-omics analyses reveal BnaA07.PAP2In-184-317 as the key gene conferring anthocyanin-based color in Brassica napus flowers. J Exp Bot 73:6630–6645. https://doi.org/10.1093/jxb/erac312
Yi B, Zeng F, Lei S, Chen Y, Yao X, Zhu Y, Wen J, Shen J, Ma C, Tu J, Fu T (2010) Two duplicate CYP704B1-homologous genes BnMs1 and BnMs2 are required for pollen exine formation and tapetal development in Brassica napus. Plant J 63:925–938. https://doi.org/10.1111/j.1365-313X.2010.04289.x
Ytterberg AJ, Peltier JB, van Wijk KJ (2006) Protein profiling of plastoglobules in chloroplasts and chromoplasts. A surprising site for differential accumulation of metabolic enzymes. Plant Physiol 140:984–997. https://doi.org/10.1104/pp.105.076083
Yuan H, Zhang JX, Nageswaran D, Li L (2015) Carotenoid metabolism and regulation in horticultural crops. Hortic Res 2:15036. https://doi.org/10.1038/hortres.2015.36
Zhang B, Lu C, Kakihara F, Kato M (2002) Effect of genome composition and cytoplasm on petal colour in resynthesized amphidiploids and sesquidiploids derived from crosses between Brassica rapa and Brassica oleracea. Plant Breed 121:297–300. https://doi.org/10.1046/j.1439-0523.2002.722295.x
Zhang B, Liu C, Wang YQ, Yao X, Wang F, Wu JS, King GJ, Liu KD (2015) Disruption of a CAROTENOID CLEAVAGE DIOXYGENASE 4 gene converts flower color from white to yellow in Brassica species. New Phytol 206:1513–1526. https://doi.org/10.1111/nph.13335
Zhang XX, Li R, Chen L, Niu SL, Chen L, Gao J, Wen J, Yi B, Ma CZ, Tu JX, Fu TD, Shen JX (2018a) Fine-mapping and candidate gene analysis of the Brassica juncea white-flowered mutant Bjpc2 using the whole-genome resequencing. Mol Genet Genomics 293:359–370. https://doi.org/10.1007/s00438-017-1390-5
Zhang XX, Li RH, Chen L, Niu SL, Li Q, Xu K, Wen J, Yi B, Ma CZ, Tu JX, Fu TD, Shen JX (2018b) Inheritance and gene mapping of the white flower trait in Brassica juncea. Mol Breed 38:20–29. https://doi.org/10.1007/s11032-017-0771-0
Zhang N, Chen L, Ma S, Wang RF, He Q, Tian M, Zhang LG (2020) Fine mapping and candidate gene analysis of the white flower gene Brwf in Chinese cabbage (Brassica rapa L.). Sci Rep 10:6080. https://doi.org/10.1038/s41598-020-63165-7
Zhu C, Yang Q, Ni X, Bai C, Sheng Y, Shi L, Capell T, Sandmann G, Christou P (2014) Cloning and functional analysis of the promoters that upregulate carotenogenic gene expression during flower development in Gentiana lutea. Physiol Plant 150:493–504. https://doi.org/10.1111/ppl.12129
Zhu C, Bai C, Sanahuja G, Yuan D, Farré G, Naqvi S, Shi L, Capell T, Christou P (2010) The regulation of carotenoid pigmentation in flowers. Arch Biochem Biophys 504:132–141. https://doi.org/10.1016/j.abb.2010.07.028
Acknowledgements
The research was supported by the National Key Research and Development Program of China (2016YFD0101007AA102602).
Funding
The research was supported by the National Key Research and Development Program of China (2016YFD0101007AA102602). Zhilin Guan, Xuewei Li, Jianshun Yang, Junwei Zhao, Kaiyue Wang, Jianlin Hu, Bao Zhang and Kede Liu were supported by the National Key Research and Development Program of China (2016YFD0101007AA102602). The funders had no role in study disign, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
ZG, XL, JY, JZ, and KW performed the experiments. ZG, XL, and KL wrote the manuscript. BW helped analyze the data. KL conceived and supervised the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose. On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethics approval
The authors declare that the experiments comply with the current laws of the country in which they were performed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guan, Z., Li, X., Yang, J. et al. The mechanism of white flower formation in Brassica rapa is distinct from that in other Brassica species. Theor Appl Genet 136, 133 (2023). https://doi.org/10.1007/s00122-023-04344-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04344-8