Abstract
Domesticated materials with well-known wild relatives provide an experimental system to reveal how human selection during cultivation affects genetic composition and adaptation to novel environments. In this paper, our goal was to elucidate how two geographically distinct domestication events modified the structure and level of genetic diversity in common bean. Specifically, we analyzed the genome-wide genetic composition at 26, mostly unlinked microsatellite loci in 349 accessions of wild and domesticated common bean from the Andean and Mesoamerican gene pools. Using a model-based approach, implemented in the software STRUCTURE, we identified nine wild or domesticated populations in common bean, including four of Andean and four of Mesoamerican origins. The ninth population was the putative wild ancestor of the species, which was classified as a Mesoamerican population. A neighbor-joining analysis and a principal coordinate analysis confirmed genetic relationships among accessions and populations observed with the STRUCTURE analysis. Geographic and genetic distances in wild populations were congruent with the exception of a few putative hybrids identified in this study, suggesting a predominant effect of isolation by distance. Domesticated common bean populations possessed lower genetic diversity, higher F ST, and generally higher linkage disequilibrium (LD) than wild populations in both gene pools; their geographic distributions were less correlated with genetic distance, probably reflecting seed-based gene flow after domestication. The LD was reduced when analyzed in separate Andean and Mesoamerican germplasm samples. The Andean domesticated race Nueva Granada had the highest F ST value and widest geographic distribution compared to other domesticated races, suggesting a very recent origin or a selection event, presumably associated with a determinate growth habit, which predominates in this race.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Phaseolus, and more specifically its economically most important species, the common bean or Phaseolus vulgaris L. (2n = 2x = 22), provides interesting features to study the process of plant domestication. Of the 70-odd species that have been recognized in the genus (Freytag and Debouck 2002), 5 have been domesticated and a few additional species show signs of incipient domestication (Delgado-Salinas et al. 2006). Domestication in common bean took place in two, already diverged ancestral gene pools distributed from northern Mexico to Colombia (Mesoamerican gene pool), on the one hand, and from southern Peru to northwestern Argentina (Andean gene pool), on the other (Gepts et al. 1986; Koenig and Gepts 1989; Khairallah et al. 1990, 1992; Koinange and Gepts 1992; Freyre et al. 1996). The two domestications led to two distinct domesticated gene pools (Singh et al. 1991b, c; Becerra Velásquez and Gepts 1994), in part because they arose from two already diverged gene pools just mentioned but also because of further selection under domestication. One consequence of this selection was the appearance of ecogeographic races in each of the two domesticated gene pools (Singh et al. 1991a; Beebe et al. 2000; Díaz and Blair 2006). Partial reproductive isolation has been identified between them, both in wild (Koinange and Gepts 1992) and domesticated populations (Gepts and Bliss 1985), suggesting that P. vulgaris may be in the process of incipient speciation.
The existence of the Andean and Mesoamerican gene pools in common bean and the multiple domestications associated with them is a unique situation among crops, rice being an exception (Vitte et al. 2004; Londo et al. 2006). The existence of these two gene pools raises a number of questions such as the origin and relationships between these two gene pools, the qualitative and quantitative differences in genetic diversity between them, the respective levels of linkage disequilibrium, and the extent to which different loci have been the subject of selection during and after the two major domestications in the species. The first question has been answered with the discovery in the 1980s of a missing link, namely wild P. vulgaris populations in Ecuador and northern Peru (Debouck et al. 1993). Based on a DNA sequence analysis of the genes for phaseolin seed protein, this segment of bean germplasm is actually the putative ancestor of the species (Kami et al. 1995). This segment also shows chloroplast DNA (cpDNA) haplotypes that closely resemble the putative ancestral cpDNA haplotype of the species (Chacón et al. 2007). From the core area on the western slope of the Andes in Ecuador and northern Peru, wild beans were dispersed northwards (to Colombia, Central America, and Mexico) and southwards (southern Peru, Bolivia, and Argentina) resulting in the Mesoamerican and Andean gene pools, respectively. The alpha-amylase inhibitor (Gepts et al. 1999) and internal transcribed spacer (Chacón et al. 2005) sequence data independently suggest that the split between Andean and Mesoamerican gene pools took place some 0.5 million years ago.
In the research reported here, we broadened the scope of earlier research on the organization of genetic diversity in common bean using microsatellite markers by examining a larger plant sample (n = 349), which included both wild and domesticated accessions from the Andean and Mesoamerican gene pools. Microsatellite markers are more polymorphic (Blair et al. 2006) than markers used earlier to characterize genetic diversity such as phaseolin seed protein (Gepts et al. 1986), allozymes (Koenig and Gepts 1989; Singh et al. 1991c), RFLP (Becerra Velásquez and Gepts 1994), and RAPD (Freyre et al. 1996). They are also more widely distributed in the bean genome (Freyre et al. 1998; Blair et al. 2003). In common bean, around 400 microsatellite markers have been developed and mapped (Yu et al. 2000; Gaitán-Solís et al. 2002; Blair et al. 2003; Masi et al. 2003; Yaish and Pérez de la Vega 2003; Guerra-Sanz 2004; Caixeta et al. 2005; Buso et al. 2006). However, population studies with microsatellites in common bean so far have been performed only in a small number of landraces or breeding lines or they have focused on certain geographic regions (Métais et al. 2002; Blair et al. 2006; Díaz and Blair 2006). Thus, an analysis of population structure among wild and domesticated accessions from Andean and Mesoamerican gene pool using microsatellites could yield significant additional insights into the organization of genetic diversity of common bean.
Specifically, we sought to determine how the two domestication processes in common bean had affected genetic diversity and differentiation in the two major gene pools (Andean vs. Mesoamerican), in their respective wild and domesticated components, and among the different domesticated ecogeographic races. We also sought to determine the level of multilocus associations (Hedrick et al. 1978) across and within gene pools and races as a prelude to future linkage disequilibrium (LD; Gupta et al. 2005) and association mapping studies (Zhu et al. 2008).
Materials and methods
Plant materials
Three hundred forty-nine wild, landraces and commercial varieties or advanced germplasm accessions from Latin American, Europe, USA, Africa, and Asia from the Phaseolus World Collection at CIAT, Cali, Colombia or from the Phaseolus collection of the USDA National Plant Germplasm System at Pullman, WA, USA, were analyzed. These samples included 100 wild and 249 domesticated accessions (supplemental Table S1). More detailed information for each accession is included in supplemental Table S1 (accession number, common name, seed weight and color, growth habit, country origin, and coordinates, with assigned gene pool and posterior membership coefficients as determined with STRUCTURE, Pritchard et al. 2000).
