Abstract
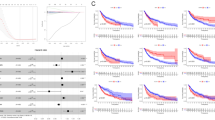
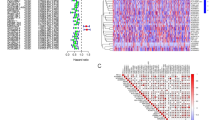
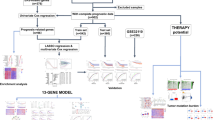
We aimed to develop endoplasmic reticulum (ER) stress-related risk signature to predict the prognosis of melanoma and elucidate the immune characteristics and benefit of immunotherapy in ER-related risk score-defined subgroups of melanoma based on a machine learning algorithm. Based on The Cancer Genome Atlas (TCGA) melanoma dataset (n = 471) and GTEx database (n = 813), 365 differentially expressed ER-associated genes were selected using the univariate Cox model and LASSO penalty Cox model. Ten genes impacting OS were identified to construct an ER-related signature by using the multivariate Cox regression method and validated with the Gene Expression Omnibus (GEO) dataset. Thereafter, the immune features, CNV, methylation, drug sensitivity, and the clinical benefit of anticancer immune checkpoint inhibitor (ICI) therapy in risk score subgroups, were analyzed. We further validated the gene signature using pan-cancer analysis by comparing it to other tumor types. The ER-related risk score was constructed based on the ARNTL, AGO1, TXN, SORL1, CHD7, EGFR, KIT, HLA-DRB1 KCNA2, and EDNRB genes. The high ER stress-related risk score group patients had a poorer overall survival (OS) than the low-risk score group patients, consistent with the results in the GEO cohort. The combined results suggested that a high ER stress-related risk score was associated with cell adhesion, gamma phagocytosis, cation transport, cell surface cell adhesion, KRAS signalling, CD4 T cells, M1 macrophages, naive B cells, natural killer (NK) cells, and eosinophils and less benefitted from ICI therapy. Based on the expression patterns of ER stress-related genes, we created an appropriate predictive model, which can also help distinguish the immune characteristics, CNV, methylation, and the clinical benefit of ICI therapy.
Key messages
-
Melanoma is the cutaneous tumor with a high degree of malignancy, the highest fatality rate, and extremely poor prognosis.
-
Model usefulness should be considered when using models that contained more features.
-
We constructed the Endoplasmic Reticulum stress-associated signature using TCGA and GEO database based on machine learning algorithm.
-
ER stress-associated signature has excellent ability for predicting prognosis for melanoma.











Similar content being viewed by others
Availability of data and materials
The data that support the findings of this study are available in The Cancer Genome Atlas (TCGA) datasets at https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga.
References
Song WM, Agrawal P, Von Itter R, Fontanals-Cirera B, Wang M, Zhou X et al (2021) Network models of primary melanoma microenvironments identify key melanoma regulators underlying prognosis. Nat Commun 12(1):1214. https://doi.org/10.1038/s41467-021-21457-0
Zhang Y, Hou J, Shi S, Du J, Liu Y, Huang P et al (2021) CSN6 promotes melanoma proliferation and metastasis by controlling the UBR5-mediated ubiquitination and degradation of CDK9. Cell Death Dis 12(1):118. https://doi.org/10.1038/s41419-021-03398-0
Wang N, Shi J, Wu C, Chu W, Tao W, Li W et al (2020) Design of DOX-GNRs-PNIPAM@PEG-PLA micelle with temperature and light dual-function for potent melanoma therapy. Front Chem 8:599740. https://doi.org/10.3389/fchem.2020.599740
