Abstract
Key message
Phenomic prediction of wheat grain yield and heading date in different multi-environmental trial scenarios is accurate. Modelling the genotype-by-environment interaction effect using phenomic data is a potentially low-cost complement to genomic prediction.
Abstract
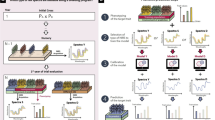
The performance of wheat cultivars in multi-environmental trials (MET) is difficult to predict because of the genotype-by-environment interactions (G × E). Phenomic selection is supposed to be efficient for modelling the G × E effect because it accounts for non-additive effects. Here, phenomic data are near-infrared (NIR) spectra obtained from plant material. While phenomic selection has recently been shown to accurately predict wheat grain yield in single environments, its accuracy needs to be investigated for MET. We used four datasets from two winter wheat breeding programs to test and compare the predictive abilities of phenomic and genomic models for grain yield and heading date in different MET scenarios. We also compared different methods to model the G × E using different covariance matrices based on spectra. On average, phenomic and genomic prediction abilities are similar in all different MET scenarios. Better predictive abilities were obtained when G × E effects were modelled with NIR spectra than without them, and it was better to use all the spectra of all genotypes in all environments for modelling the G × E. To facilitate the implementation of phenomic prediction, we tested MET designs where the NIR spectra were measured only on the genotype–environment combinations phenotyped for the target trait. Missing spectra were predicted with a weighted multivariate ridge regression. Intermediate predictive abilities for grain yield were obtained in a sparse testing scenario and for new genotypes, which shows that phenomic selection is an efficient and practicable prediction method for dealing with G × E.



Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are not publicly available due to the breeding program privacy policy, but are available from the corresponding author on reasonable request.
Code availability
Code used to run the analysis is available from the corresponding author on request.
References
Albrecht T, Wimmer V, Auinger H-J et al (2011) Genome-based prediction of testcross values in maize. Theor Appl Genet 123:339–350. https://doi.org/10.1007/s00122-011-1587-7
Allard RW, Bradshaw AD (1964) Implications of genotype-environmental interactions in applied plant breeding 1. Crop Sci 4:503–508. https://doi.org/10.2135/cropsci1964.0011183X000400050021x
Azodi CB, Pardo J, VanBuren R et al (2020) Transcriptome-based prediction of complex traits in maize. Plant Cell 32:139–151. https://doi.org/10.1105/tpc.19.00332
Bernardo R (1994) Prediction of maize single-cross performance using RFLPs and information from related hybrids. Crop Sci 34:20–25. https://doi.org/10.2135/cropsci1994.0011183X003400010003x
Brault C, Lazerges J, Doligez A et al (2021) Interest of phenomic prediction as an alternative to genomic prediction in grapevine. Genetics 31:277
Burgueño J, de los Campos G, Weigel K, Crossa J (2012) Genomic prediction of breeding values when modeling genotype × environment interaction using pedigree and dense molecular markers. Crop Sci 52:707–719. https://doi.org/10.2135/cropsci2011.06.0299
Comstock RE, Moll RH (1963) Genotype x Environment Interactions. Stat Genet Plant Breed
Covarrubias-Pazaran G (2016) Genome-assisted prediction of quantitative traits using the R package sommer. PLoS ONE 11:e0156744. https://doi.org/10.1371/journal.pone.0156744
Crossa J, de los Campos G, Pérez P et al (2010) Prediction of genetic values of quantitative traits in plant breeding using pedigree and molecular markers. Genetics 186:713–724. https://doi.org/10.1534/genetics.110.118521
Cuevas J, Crossa J, Soberanis V et al (2016) Genomic prediction of genotype × environment interaction kernel regression models. Plant Genome. https://doi.org/10.3835/plantgenome2016.03.0024
Cuevas J, Montesinos-López O, Juliana P et al (2019) Deep kernel for genomic and near infrared predictions in multi-environment breeding trials. G3 Genes Genomes Genetics 9:2913–2924. https://doi.org/10.1534/g3.119.400493
Daetwyler HD, Calus MPL, Pong-Wong R et al (2013) Genomic prediction in animals and plants: simulation of data, validation, reporting, and benchmarking. Genetics 193:347–365. https://doi.org/10.1534/genetics.112.147983