Genomic DNA extraction and genotyping microsatellite
Genomic DNA was extracted from young leaves of greenhouse-grown plants using the CTAB method (Doyle and Doyle 1987). Twenty-six microsatellite markers from all 11 linkage groups were selected based on their dispersed map location (Yu et al. 2000; Blair et al. 2003; Pedrosa-Harand et al. 2008). With the exception of marker pairs BM146-BM157 (linkage group 1) and BMd142-BM212 (linkage group 10), which were each linked at approximately 10 cM, all other pairs were distant by 50 cM or more. Markers originated in equal proportions from genic and non-genic sequences (supplemental Table S2). Forward primers were designed with a 5′-TGTAAACGACGGCCAGTATGC M-13 reverse sequence tail added to the 5′ end of the forward primer. The genetic linkage map location, repeat motif, and primer sequences, can be found in the original publications (Bmd: Blair et al. 2003; Pv: Yu et al. 2000; BM: Gaitán-Solís et al. 2002). Except for SSR markers BM146 and BM157, two independent PCR reactions were performed. For the primary PCR, the pairs of forward and reverse primers were used to amplify microsatellite fragments. Thus, the fragments amplified in the primary PCR included the M13 sequence extension at forward primer site. The secondary PCR reactions were performed with the reverse primer and the M-13 primer labeled with the 6-FAM, NED, PET or VIC fluorescence dyes. For the primary PCR reaction, PCR reaction mixtures contained approximately 30 ng of total genomic DNA, 200 mM of dNTP, 0.2 μM of forward primer and reverse primer, the standard Taq buffer with 1.5 mM MgCl2, and 1 unit of Taq polymerase (New England Biolabs) in a 20 μl total reaction volume. The primary PCR cycle consisted of 2 min at 94°C and 35 cycles of 30 s at 94°C, 1 min at 47°C (BMd45, BMd10, BMd1, Pv-ctt001, BMd53, BMd37, BMd12, BMd25, BMd42 and BMd41), 49°C (Pv-ag003, BM143, BM172, Pv-ag004, BM151, Pv-at007, and BM212), 52°C (GATS91, BM160 and BM210), 55°C (BM188), 57°C (Pv-ag001), or 60°C (BM53 and BMd20) and then 40 s at 72°C followed by a 3 min extension at 72°C. For the secondary PCR reaction, the PCR reaction mixtures contained 1 μl of primary PCR product, 0.2 μM of florescence labeled M13 universal primer and reverse primer, 0.34 μM of forward primer and standard Taq buffer with 1.5 mM MgCl2, and 1 unit of Taq polymerase in a total volume 20 μl reaction. For M13 primer labeling, the choice of 6-FAM, NED, PET or VIC dye was attached to the 5′ end of the 5′-TGTAAAACGACGGCCAGT-3′ M-13 universal primer sequence. The secondary PCR cycle consisted of 2 min at 94°C and 30 cycles of 30 s at 94°C, 45 s at 56°C and 45 s at 72°C followed by 8 cycles of 30 s at 94°C, 45 s at 53°C, and 45 s at 72°C, and then 3 min at 72°C for the final extension. For the BM146 and BM157 amplification, PCR reaction mixtures contained approximately 30 ng of total genomic DNA, 200 mM of dNTP, 0.16 μM of labeled M-13 universal primer and reverse primer, 0.04 μM of reverse primer, standard Taq buffer with 1.5 mM MgCl2, and 1 unit of Taq polymerase in a 20 μl total reaction volume. PCR cycles consisted of 5 min at 94°C and 30 cycles of 30 s at 94°C, 45 s at 56°C and 45 s at 72°C followed by 8 cycles of 30 s at 94°C, 45 s at 53°C and 45 s at 72°C, and a final extension of 3 min at 72°C. The amplified fragments were multiplexed depending on their size variation and analyzed in an ABI 3730 (Applied Biosystems). Genotypes of makers were determined using the GeneMarker program (version1.51; SoftGenetics) (supplemental Table S2).
Analysis of population structure
As a preliminary step, STRUCTURE (Pritchard et al. 2000) was run a single time for each K value ranging from 2 to 20. Each run was performed using the admixture model and 1,000 replicates for burn-in and 3,000 during the analysis. To distinguish between Andean and Mesoamerican accessions, the K = 2 analysis was of particular interest. Five independent runs were performed using the admixture model and 5,000 replicates for burn-in and 50,000 replicates during analysis. The clustering in different runs was almost identical (similarity coefficient 0.9969). Among the five runs, the run with the lowest likelihood value was selected and the accessions with more than 50% posterior assignment probability for the Mesoamerican cluster were assigned to the Mesoamerican gene pool (and vice versa for the Andean gene pool) (supplemental Table S1). Low values of posterior assignment probabilities (e.g., between 50 and 80%) may actually indicate hybrids rather than “pure” accessions; however, such accessions are also of interest to understand the origin of the bean gene pool and in breeding. Therefore, we included such accessions in the K = 2 analysis.
Subsequently, 20 simulations per K value were then performed from K = 6 to 12 using 5,000 replicates for burn-in and 50,000 replicates during the analysis. The Δ statistical test using the Structure-sum program showed that K = 9 was optimal in this analysis (Rosenberg et al. 2002; Evanno et al. 2005; Ehrich 2006). At K = 9, the membership coefficient from the run with the lowest likelihood value (−17458.8) was used to assign each accession to the K = 1 to 9 populations for each accession based on the highest membership coefficient (supplemental Table S1). Accessions with a membership coefficient less than 0.8 or 0.9 were identified as putative hybrids. A graphical bar plot of membership coefficients was generated using the Distruct program (Fig. 1; Rosenberg 2004). STRUCTURE was also used to calculate F ST coefficients among the nine populations that were eventually selected.