Kim TW, Hong DW, Hong SH (2020) CB13, a novel PPARgamma ligand, overcomes radio-resistance via ROS generation and ER stress in human non-small cell lung cancer. Cell Death Dis 11(10):848. https://doi.org/10.1038/s41419-020-03065-w
Oakes SA, Papa FR (2015) The role of endoplasmic reticulum stress in human pathology. Annu Rev Pathol 10:173–194. https://doi.org/10.1146/annurev-pathol-012513-104649
Chen X, Cubillos-Ruiz JR (2021) Endoplasmic reticulum stress signals in the tumour and its microenvironment. Nat Rev Cancer 21(2):71–88. https://doi.org/10.1038/s41568-020-00312-2
Oakes SA (2020) Endoplasmic reticulum stress signaling in cancer cells. Am J Pathol 190(5):934–946. https://doi.org/10.1016/j.ajpath.2020.01.010
Szasz I, Koroknai V, Patel V, Hajdu T, Kiss T, Adany R et al (2021) Cell proliferation is strongly associated with the treatment conditions of an ER stress inducer new anti-melanoma drug in melanoma cell lines. Biomedicines 9(2). https://doi.org/10.3390/biomedicines9020096
Kumari N, Reabroi S, North BJ (2021) Unraveling the molecular nexus between GPCRs, ERS, and EMT. Mediators Inflamm 2021:6655417. https://doi.org/10.1155/2021/6655417
Raines LN, Zhao H, Wang Y, Chen HY, Gallart-Ayala H, Hsueh PC et al (2022) PERK is a critical metabolic hub for immunosuppressive function in macrophages. Nat Immunol 23(3):431–445. https://doi.org/10.1038/s41590-022-01145-x
Cubillos-Ruiz JR, Bettigole SE, Glimcher LH (2017) Tumorigenic and immunosuppressive effects of endoplasmic reticulum stress in cancer. Cell 168(4):692–706. https://doi.org/10.1016/j.cell.2016.12.004
Zhang Q, Guan G, Cheng P, Cheng W, Yang L, Wu A (2021) Characterization of an endoplasmic reticulum stress-related signature to evaluate immune features and predict prognosis in glioma. J Cell Mol Med 25(8):3870–3884. https://doi.org/10.1111/jcmm.16321
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S et al (2016) Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165(1):35–44. https://doi.org/10.1016/j.cell.2016.02.065
Riaz N, Havel JJ, Makarov V, Desrichard A, Urba WJ, Sims JS et al (2017) Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171(4):934–49e16. https://doi.org/10.1016/j.cell.2017.09.028
Geeleher P, Cox NJ, Huang RS (2014) Clinical drug response can be predicted using baseline gene expression levels and in vitro drug sensitivity in cell lines. Genome Biol 15(3):R47. https://doi.org/10.1186/gb-2014-15-3-r47
Zhou X, Du J, Liu C, Zeng H, Chen Y, Liu L et al (2021) A pan-cancer analysis of CD161, a potential new immune checkpoint. Front Immunol 12:688215. https://doi.org/10.3389/fimmu.2021.688215
Ju M, Bi J, Wei Q, Jiang L, Guan Q, Zhang M et al (2021) Pan-cancer analysis of NLRP3 inflammasome with potential implications in prognosis and immunotherapy in human cancer. Brief Bioinform 22(4). https://doi.org/10.1093/bib/bbaa345
Zhang J, Jiang H, Du K, Xie T, Wang B, Chen C et al (2021) Pan-cancer analysis of genomic and prognostic characteristics associated with coronavirus disease 2019 regulators. Front Med (Lausanne) 8:662460. https://doi.org/10.3389/fmed.2021.662460
Chan TA, Yarchoan M, Jaffee E, Swanton C, Quezada SA, Stenzinger A et al (2019) Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol 30(1):44–56. https://doi.org/10.1093/annonc/mdy495
Palmeri M, Mehnert J, Silk AW, Jabbour SK, Ganesan S, Popli P et al (2022) Real-world application of tumor mutational burden-high (TMB-high) and microsatellite instability (MSI) confirms their utility as immunotherapy biomarkers. ESMO Open 7(1):100336. https://doi.org/10.1016/j.esmoop.2021.100336