Damesa T, Worku M, Möhring J, Piepho HP (2017) One step at a time: stage-wise analysis of a series of experiments. Agron J 109:845–857. https://doi.org/10.2134/agronj2016.07.0395
De Los Campos G, Gianola D, Rosa GJM et al (2010) Semi-parametric genomic-enabled prediction of genetic values using reproducing kernel Hilbert spaces methods. Genet Res 92:295–308. https://doi.org/10.1017/S0016672310000285
De Los Campos G, Naya H, Gianola D et al (2009) Predicting quantitative traits with regression models for dense molecular markers and pedigree. Genetics 182:375–385. https://doi.org/10.1534/genetics.109.101501
Endelman JB (2011) Ridge regression and other kernels for genomic selection with R package rrBLUP. Plant Genome 4:250–255. https://doi.org/10.3835/plantgenome2011.08.0024
Endelman JB, Jannink J-L (2012) Shrinkage estimation of the realized relationship matrix. G3 Genes Genomes Genetics 2:1405–1413. https://doi.org/10.1534/g3.112.004259
Forni S, Aguilar I, Misztal I (2011) Different genomic relationship matrices for single-step analysis using phenotypic, pedigree and genomic information. Genet Sel Evol 43:1. https://doi.org/10.1186/1297-9686-43-1
Friedman J, Hastie T, Tibshirani R (2010) Regularization paths for generalized linear models via coordinate descent. J Stat Softw. https://doi.org/10.18637/jss.v033.i01
Frisch M, Thiemann A, Fu J et al (2010) Transcriptome-based distance measures for grouping of germplasm and prediction of hybrid performance in maize. Theor Appl Genet 120:441–450. https://doi.org/10.1007/s00122-009-1204-1
Fu J, Falke KC, Thiemann A et al (2012) Partial least squares regression, support vector machine regression, and transcriptome-based distances for prediction of maize hybrid performance with gene expression data. Theor Appl Genet 124:825–833. https://doi.org/10.1007/s00122-011-1747-9
Galán RJ, Bernal-Vasquez A-M, Jebsen C et al (2020) Integration of genotypic, hyperspectral, and phenotypic data to improve biomass yield prediction in hybrid rye. Theor Appl Genet 133:3001–3015. https://doi.org/10.1007/s00122-020-03651-8
Galán RJ, Bernal-Vasquez A-M, Jebsen C et al (2021) Early prediction of biomass in hybrid rye based on hyperspectral data surpasses genomic predictability in less-related breeding material. Theor Appl Genet 134:1409–1422. https://doi.org/10.1007/s00122-021-03779-1
Gemmer MR, Richter C, Jiang Y et al (2020) Can metabolic prediction be an alternative to genomic prediction in barley? PLoS ONE 15:e0234052. https://doi.org/10.1371/journal.pone.0234052
Guo Z, Magwire MM, Basten CJ et al (2016) Evaluation of the utility of gene expression and metabolic information for genomic prediction in maize. Theor Appl Genet 129:2413–2427. https://doi.org/10.1007/s00122-016-2780-5
Heffner EL, Jannink J-L, Iwata H et al (2011) Genomic selection accuracy for grain quality traits in biparental wheat populations. Crop Sci 51:2597–2606. https://doi.org/10.2135/cropsci2011.05.0253
Heslot N, Akdemir D, Sorrells ME, Jannink J-L (2014) Integrating environmental covariates and crop modeling into the genomic selection framework to predict genotype by environment interactions. Theor Appl Genet 127:463–480. https://doi.org/10.1007/s00122-013-2231-5
Heslot N, Jannink J-L, Sorrells ME (2013) Using genomic prediction to characterize environments and optimize prediction accuracy in applied breeding data. Crop Sci 53:921–933. https://doi.org/10.2135/cropsci2012.07.0420
Jarquín D, Crossa J, Lacaze X et al (2014) A reaction norm model for genomic selection using high-dimensional genomic and environmental data. Theor Appl Genet 127:595–607. https://doi.org/10.1007/s00122-013-2243-1
Jarquín D, Lemes da Silva C, Gaynor RC et al (2017) Increasing genomic-enabled prediction accuracy by modeling genotype × environment interactions in Kansas wheat. Plant Genome. https://doi.org/10.3835/plantgenome2016.12.0130
Kang HM, Sul JH, Service SK et al (2010) Variance component model to account for sample structure in genome wide association studies. Nat Genet 42:348–354. https://doi.org/10.1038/ng.548
Kitt J, Danguy des Désert A, Bouchet S, Servin B, Rimbert H, de Oliveira R, Choulet F, Balfourier F, Sourdille P, Paux E (2021) Genotyping of 4,506 bread wheat accessions with the TaBW410K SNP array. Zenodo. https://doi.org/10.5281/zenodo.4518374