Hierarchical organization of genetic relatedness of 349 common bean accessions based on 26 microsatellite markers and analyzed by the STRUCTURE program as described in “Materials and methods” for K = 2 to 9. Bar graphs were developed with the program DISTRUCT
Analysis of genetic diversity and geographic distribution
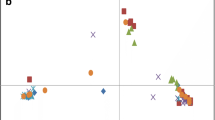
The average number of alleles and gene diversity, heterozygosity, and polymorphism information content (PIC) were calculated for each microsatellite locus using Powermarker version 3.25 (Liu and Muse 2005). Genetic distances among accessions were calculated using the C.S. Chord distance (Cavalli-Sforza and Edwards 1967); a neighbor-joining (NJ) tree was constructed with Powermaker (Fig. 2). The genetic relationship among entire accessions as well as among wild accessions was analyzed by principal coordinate analysis (PCoA) using the GenAlEx 6 program (Fig. 3: see supplemental Table S3 for coordinates; Peakall and Smouse 2006). The geographic distribution of wild accessions was visualized with the DIVA-GIS program (Fig. 4; Hijmans et al. 2001).
Neighbor-joining tree of microsatellite diversity based on the C. S. Chord distance implemented in the Powermarker program. Each branch is color-coded according to membership into the K = 9 groups identified by STRUCTURE (same colors as in Fig. 1). Branches ending with black dots represent domesticated accessions, while those without dots are wild accessions
Geographical and genetic distributions of wild common bean accessions. The lower left plot is the result of a principal coordinate analysis involving wild accessions only (for which precise coordinates are available). The lines link positions of accessions in the PCA graph and their geographic origin on the map. The colors indicate population membership identified using STRUCTURE (same colors as in Fig. 1)
Multilocus associations in common bean
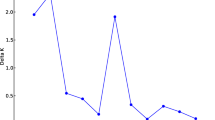
To characterize the frequency of significant multilocus associations in common bean, the microsatellite data were transformed to haplotype data after heterozygous genotypes were treated as missing data. Haplotype frequencies were estimated from 25 microsatellite genotype data using an expectation-maximization (EM) algorithm in ARLEQUIN 3.11 (Excoffier et al. 2005). The EM algorithm estimated haplotype frequencies by a 1,000 permutation procedure. The marker BM157 was removed because of a high frequency of missing data (9%). Forty-three accessions that had any missing values for the remaining 25 markers were also excluded for this analysis. The pairwise LD among microsatellite marker pairs was tested using a likelihood-ratio test between the likelihood of the data assuming linkage equilibrium and the likelihood of the data assuming linkage disequilibrium obtained by estimated haplotypes frequencies (ARLEQUIN program: Excoffier et al. 2005; Excoffier and Slatkin 1998). To further evaluate LD, the standardized disequilibrium coefficient D′ and r 2 were calculated for all accessions with 26 markers using the TASSEL program (Bradbury et al. 2007; http://www.maizegenetics.net/tassel). These LD parameters were calculated for the entire sample, the Andean and Mesoamerican gene pools, and for the different K = 9 groups. Because the sample size differed among gene pools and K = 9 groups, the effect of sample size differences on LD was analyzed by calculation of averages for D′, r 2, and percentage marker pairs in LD from ten random replicated samples generated from the entire sample without replacement and whose size equaled that of the individual groups aforementioned (Fig. 5).
Marker pairs in LD (%) and r 2 and D′ values. Black circle average of each value calculated from ten random samples. Black diamond observed LD values in the entire sample (n = 349); Red open diamond observed LD values in the Mesoamerican gene pool (n = 155); Blue open diamond observed LD values from Andean gene pool (n = 194); Filled diamonds observed LD from K = 9 populations and same colors as in Fig. 1
Results
Microsatellite diversity in common bean
In this study, 26 microsatellite markers distributed over all 11 genetic linkage groups in common bean were genotyped in 349 wild and domesticated accessions. The overall mean genetic diversity in common bean was 0.66 and the average number of alleles per microsatellite locus was 16, ranging from four (BMd45 and BMd1) to 56 alleles (BM53). The PIC values ranged from 0.09 to 0.91 with an average of 0.62 (Table 1). Overall, heterozygosity was below 1–3%, consistent with the predominantly self-pollinated nature of the species (Table 2). The genetic diversity in Mesoamerica was slightly higher than in the Andean group (0.60 and 0.52, respectively; Table 2). With the exception of the Ancestral Peruvian and Ecuadoran wild group (K1), the combined wild groups from both gene pools (K3, K5, and K7) had higher genetic diversity and higher heterozygosity than the combined domesticated groups (K6, K9, K2, K4 and K8). Within both Andean and Mesoamerican gene pools, domestication induced a reduction in genetic diversity of about 10%, whether measured by gene diversity or PIC values (Table 2). Although race Nueva Granada had the largest sample size (94) in this study, its genetic diversity (0.41) was the lowest among the nine groups.
Population structure in common bean
The population subdivision (as determined by STRUCTURE) (Fig. 1), the NJ tree based on genetic distance (Fig. 2), and the PCoA (Fig. 3) showed significant Andean–Mesoamerican gene pool divergence as well as racial differentiation within gene pools. The identification of gene pool of origin (Andean vs. Mesoamerican) for each accession was accomplished as described in “Materials and methods” for K = 2. At K = 2, 155 and 194 accessions fell into the Mesoamerican and Andean groups, respectively, based on posterior assignment probabilities of P > 0.50. This split was generally maintained from K = 2 to 9, with the exception of K = 3 and 6 (Fig. 1). For K = 3, the group of wild, presumably ancestral beans from Ecuador and northern Peru showed a mixed membership between the Mesoamerican (as defined in K = 2) and Andean gene pool (the latter including wild Andean types and domesticated race Peru). For K = 6, the wild, presumably ancestral group clustered with the Andean wild beans. The same group of wild, presumably ancestral grouped with other Mesoamerican accessions at all other K levels. Such membership switching of the presumably ancestral group between the Andean and Mesoamerican gene pools has been observed for allozyme and RAPD data as well (Koenig and Gepts 1989; Freyre et al. 1996).
For K = 9, the groups were identified as Ancestral Northern Peruvian and Ecuadoran wild (K1), Mesoamerican and Colombian wild (K3), Mexican wild (K5), Race Mesoamerica (K6), Races Jalisco and Durango (K9), Andean wild (K7), Race Peru (K2), Race Chile (K4) and Race Nueva Granada (K8) (Fig. 1). The first five groups belong to the Mesoamerican gene pool and the last four groups to the Andean gene pool. This population structure is similar to that encountered in previous population studies in common bean (Gepts et al. 1986; Koenig and Gepts 1989; Singh et al. 1991a; Díaz and Blair 2006). On average, F ST values for wild populations (K1, K3, K5, and K7) were lower (0.22) compared to those of domesticated races (K2, K4, K6, K8, and K9: 0.28) (Table 3). Furthermore, the F ST value for race Nueva Granada was higher than those observed for all other races, whether Andean or Mesoamerican (Table 3).