Sharma P, Allison JP (2015) The future of immune checkpoint therapy. Science 348(6230):56–61. https://doi.org/10.1126/science.aaa8172
Sunshine J, Taube JM (2015) PD-1/PD-L1 inhibitors. Curr Opin Pharmacol 23:32–38. https://doi.org/10.1016/j.coph.2015.05.011
Stevenson VB, Perry SN, Todd M, Huckle WR, LeRoith T (2021) PD-1, PD-L1, and PD-L2 gene expression and tumor infiltrating lymphocytes in canine melanoma. Vet Pathol 58(4):692–698. https://doi.org/10.1177/03009858211011939
Yang Y, Long X, Li K, Li G, Yu X, Wen P et al (2021) Development and validation of an oxidative stress-associated prognostic risk model for melanoma. Peer J 9:e11258. https://doi.org/10.7717/peerj.11258
Huang R, Mao M, Lu Y, Yu Q, Liao L (2020) A novel immune-related genes prognosis biomarker for melanoma: associated with tumor microenvironment. Aging (Albany NY) 12(8):6966–6980. https://doi.org/10.18632/aging.103054
Marzagalli M, Ebelt ND, Manuel ER (2019) Unraveling the crosstalk between melanoma and immune cells in the tumor microenvironment. Semin Cancer Biol 59:236–250. https://doi.org/10.1016/j.semcancer.2019.08.002
Eddy K, Chen S (2020) Overcoming immune evasion in melanoma. Int J Mol Sci 21(23). https://doi.org/10.3390/ijms21238984
Thrane K, Eriksson H, Maaskola J, Hansson J, Lundeberg J (2018) Spatially resolved transcriptomics enables dissection of genetic heterogeneity in stage III cutaneous malignant melanoma. Cancer Res 78(20):5970–5979. https://doi.org/10.1158/0008-5472.CAN-18-0747
Rather RA, Bhagat M, Singh SK (2020) Oncogenic BRAF, endoplasmic reticulum stress, and autophagy: crosstalk and therapeutic targets in cutaneous melanoma. Mutat Res Rev Mutat Res 785:108321. https://doi.org/10.1016/j.mrrev.2020.108321
Cerezo M, Lehraiki A, Millet A, Rouaud F, Plaisant M, Jaune E et al (2016) Compounds triggering ER stress exert anti-melanoma effects and overcome BRAF inhibitor resistance. Cancer Cell 29(6):805–819. https://doi.org/10.1016/j.ccell.2016.04.013
Urra H, Dufey E, Avril T, Chevet E, Hetz C (2016) Endoplasmic reticulum stress and the hallmarks of cancer. Trends Cancer 2(5):252–262. https://doi.org/10.1016/j.trecan.2016.03.007
Yao X, Tu Y, Xu Y, Guo Y, Yao F, Zhang X (2020) Endoplasmic reticulum stress-induced exosomal miR-27a-3p promotes immune escape in breast cancer via regulating PD-L1 expression in macrophages. J Cell Mol Med 24(17):9560–9573. https://doi.org/10.1111/jcmm.15367
de Assis LVM, Kinker GS, Moraes MN, Markus RP, Fernandes PA, Castrucci AML (2018) Expression of the circadian clock gene BMAL1 positively correlates with antitumor immunity and patient survival in metastatic melanoma. Front Oncol 8:185. https://doi.org/10.3389/fonc.2018.00185
Peng H, Zhang J, Zhang PP, Chen L, Tang LL, Yang XJ et al (2019) ARNTL hypermethylation promotes tumorigenesis and inhibits cisplatin sensitivity by activating CDK5 transcription in nasopharyngeal carcinoma. J Exp Clin Cancer Res 38(1):11. https://doi.org/10.1186/s13046-018-0997-7
Xu GY, Tang XJ (2017) Troxerutin (TXN) potentiated 5-Fluorouracil (5-Fu) treatment of human gastric cancer through suppressing STAT3/NF-kappaB and Bcl-2 signaling pathways. Biomed Pharmacother 92:95–107. https://doi.org/10.1016/j.biopha.2017.04.059
Cai WQ, Zeng LS, Wang LF, Wang YY, Cheng JT, Zhang Y et al (2020) The latest battles between EGFR monoclonal antibodies and resistant tumor cells. Front Oncol 10:1249. https://doi.org/10.3389/fonc.2020.01249
Wei F, Ge Y, Li W, Wang X, Chen B (2020) Role of endothelin receptor type B (EDNRB) in lung adenocarcinoma. Thorac Cancer 11(7):1885–1890. https://doi.org/10.1111/1759-7714.13474