Krause MR, González-Pérez L, Crossa J et al (2019) Hyperspectral reflectance-derived relationship matrices for genomic prediction of grain yield in wheat. G3 Genes Genomes Genetics. https://doi.org/10.1534/g3.118.200856
Lado B, Barrios PG, Quincke M et al (2016) Modeling genotype × environment interaction for genomic selection with unbalanced data from a wheat breeding program. Crop Sci 56:2165–2179. https://doi.org/10.2135/cropsci2015.04.0207
Lane HM, Murray SC, Montesinos-López OA et al (2020) Phenomic selection and prediction of maize grain yield from near-infrared reflectance spectroscopy of kernels. Plant Phenome J. https://doi.org/10.1002/ppj2.20002
Legarra A, Aguilar I, Misztal I (2009) A relationship matrix including full pedigree and genomic information. J Dairy Sci 92:4656–4663. https://doi.org/10.3168/jds.2009-2061
Lopez-Cruz M, Crossa J, Bonnett D et al (2015) Increased prediction accuracy in wheat breeding trials using a marker × environment interaction genomic selection model. G3 Genes Genomes Genetics 5:569–582. https://doi.org/10.1534/g3.114.016097
Ly D, Chenu K, Gauffreteau A et al (2017) Nitrogen nutrition index predicted by a crop model improves the genomic prediction of grain number for a bread wheat core collection. Field Crops Res 214:331–340. https://doi.org/10.1016/j.fcr.2017.09.024
Ly D, Huet S, Gauffreteau A et al (2018) Whole-genome prediction of reaction norms to environmental stress in bread wheat (Triticum aestivum L.) by genomic random regression. Field Crops Res 216:32–41. https://doi.org/10.1016/j.fcr.2017.08.020
Malosetti M, Bustos-Korts D, Boer MP, van Eeuwijk FA (2016) Predicting responses in multiple environments: issues in relation to genotype × environment interactions. Crop Sci 56:2210. https://doi.org/10.2135/cropsci2015.05.0311
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829. https://doi.org/10.1093/genetics/157.4.1819
Michel S, Wagner C, Nosenko T et al (2021) Merging genomics and transcriptomics for predicting fusarium head blight resistance in wheat. Genes 12:114. https://doi.org/10.3390/genes12010114
Montesinos-López A, Montesinos-López OA, Cuevas J et al (2017) Genomic Bayesian functional regression models with interactions for predicting wheat grain yield using hyper-spectral image data. Plant Methods. https://doi.org/10.1186/s13007-017-0212-4
Osborne BG (2006) Applications of near infrared spectroscopy in quality screening of early-generation material in cereal breeding programmes. J Infrared Spectrosc 14:93–101. https://doi.org/10.1255/jnirs.595
Parmley K, Nagasubramanian K, Sarkar S et al (2019) Development of optimized phenomic predictors for efficient plant breeding decisions using phenomic-assisted selection in soybean. Plant Phenomics 2019:1–15. https://doi.org/10.34133/2019/5809404
Pérez P, de los Campos G (2014) Genome-wide regression and prediction with the BGLR statistical package. Genetics 198:483–495. https://doi.org/10.1534/genetics.114.164442
Pérez-Rodríguez P, Crossa J, Rutkoski J et al (2017) Single-step genomic and pedigree genotype × environment interaction models for predicting wheat lines in international environments. Plant Genome. https://doi.org/10.3835/plantgenome2016.09.0089
Pszczola M, Strabel T, Mulder HA, Calus MPL (2012) Reliability of direct genomic values for animals with different relationships within and to the reference population. J Dairy Sci 95:389–400. https://doi.org/10.3168/jds.2011-4338
Riedelsheimer C, Czedik-Eysenberg A, Grieder C et al (2012) Genomic and metabolic prediction of complex heterotic traits in hybrid maize. Nat Genet 44:217
Rimbert H, Darrier B, Navarro J et al (2018) High throughput SNP discovery and genotyping in hexaploid wheat. PLoS ONE 13:e0186329. https://doi.org/10.1371/journal.pone.0186329
Rincent R, Charpentier J-P, Faivre-Rampant P et al (2018) Phenomic selection is a low-cost and high-throughput method based on indirect predictions: proof of concept on wheat and poplar. G3 Genes Genomes Genetics. https://doi.org/10.1534/g3.118.200760
Rincent R, Laloë D, Nicolas S et al (2012) Maximizing the reliability of genomic selection by optimizing the calibration set of reference individuals: comparison of methods in two diverse groups of maize inbreds (Zea mays L.). Genetics 192:715–728. https://doi.org/10.1534/genetics.112.141473
Rincent R, Malosetti M, Ababaei B et al (2019) Using crop growth model stress covariates and AMMI decomposition to better predict genotype-by-environment interactions. Theor Appl Genet 132:3399–3411. https://doi.org/10.1007/s00122-019-03432-y