This study also allowed us to quantify population admixture for each accession (Fig. 1; supplemental Table S1). The Mesoamerican gene pool had a higher proportion of non-hybrid accessions than the Andean gene pool (75 and 64% at the 0.8 cutoff, respectively; Table 4). The proportion of non-hybrid accessions in each K group ranged from 48% (race Chile) to 89% (Mesoamerican and Colombian wild) at the 0.8 cutoff value. The majority of hybrid accessions had an ancestry involving the domesticated groups in the Mesoamerican gene pool (races Jalisco and Durango; race Mesoamerica). In addition, there were admixed accessions in the Andean group involving (1) races Chile and Nueva Granada, and (2) Andean wild types and the domesticated race Peru. Using a cutoff of 0.9 revealed comparable trends.
A similar population structure was uncovered with the NJ tree, in particular the subdivision into Andean and Mesoamerican gene pools, the membership of the ancestral wild group (from Ecuador and northern Peru) in the Mesoamerican gene pool, and the close relationship between races Jalisco and Durango (Fig. 2). Furthermore, Andean and Mesoamerican populations were also well separated according to principal coordinate 1 (53%) (Fig. 3). The presumed Ancestral wild population from northern Peru and Ecuador was positioned between the Andean and Mesoamerican gene pools, but skewed towards the Mesoamerican. While Mesoamerican wild and domesticated populations were well separated on principal coordinate 2 (13%), Andean groups were not well resolved.
Genetic relationship of wild and domesticated accessions according to their geographic distribution
The genetic relationship of wild accessions reflected their geographic distribution with exceptions that had been identified previously as hybrids mainly between local wild types and domesticated accessions from another gene pool or race (Fig. 4). All domesticated groups had a wide geographical distribution with close genetic relationships within groups. For example, accessions of race Mesoamerica (Group K6) were closely related genetically (Fig. 2), but they had a broad geographical distribution that included Bolivia, Brazil, Colombia, Costa Rica, Ecuador, Guatemala, Honduras, Mexico, Nicaragua, Venezuela, and the United States (supplemental Table S1). Accessions classified in the Andean gene pool but originating outside the Americas belonged to race Chile (K4; 9 accessions) and race Nueva Granada (K8; 26 accessions). The absence of Race Peru outside the Americas was noted before (Gepts and Bliss 1988; Zeven et al. 1999).
Multilocus associations in common bean
A very high proportion (95%) of marker pairs among the 26 microsatellites showed a significant LD when considering the entire plant sample of 349 accessions. Marker pairs in LD included both markers in the same or different linkage groups (supplemental Table S4). Calculating LD in “hybrid” versus “non-hybrid” accessions (between the Andean and Mesoamerican gene pools, as determined posterior membership probabilities thresholds of 0.80 or 0.90 in a K = 2 STRUCTURE analysis) showed a limited reduction in LD (supplemental Table S4). To further test the effect of population structure on genome-wide LD, the proportion of pairs in LD and the extent of LD measured by r 2 and D′ were calculated in both gene pools and the nine groups identified in this study. An analysis of LD in separate Andean and Mesoamerican samples (as defined by STRUCTURE) lowered the number of locus pairs in LD from 95 to 68 and 75%, respectively. The LD in the nine groups identified by STRUCTURE was reduced further to 30–40% depending on the group (supplemental Table S4). When measured by r 2 or D′, LD was reduced in “hybrid” accessions (80 or 90% thresholds) compared to “non-hybrid” accessions, and in the Andean or Mesoamerican groups compared to the entire sample (supplemental Table S4). Further subdivision of the Andean or Mesoamerican groups into constituent K groups as defined by STRUCTURE, however, increased values for r 2 and D′, in contrast with the observation for the proportion of pairs in LD.
This apparent contradiction between percentage of locus pairs in LD, on the one hand, and r 2 and D′ could be due to the effect of sample size, to which both r 2 and D′ are sensitive. To examine the possible role of sample size in affecting LD measures, a resampling experiment was conducted. Averages for the proportion of pairs in LD, r 2, and D′ were calculated from 10 independent samples of the same size as each of the Andean, Mesoamerican and K groups defined by STRUCTURE. Each resampling was obtained by sampling the entire plant panel without replacement. The results show that subdivisions of the entire sample used in this study lead to underestimation of LD, whether measured by the percentage of markers pairs in LD, r 2, or D′ (Fig. 5). More specifically, LD (calculated over the entire sample: black diamond) shows a very high proportion (>95%) of marker pairs in non-random association as measured by a likelihood ratio test. Modeling studies with resampling of smaller samples (black-filled circles) with sizes corresponding to those of subdivisions (gene pools or STRUCTURE groups) lead, surprisingly, to smaller proportions of marker pairs in LD, whereas for r 2 and D′ smaller samples sizes lead to the expected increase in LD. In all three graphs, the relationship between sample size and LD is asymptotic. Visual inspection of the graphs to compare LD in the actual total sample (black diamond) and simulated samples (black-filled circles) suggest that a sample of 150–200 is about the minimum size beyond which measures of LD appear to be minimally affected by sample size. In our study, this corresponds to the entire sample (n = 349) and the Andean (n = 194) and Mesoamerican (n = 155) groups. Further subdivisions based on the STRUCTURE groups become too small (n = 9 to 94) to accurately measure LD.
Nevertheless, a comparison of LD in observed samples (colored shapes) and simulated samples of the same size (black-filled circles) suggest that the high levels of LD observed in the entire sample studied here is due to the divergence between the Andean and Mesoamerican gene pools. Subdivision of the entire sample into subsamples that contain entries belonging only to the Andean (open blue diamond) or Mesoamerican gene pool (open red diamond) lowers LD significantly as observed earlier.