Chu X, Guo X, Jiang Y, Yu H, Liu L, Shan W et al (2017) Genotranscriptomic meta-analysis of the CHD family chromatin remodelers in human cancers - initial evidence of an oncogenic role for CHD7. Mol Oncol 11(10):1348–1360. https://doi.org/10.1002/1878-0261.12104
Machado RAC, Schneider H, DeOcesano-Pereira C, Lichtenstein F, Andrade F, Fujita A et al (2019) CHD7 promotes glioblastoma cell motility and invasiveness through transcriptional modulation of an invasion signature. Sci Rep 9(1):3952. https://doi.org/10.1038/s41598-019-39564-w
Wu G, Fan F, Hu P, Wang C (2020) AGO1 enhances the proliferation and invasion of cholangiocarcinoma via the EMT-associated TGF-beta signaling pathway. Am J Transl Res 12(6):2890–2902
Fang X, Yin Z, Li X, Xia L, Zhou B (2016) Polymorphisms in GEMIN4 and AGO1 genes are associated with the risk of lung cancer: a case-control study in Chinese female non-smokers. Int J Environ Res Public Health 13(10). https://doi.org/10.3390/ijerph13100939
Kanematsu S, Tanimoto K, Suzuki Y, Sugano S (2014) Screening for possible miRNA-mRNA associations in a colon cancer cell line. Gene 533(2):520–531. https://doi.org/10.1016/j.gene.2013.08.005
Xu Y, Chen D, Shen L, Huang X, Chen Y, Su H (2021) Identification and mechanism of the PD-1/PD-L1 genomic signature SORL1 as protective factor in bladder cancer. Front Genet 12:736158. https://doi.org/10.3389/fgene.2021.736158
Kim JO, Kim KH, Baek EJ, Park B, So MK, Ko BJ et al (2022) A novel anti-c-Kit antibody-drug conjugate to treat wild-type and activating-mutant c-Kit-positive tumors. Mol Oncol 16(6):1290–1308. https://doi.org/10.1002/1878-0261.13084
Chengcheng L, Wenwen Q, Ningyue G, Fangyuan Z, Runtong X, Zhenxiao T et al (2021) Identification of the immune-related genes in tumor microenvironment that associated with the recurrence of head and neck squamous cell carcinoma. Front Cell Dev Biol 9:723721. https://doi.org/10.3389/fcell.2021.723721
Correale P, Saladino RE, Giannarelli D, Giannicola R, Agostino R, Staropoli N et al (2020) Distinctive germline expression of class I human leukocyte antigen (HLA) alleles and DRB1 heterozygosis predict the outcome of patients with non-small cell lung cancer receiving PD-1/PD-L1 immune checkpoint blockade. J Immunother Cancer 8(1). https://doi.org/10.1136/jitc-2020-000733
Ribatti D, Solimando AG, Pezzella F (2021) The anti-VEGF(R) drug discovery legacy: improving attrition rates by breaking the vicious cycle of angiogenesis in cancer. Cancers (Basel) 13(14). https://doi.org/10.3390/cancers13143433
Mahoney FI, Barthel DW (1965) Functional evaluation: the Barthel index. Md State Med J 14:61–65
Walter LC, Brand RJ, Counsell SR, Palmer RM, Landefeld CS, Fortinsky RH et al (2001) Development and validation of a prognostic index for 1-year mortality in older adults after hospitalization. JAMA 285(23):2987–2994. https://doi.org/10.1001/jama.285.23.2987
Lee Y (2000) The predictive value of self assessed general, physical, and mental health on functional decline and mortality in older adults. J Epidemiol Community Health 54(2):123–129. https://doi.org/10.1136/jech.54.2.123
Mossey JM, Shapiro E (1982) Self-rated health: a predictor of mortality among the elderly. Am J Public Health 72(8):800–808. https://doi.org/10.2105/ajph.72.8.800
Vuorisalmi M, Lintonen T, Jylha M (2005) Global self-rated health data from a longitudinal study predicted mortality better than comparative self-rated health in old age. J Clin Epidemiol 58(7):680–687. https://doi.org/10.1016/j.jclinepi.2004.11.025
