Abstract.
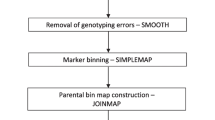
Mapping in forest trees generally relies on outbred pedigrees in which genetic segregation is the result of meiotic recombination from both parents. The currently available mapping packages are not optimal for outcrossed pedigrees as they either cannot order phase-ambiguous data or only use pairwise information when ordering loci within linkage groups. A new package, OUTMAP, has been developed for mapping codominant loci in outcrossed trees. A comparison of maps produced using linkage data from two pedigrees of Acacia mangium Willd demonstrated that the marker orders produced using OUTMAP were consistently of higher likelihood than those produced by JOINMAP. In addition, the maps were produced more efficiently, without the need for recoding data or the detailed investigation of pairwise recombination fractions which was necessary to select the optimal marker order using JOINMAP. Distances between markers often varied from those calculated by JOINMAP, resulting in an increase in the estimated genome length. OUTMAP can be used with all segregation types to determine phase and to calculate the likelihood of alternative marker orders, with a choice of three optimisation methods.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Butcher, .P., Williams, .E., Whitaker, .D. et al. Improving linkage analysis in outcrossed forest trees – an example from Acacia mangium . Theor Appl Genet 104, 1185–1191 (2002). https://doi.org/10.1007/s00122-001-0820-1
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00122-001-0820-1