Discussion
Current knowledge of population structure and domestication origin of common bean is based on studies that relied on several types of molecular markers (seed proteins: Gepts et al. 1986, Gepts and Bliss 1986; allozymes: Koenig and Gepts 1989, Singh et al. 1991c; RFLPs: Becerra-Velásquez and Gepts1994; RAPDs: Freyre et al. 1996; AFLPs: Tohme et al. 1996; Papa and Gepts 2003; SSRs: Blair et al. 2006; Díaz and Blair 2006) and morphological characteristics (Singh et al. 1991a, b; Gepts 1998). However, in many of these studies, the low level of polymorphism of the markers and the reduced number of markers (Gepts et al. 1986; Becerra Velásquez and Gepts 1994; McClean et al. 2004) precluded a more detailed quantification of the population structure and genetic relationships within the common bean germplasm. For example, electrophoretic variation for phaseolin, the major seed storage protein of common bean, has been instrumental in identifying the geographic pattern of multiple domestications of common bean (Gepts 1988). Nevertheless, phaseolin is coded by a single, albeit complex, locus and its relative lack of polymorphism in the domesticated gene pool prevented the detection of more subtle genetic differences between closely related landraces or cultivars. Furthermore, the phaseolin locus or a locus close linked to it, has since been implicated in the control of seed weight (Johnson et al. 1996; Koinange et al. 1996) and might, thus, be affected by selection for seed size during domestication (Paredes and Gepts 1995). Presumably neutral markers such as microsatellites would therefore be more desirable to assess the genetic structure of the common bean gene pool.
The population structure identified in this study is generally consistent with the current hierarchical scheme of gene pools and ecogeographic racial structure within gene pools (Gepts 1998). First, the differentiation into Andean and Mesoamerican gene pools is well supported in this analysis. In the NJ tree and the PCoA analysis, the Mesoamerican and Andean gene pools are divided into two different clusters (Fig. 1). A stepwise increase in the K number in the STRUCTURE analysis generally leads to subdivisions within the two major gene pools but not to groups of accessions from both gene pools (Fig. 1). The split between the two major geographic gene pools has now been documented repeatedly based on both phenotypic and molecular information and suggests that P. vulgaris may be undergoing incipient speciation. The existence of partial reproductive isolation, including hybrid weakness in the F1 (Shii et al. 1980; Gepts and Bliss 1985; Koinange and Gepts 1992) and later generations (Singh and Molina 1996), further confirms this hypothesis.
Second, the five domesticated groups identified by STRUCTURE generally corresponded to the racial structure of common bean identified by Singh et al. (1991a) except that races Jalisco and Durango constituted a single group in this study. Race Jalisco consists mainly of climbing varieties distributed in the subhumid highlands in the states of Jalisco, Guanajuato, Michoacán, Mexico, Puebla, and Oaxaca. Race Durango includes prostrate varieties originating mainly in the semiarid highlands of northern Mexico (Singh et al. 1991a). Although these races can be distinguished by their distribution, plant and seed morphology, and disease resistance (Singh et al. 1991a), they were not well differentiated at the molecular level in this study. Pallottini et al. (2004) and Díaz and Blair (2006) made a similar observation based on AFLP and microsatellite data, respectively. The closeness of the two races may be due to a recent divergence, high gene flow between them, differentiation of the two races limited to a few major genes controlling plant and seed morphology, or a combination of these factors.
Among the nine STRUCTURE groups, four groups consisted predominantly of wild accessions. First, the K1 group consisted of wild accessions from northern Peruvian and Ecuadoran accessions, which had previously been identified as the presumed ancestral population of P. vulgaris based on the presence of I phaseolin genes without tandem direct repeats (Kami and Gepts 1994; Kami et al. 1995). Although this group was more closely related to Mesoamerican wild types (Figure 2), it was positioned between Andean and Mesoamerican accessions along coordinate 1 (53%) in PCoA plots (Fig. 3). This population was differentiated from other wild populations on coordinate 2 (13%; Fig. 4) and was composed of only nine accessions from a relatively narrow habitat (Debouck et al. 1993). In addition, this population showed lower gene diversity than other wild populations (Table 4) and a single phaseolin type (Debouck et al. 1993). Thus, this population may be a relic that only represents a fraction of the genetic diversity of the ancestral population. Alternatively, the reduced genetic diversity may also reflect the narrow ecological amplitude of this group on the Pacific slope of the Andes (Debouck et al. 1993).
The STRUCTURE analysis detected two Mesoamerican wild populations: Colombian and Mesoamerican wild (K3) and Mexican wild (K5). The Mesoamerican and Colombian wild group (K3) was distributed from Colombia through Guatemala to the central part of Mexico and formed a large cloud of points in the PCA analysis, suggesting a broadly diverse group (Fig. 4). The Mexican wild group (K5) was composed of accessions from Mexico only and, unlike the K3 group, also included accessions from northern Mexico. Compared to the K3 wild group, the K5 wild group was not as dispersed in the PCA plot, suggesting it is a genetically more homogeneous group. In the same PCA plot, Colombian accessions were located in an intermediate position between the Ancestral Peruvian and Ecuadoran population and the Mexican wild populations as observed earlier with RAPD markers (Fig. 4; Freyre et al. 1996).
The 24 Andean wild accessions, which originated in southern Peru, Bolivia, and Argentina, were assigned to one group (K7). However, K7 also includes ten domesticated accessions from Bolivia, Ecuador, Peru and Mexico (supplemental Table S1). Except for three accessions from Peru (G12587, G12588, and G12632), however, these domesticated accessions had low posterior membership probability values (less than 0.8) for K7. Thus, most domesticated accessions in K7 may actually result from hybridization between the wild Andean ancestor and a domesticated descendant. Alternatively, these accessions may represent descendants of the earliest Andean bean domesticates. For example, G12587 and G12588 are nuña or popping beans, which may be among the oldest domesticated beans in the Andes as they can be cooked simply by heating but do not require boiling in ceramics or other types of vessels.
The genetic relatedness among wild accessions correlated well with their geographic distribution except for a few putative hybrids (Fig. 4). When considering simultaneously geographical information, genetic distance, and calculated ancestry using STRUCTURE, the identification of potential hybrids and their ancestry is possible. In Fig. 4, some accessions show a discordance between genetic position based on PCA and geographic location. For example, the Peruvian wild accession G7225 was assigned by STRUCTURE to the K3 group (membership coefficient: 0.545), the group including Colombian and Mesoamerican wild types, in spite of its geographic origin, which suggests membership in the K1 group of southern Andean wild beans (K1 membership coefficient 0.252). Furthermore, G7225 also shows an S-type phaseolin, characteristic of the Mesoamerican gene pool in addition to the T phaseolin, observed in the Andean gene pool (Gepts et al. 1986). Thus, this discordance between geographic and genetic position can be explained by a hybridization event. Wild accession G23580, while originating in Ecuador, grouped with the wild beans from the southern Andes in the PCA. G23580 has both a T (Andean) and an I (ancestral) type of phaseolin (Debouck et al. 1993), suggesting a probable case of outcrossing between a local wild population (I) and an Andean domesticate (T).