Robert P, Auzanneau J, Goudemand E et al (2022a) Phenomic selection in wheat breeding: identification and optimisation of factors influencing prediction accuracy and comparison to genomic selection. Theor Appl Genet. https://doi.org/10.1007/s00122-021-04005-8
Robert P, Brault C, Rincent R, Segura V (2022b) Phenomic selection: a new and efficient alternative to genomic selection. In: Ahmadi N, Bartholomé J (eds) Complex trait prediction. Springer, New York, pp 397–420
Robert P, Le Gouis J, Rincent R (2020) Combining crop growth modeling with trait-assisted prediction improved the prediction of genotype by environment interactions. Front Plant Sci 11:827. https://doi.org/10.3389/fpls.2020.00827
Schrag TA, Westhues M, Schipprack W et al (2018) Beyond genomic prediction: combining different types of omics data can improve prediction of hybrid performance in maize. Genetics 208:1373–1385. https://doi.org/10.1534/genetics.117.300374
Wang S, Wei J, Li R et al (2019) Identification of optimal prediction models using multi-omic data for selecting hybrid rice. Heredity 123:395–406. https://doi.org/10.1038/s41437-019-0210-6
Westhues M, Schrag TA, Heuer C et al (2017) Omics-based hybrid prediction in maize. Theor Appl Genet 130:1927–1939. https://doi.org/10.1007/s00122-017-2934-0
Whittaker JC, Thompson R, Denham MC (2000) Marker-assisted selection using ridge regression. Genet Res 75:249–252. https://doi.org/10.1017/S0016672399004462
Xu S, Xu Y, Gong L, Zhang Q (2016) Metabolomic prediction of yield in hybrid rice. Plant J 88:219–227. https://doi.org/10.1111/tpj.13242
Yamada Y, Itoh Y, Sugimoto I (1988) Parametric relationships between genotype x environment interaction and genetic correlation when two environments are involved. Theor Appl Genet 76:850–854. https://doi.org/10.1007/BF00273671
Zenke-Philippi C, Frisch M, Thiemann A et al (2017) Transcriptome-based prediction of hybrid performance with unbalanced data from a maize breeding programme. Plant Breed 136:331–337. https://doi.org/10.1111/pbr.12482
Zhu X, Leiser WL, Hahn V, Würschum T (2021a) Phenomic selection is competitive with genomic selection for breeding of complex traits. Plant Phenome J. https://doi.org/10.1002/ppj2.20027
Zhu X, Maurer HP, Jenz M et al (2021b) The performance of phenomic selection depends on the genetic architecture of the target trait. Theor Appl Genet. https://doi.org/10.1007/s00122-021-03997-7
Acknowledgements
The authors thank INRAE experimental units (UE PHACC Clermont-Ferrand, UE GCIE Estrées-Mons, IE UMR GQE Le Moulon, UE Domaine de la Motte Rennes), breeders from Agri-Obtentions and Florimond Desprez. The authors are grateful to Agri-Obtentions, Florimond-Desprez and the Association Nationale de la Recherche et de la Technologie (ANRT, Grant Number 2019/0060) which supported this PhD work. The authors also thank Rachel Carol (Bioscience Editing, France) for proofreading. The authors thank the editor and two anonymous reviewers for their fruitful comments.
Funding
This work was funded by Agri-Obtentions, Florimond-Desprez, and the Association Nationale de la Recherche et de la Technologie (ANRT, Grant Number 2019/0060).
Author information
Authors and Affiliations
Contributions
JA, FXO, BR and EH: designed the field trials and collected the phenotypic data from Agri-Obtentions and INRAE. EGD: provided the phenotypic data and genotyping data from the Florimond Desprez company. SB: provided the genotyping data from Agri-Obtentions and INRAE and participated in discussions of this study. AC and TMH: developed the method of multivariate weighted ridge regression prediction of NIR spectra. RR: initiated the project and with JLG: supervised the study and helped improving the manuscript. PR: analysed the data and wrote the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Co-author Ellen Goudemand was employed by Florimond-Desprez, and co-author Jérôme Auzanneau was employed by Agri-Obtentions. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Communicated by Daniela Bustos-Korts.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Robert, P., Goudemand, E., Auzanneau, J. et al. Phenomic selection in wheat breeding: prediction of the genotype-by-environment interaction in multi-environment breeding trials. Theor Appl Genet 135, 3337–3356 (2022). https://doi.org/10.1007/s00122-022-04170-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04170-4