Benyamini Y, Blumstein T, Lusky A, Modan B (2003) Gender differences in the self-rated health-mortality association: is it poor self-rated health that predicts mortality or excellent self-rated health that predicts survival? Gerontologist 43(3):396–405; discussion 372–5. https://doi.org/10.1093/geront/43.3.396
Cochran WG (1968) The effectiveness of adjustment by subclassification in removing bias in observational studies. Biometrics 24(2):295–313
Austin PC, Mamdani MM, Stukel TA, Anderson GM, Tu JV (2005) The use of the propensity score for estimating treatment effects: administrative versus clinical data. Stat Med 24(10):1563–1578. https://doi.org/10.1002/sim.2053
Sturmer T, Schneeweiss S, Brookhart MA, Rothman KJ, Avorn J, Glynn RJ (2005) Analytic strategies to adjust confounding using exposure propensity scores and disease risk scores: nonsteroidal antiinflammatory drugs and short-term mortality in the elderly. Am J Epidemiol 161(9):891–898. https://doi.org/10.1093/aje/kwi106
Reeves BC, van Binsbergen J, van Weel C (2005) Systematic reviews incorporating evidence from nonrandomized study designs: reasons for caution when estimating health effects. Eur J Clin Nutr 59(Suppl 1):S155–S161. https://doi.org/10.1038/sj.ejcn.1602190
Brookhart MA, Schneeweiss S, Rothman KJ, Glynn RJ, Avorn J, Sturmer T (2006) Variable selection for propensity score models. Am J Epidemiol 163(12):1149–1156. https://doi.org/10.1093/aje/kwj149
Acknowledgements
We also acknowledge TCGA and GEO database for providing their platforms and contributors for uploading their meaningful datasets.
Funding
This work was supported by the Projects of the National Natural Science Foundation of China (grant number 82073019 and 82073018), the Shenzhen Science and Technology Innovation Commission, China (Natural Science Foundation of Shenzhen, grant number JCYJ20210324113001005), Scientific Research Fund of Hunan Provincial Education Department (grant number 20C0402), and Hunan First Normal University (grant number XYS16N03), and Natural Science Foundation of Hunan Province China under Grants 2022JJ30673;.
Author information
Authors and Affiliations
Contributions
Conceptualization: Alphonse Houssou Hounye; investigation: Bingqian Hu; writing—original draft: Alphonse Houssou Hounye and Bingqian Hu; writing—review and editing: Alphonse Houssou Hounye, Bingqian Hu, Zheng Wang, Jiaoju Wang, Cong Cao, Min Qi, Jianglin Zhang, and Muzhou Hou; visualization: Alphonse Houssou Hounye; supervision: Zheng Wang, Min Qi, Jianglin Zhang, and Muzhou Hou; funding acquisition: Min Qi and Jianglin Zhang.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The current study followed data requirements and Clinical Medical Ethics Committee approval, Xiangya Hospital, Central South University with ethic number: 202112271.
Consent for publication
All authors consent to the publication of the article in Journal of Molecular Medicine.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Alphonse Houssou Hounye and Bingqian Hu contributed equally to this work
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hounye, A.H., Hu, B., Wang, Z. et al. Evaluation of drug sensitivity, immunological characteristics, and prognosis in melanoma patients using an endoplasmic reticulum stress-associated signature based on bioinformatics and pan-cancer analysis. J Mol Med 101, 1267–1287 (2023). https://doi.org/10.1007/s00109-023-02365-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-023-02365-w