The independent domestications in Andean and Mesoamerica region are well-documented (Gepts 1998; Gepts et al. 1986). This study also indicates two different origins for domestication as the Andean and Mesoamerican domesticates are more closely related to wild types of their respective regions (Figs. 1, 2, 3). In the Mesoamerican gene pool, a single cluster groups most of the domesticated type, which confirms previous observations suggesting a single domestication located in the state of Jalisco form this gene pool (Gepts et al. 1986; Papa and Gepts 2003; Kwak et al. 2009). However, it was not possible to reach a conclusion as the domestication pattern within the Andean gene pool in this study as Andean wild or domesticated accessions show less geographic structure than Mesoamerican accessions (Fig. 2). Thus, further study with additional wild accessions or markers should be performed to determine whether Andean beans results from a single or multiple domestications.
The higher F ST values in domesticated types compared to wild types were expected given the relatively higher age of wild populations in relation to their domesticated descendants. A higher age would provide more opportunity for gene exchange among populations. In contrast, the ecogeographic races appeared after domestication because of both drift and selection for adaptation to local conditions, leading to a higher differentiation among landraces. The highest F ST value observed for race Nueva Granada may be due to the predominance of the bush determinate growth habit (type I habit; Singh 1982) or a recent expansion of this group, possibly associated with this growth habit, which is very frequent in race Nueva Granada (Singh et al. 1991a, b, c). In this growth habit, determinacy causes a termination of the modular growth habit of the bean plant (Tanaka and Fujita 1979) and, therefore, leads to earliness, which is often selected by farmers.
The low frequency of non-hybrid accessions in race Chile confirmed the findings of Paredes and Gepts (1995) based on allozyme and phaseolin data that up to 70% of Chilean landraces may have a hybrid origin. The identification of marker alleles as primarily Andean or Mesoamerican in large samples in this study and that of Paredes and Gepts (1995) allowed us to better track potential cases of hybridization, unlike the study of Johns et al. (1997), which used RAPD markers.
The LD has been proposed as a method to identify selection episodes during domestication (Garris et al. 2003; Remington et al. 2001; Thornsberry et al. 2001) and candidate genes (or loci) for agronomically important genes through association mapping (Thornsberry et al. 2001). This genome-wise LD study gives guidelines for further analysis of LD in common bean. First, association mapping should be conducted in separate samples for the Andean and Mesoamerican germplasm. Factoring out the Andean and Mesoamerican structure of the common bean gene pool reduced the percentage of marker pairs in LD and increased the r 2 and D′ values (supplementary Table S4). If population structure is a major variable affecting r 2 and D′ in this study, values of r 2 and D′ should be reduced as the number of groups is increased from K = 2 to 9. Instead, an increase in these values was observed here. Increased r 2 and D′ values in populations of limited size were also observed in durum wheat and barley populations (Maccaferri et al. 2005; Malysheva-Otto et al. 2006). To resolve this apparent contradiction, a comparison was made between r 2 and D′ values obtained in this study with those of randomly sampled populations of the same size. The LD values in Mesoamerican and Andean populations were lower than those of the random sample of the same size, indicating population structure associated with major geographic gene pools has a major effect on LD in common bean (Fig. 5). However, LD in further subdivisions below the gene pool level, especially smaller sample size populations (K1, K3, K5, K6, K9, K7 and K4 in D′ and K1 and K3 in r 2) were similar to LD estimates from random samples. Thus, further subdivisions below the gene pool level may lead to overestimates of D′ and r 2 values because sample sizes of current K groups (wild or domesticated) are too small as shown by the modeling studies performed here.
Second, identification of presumed hybrid accessions between the Andean and Mesoamerican gene pools for the purpose of association mapping does not appear to be a solution gene flow among populations contributes to reducing LD through recombination after hybridization events. The hypothetical population of potential hybrid accessions with membership coefficient values less than 0.9 or 0.8 had fewer locus pairs in LD and lower r 2 and D′ values than putatively non-hybrid accessions (membership coefficients above 0.9) (supplementary Table S4). However, hypothetical hybrid populations still had more than 80% locus pairs in LD. This high frequency of LD is probably caused by the geographic isolation between the two major gene pools, which is further reinforced by partial reproductive isolation. Lastly, this study will provide a Q (genetic background) matrix for further LD studies in common bean (Pritchard et al. 2000; Thornsberry et al. 2001). A more densely populated molecular map will be necessary to conduct more detailed LD mapping and population genomics as proposed by Papa et al. (2005, 2007).
In conclusion, we showed that the ecogeographic races identified with morphological and geographical characteristics are generally congruent with the population structure identified by microsatellite markers. The genetic composition of wild accessions was correlated with their geographic distribution and the ancestry of some wild accessions provided evidence for occasional hybridization with domesticated beans. In addition, we provided evidence of gene flow between races and gene pools through quantification of their ancestry using a model-based approach. Lastly, we showed that association mapping should be performed separately in Andean or Mesoamerican germplasm because a marked reduction in LD is observed by analyzing separate gene pools.
References
Becerra Velásquez VL, Gepts P (1994) RFLP diversity in common bean (Phaseolus vulgaris L.). Genome 37:256–263
Beebe S, Skroch PW, Tohme J, Duque MC, Pedraza F, Nienhuis J (2000) Structure of genetic diversity among common bean landraces of Middle American origin based on correspondence analysis of RAPD. Crop Sci 40:264–273
Blair MW, Giraldo MC, Buendia HF, Tovar E, Duque MC, Beebe SE (2006) Microsatellite marker diversity in common bean (Phaseolus vulgaris L.). Theor Appl Genet 113:100–109
Blair MW, Pedraza F, Buendia HF, Gaitán-Solís E, Beebe SE, Gepts P, Tohme J (2003) Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor Appl Genet 107:1362–1374
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Buso GSC, Amaral ZPS, Brondani RPV, Ferreira ME (2006) Microsatellite markers for the common bean—Phaseolus vulgaris. Mol Ecol Notes 6:252–254
Caixeta ET, Borém A, Kelly JD (2005) Development of microsatellite markers based on BAC common bean clones. Crop Breed Appl Biotech 5:125–133
Cavalli-Sforza LL, Edwards AWF (1967) Phylogenetic analysis: models and estimation procedures. Am J Hum Genet 19:233–257
Chacón SMI, Pickersgill B, Debouck DG (2005) Domestication patterns in common bean (Phaseolus vulgaris L.) and the origin of the Mesoamerican and Andean cultivated races. Theor Appl Genet 110:432–444
Chacón SMI, Pickersgill B, Debouck DG, Arias JS (2007) Phylogeographic analysis of the chloroplast DNA variation in wild common bean (Phaseolus vulgaris L.) in the Americas. Plant Syst Evol 266:175–195
Debouck DG, Toro O, Paredes OM, Johnson WC, Gepts P (1993) Genetic diversity and ecological distribution of Phaseolus vulgaris in northwestern South America. Econ Bot 47:408–423
Delgado-Salinas A, Bibler R, Lavin M (2006) Phylogeny of the genus Phaseolus (Leguminosae): a recent diversification in an ancient landscape. Syst Bot 31:779–791
Díaz LM, Blair MW (2006) Race structure within the Mesoamerican gene pool of common bean (Phaseolus vulgaris L.) as determined by microsatellite markers. Theor Appl Genet 114:143–154
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Ehrich D (2006) AFLPDAT: a collection of R functions for convenient handling of AFLP data. Mol Ecol Notes 6:603–604
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Excoffier L, Slatkin M (1998) Incorporating genotypes of relatives into a test of linkage disequilibrium. Am J Hum Genet 62:171–180
Freyre R, Ríos R, Guzmán L, Debouck D, Gepts P (1996) Ecogeographic distribution of Phaseolus spp. (Fabaceae) in Bolivia. Econ Bot 50:195–215
Freyre R, Skroch P, Geffroy V, Adam-Blondon A-F, Shirmohamadali A, Johnson W, Llaca V, Nodari R, Pereira P, Tsai S-M, Tohme J, Dron M, Nienhuis J, Vallejos C, Gepts P (1998) Towards an integrated linkage map of common bean. 4. Development of a core map and alignment of RFLP maps. Theor Appl Genet 97:847–856
Freytag GF, Debouck DG (2002) Taxonomy, distribution, and ecology of the genus Phaseolus (Leguminosae—Papilionoideae) in North America, Mexico and Central America. Botanical Research Institute of Texas, Forth Worth
Gaitán-Solís E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in common bean (Phaseolus vulgaris): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Sci 42:2128–2136
Garris AJ, McCouch SR, Kresovich S (2003) Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.). Genetics 165:759–769
Gepts P (1988) Phaseolin as an evolutionary marker. In: Gepts P (ed) Genetic resources of Phaseolus beans. Kluwer, Dordrecht, pp 215–241
Gepts P (1998) Origin and evolution of common bean: past events and recent trends. HortScience 33:1124–1130
Gepts P, Bliss FA (1985) F1 hybrid weakness in the common bean: differential geographic origin suggests two gene pools in cultivated bean germplasm. J Hered 76:447–450
Gepts P, Bliss FA (1986) Phaseolin variability among wild and cultivated common beans (Phaseolus vulgaris) from Colombia. Econ Bot 40:469–478
Gepts P, Bliss FA (1988) Dissemination pathways of common bean (Phaseolus vulgaris, Fabaceae) deduced from phaseolin electrophoretic variability. II. Europe and Africa. Econ Bot 42:86–104
Gepts P, Osborn TC, Rashka K, Bliss FA (1986) Phaseolin-protein variability in wild forms and landraces of the common bean (Phaseolus vulgaris): evidence for multiple centers of domestication. Econ Bot 40:451–468
Gepts P, Papa R, Coulibaly S, Gonzalez Mejía A, Pasquet RS (1999) Wild legume diversity: insights from molecular methods.. Ministry of Agriculture, Forestry and Fisheries (MAFF) International Workshop on Genetic Resources, National Institute of Agrobiological Resources, Tsukuba, pp 19–31
Gupta PK, Rustgi S, Kulwal PL (2005) Linkage disequilibrium and association studies in higher plants: Present status and future prospects. Plant Mol Biol 57:461–485
Guerra-Sanz JM (2004) New SSR markers of Phaseolus vulgaris from sequence databases. Plant Breed 123:87–89
Hedrick P, Jain S, Holden L (1978) Multilocus systems in evolution. Evol Biol 11:101–182
Hijmans RJ, Guarino L, Cruz M, Rojas E (2001) Computer tools for spatial analysis of plant genetic resources data: 1. DIVA-GIS. Plant Genet Res Newsl 127:15–19
Johns M, Skroch P, Nienhuis P, Hinrichsen P, Bascur G, Muñoz-Schick C (1997) Gene pool classification of common bean landraces from Chile based on RAPD and morphological data. Crop Sci 37:605–613
Johnson WC, Menéndez C, Nodari RO, Koinange EMK, Magnusson S, Singh SP, Gepts P (1996) Association of a seed weight factor with the phaseolin seed storage protein locus across genotypes, environments, and genomes in Phaseolus–Vigna spp.: Sax (1923) revisited. J Agric Genomics (previously Journal of Quantitative Trait Loci) 2:Article 5, http://www.plantsciences.ucdavis.edu/gepts/Sax.htm
Kami J, Becerra Velásquez B, Debouck DG, Gepts P (1995) Identification of presumed ancestral DNA sequences of phaseolin in Phaseolus vulgaris. Proc Natl Acad Sci USA 92:1101–1104
Kami JA, Gepts P (1994) Phaseolin nucleotide sequence diversity in Phaseolus. I. Intraspecific diversity in Phaseolus vulgaris. Genome 37:751–757
Khairallah MM, Adams MW, Sears BB (1990) Mitochondrial DNA polymorphisms of Malawian bean lines: further evidence for two major gene pools. Theor Appl Genet 80:753–761
Khairallah MM, Sears BB, Adams MW (1992) Mitochondrial restriction fragment polymorphisms in wild Phaseolus vulgaris—insights in the domestication of common bean. Theor Appl Genet 84:915–922
Koenig R, Gepts P (1989) Allozyme diversity in wild Phaseolus vulgaris: further evidence for two major centers of diversity. Theor Appl Genet 78:809–817
Koinange EMK, Gepts P (1992) Hybrid weakness in wild Phaseolus vulgaris L. J Hered 83:135–139
Koinange EMK, Singh SP, Gepts P (1996) Genetic control of the domestication syndrome in common-bean. Crop Sci 36:1037–1045
Kwak M, Kami JA, Gepts P (2009) The putative Mesoamerican domestication center of Phaseolus vulgaris is located in the Lerma-Santiago basin of Mexico. Crop Sci (in press)
Liu KJ, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Londo JP, Chiang YC, Hung KH, Chiang TY, Schaal BA (2006) Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa. Proc Natl Acad Sci USA 103:9578–9583
Maccaferri M, Sanguineti MC, Noli E, Tuberosa R (2005) Population structure and long-range linkage disequilibrium in a durum wheat elite collection. Mol Breed 15:271–289
Malysheva-Otto LV, Ganal MW, Roder MS (2006) Analysis of molecular diversity, population structure and linkage disequilibrium in a worldwide survey of cultivated barley germplasm (Hordeum vulgare L.). BMC Genetics 7, Article 6
Masi P, Spagnoletti Zeuli PL, Donini P (2003) Development and analysis of multiplex microsatellite markers sets in common bean (Phaseolus vulgaris L.). Mol Breed 11:303–313
McClean PE, Lee RK, Miklas PN (2004) Sequence diversity analysis of dihydroflavonol 4-reductase intron 1 in common bean. Genome 47:266–280
Métais I, Hamon B, Jalouzot R, Peltier D (2002) Structure and level of genetic diversity in various bean types evidenced with microsatellite markers isolated from a genomic enriched library. Theor Appl Genet 104:1346–1352
Pallottini L, Garcia E, Kami J, Barcaccia G, Gepts P (2004) The genetic anatomy of a patented yellow bean. Crop Sci 44:968–977
Papa R, Acosta J, Delgado-Salinas A, Gepts P (2005) A genome-wide analysis of differentiation between wild and domesticated Phaseolus vulgaris from Mesoamerica. Theor Appl Genet 111:1147–1158
Papa R, Bellucci E, Rossi M, Leonardi S, Rau D, Gepts P, Nanni L, Attene G (2007) Tagging the signatures of domestication in common bean (Phaseolus vulgaris) by means of pooled DNA samples. Ann Bot 100:1039–1051
Papa R, Gepts P (2003) Asymmetry of gene flow and differential geographical structure of molecular diversity in wild and domesticated common bean (Phaseolus vulgaris L.) from Mesoamerica. Theor Appl Genet 106:239–250
Paredes OM, Gepts P (1995) Extensive introgression of Middle American germplasm into Chilean common bean cultivars. Genet Res Crop Evol 42:29–41
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pedrosa-Harand A, Porch T, Gepts P (2008) Standard nomenclature for common bean chromosomes and linkage groups. Bean Improvement Cooperative, East Lansing, MI, USA. http://www.css.msu.edu/bic/PDF/Standardized%20Genetic%20&%20Physical%20Bean%20Map%202008.pdf Accessed 21 Mar 2008
Pritchard J, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Remington DL, Thornsberry JM, Matsuoka Y, Wilson LM, Whitt SR, Doeblay J, Kresovich S, Goodman MM, Buckler ES (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc Natl Acad Sci USA 98:11479–11484
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Rosenberg NA, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA, Feldman MW (2002) Genetic structure of human populations. Science 298:2381–2385
Shii CT, Mok MC, Temple SR, Mok DWS (1980) Expression of developmental abnormalities in hybrids of Phaseolus vulgaris L. J Hered 71:218–222
Singh SP (1982) A key for identification of different growth habits of Phaseolus vulgaris L. Ann Rep Bean Improv Coop 25:92–95
Singh S, Molina A (1996) Inheritance of crippled trifoliolate leaves occurring in interracial crosses of common bean and its relationship with hybrid dwarfism. J Hered 87:464–469
Singh SP, Gepts P, Debouck DG (1991a) Races of common bean (Phaseolus vulgaris L., Fabaceae). Econ Bot 45:379–396
Singh SP, Gutiérrez JA, Molina A, Urrea C, Gepts P (1991b) Genetic diversity in cultivated common bean: II. Marker-based analysis of morphological and agronomic traits. Crop Sci 31:23–29
Singh SP, Nodari R, Gepts P (1991c) Genetic diversity in cultivated common bean. I. Allozymes. Crop Sci 31:19–23
Tanaka A, Fujita K (1979) Photosynthesis and yield components in relation to grain yield of the field beans. J Fac Agri Hokkaido Univ 59:145–238
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler ES (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28:286–289
Tohme J, Gonzalez DO, Beebe S, Duque MC (1996) AFLP analysis of gene pools of a wild bean core collection. Crop Sci 36:1375–1384
Vitte C, Ishii T, Lamy F, Brar D, Panaud O (2004) Genomic paleontology provides evidence for two distinct origins of Asian rice (Oryza sativa L.). Mol Genet Genom 272:504–511
Yaish MWF, Pérez de la Vega M (2003) Isolation of (GA)(n) microsatellite sequences and description of a predicted MADS-box sequence isolated from common bean (Phaseolus vulgaris L.). Genet Mol Biol 26:337–342
Yu K, Park S, Poysa V, Gepts P (2000) Integration of simple sequence repeat (SSR) markers into a molecular linkage map of common bean (Phaseolus vulgaris L.). J Hered 91:429–434
Zeven AC, Waninge J, van Hintum T, Singh SP (1999) Phenotypic variation in a core collection of common bean (Phaseolus vulgaris L.) in the Netherlands. Euphytica 109:93–106
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgments
This research was supported by the United States Department of Agriculture Cooperative State Research Education and Extension Service—National Research Initiative Plant Genome Program. MK is the recipient of a Department of Plant Sciences graduate student fellowship. We thank D. Debouck and O. Toro, and M. Welsh for providing seed samples from the gene banks at CIAT (Cali, Colombia) and the USDA Western Regional Plant Introduction Station (Pullman, WA, USA), respectively. We thank three anonymous reviewers for their useful suggestions.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Yu.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Kwak, M., Gepts, P. Structure of genetic diversity in the two major gene pools of common bean (Phaseolus vulgaris L., Fabaceae). Theor Appl Genet 118, 979–992 (2009). https://doi.org/10.1007/s00122-008-0955-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-008-0955-4