Abstract
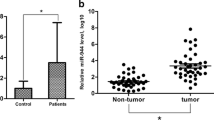
The modulation of mitochondrial functions is important for maintaining cellular homeostasis. Mitochondria essentially depend on the import of RNAs and proteins encoded by the nuclear genome. MicroRNAs encoded in the nucleus can translocate to mitochondria and target the genome, affecting mitochondrial function. Here, we analyzed the role of miR-4485 in the regulation of mitochondrial functions. We showed that miR-4485 translocated to mitochondria where its levels varied in response to different stress conditions. A direct binding of miR-4485 to mitochondrial 16S rRNA was demonstrated. MiR-4485 regulated the processing of pre-rRNA at the 16S rRNA-ND1 junction and the translation of downstream transcripts. MiR-4485 modulated mitochondrial complex I activity, the production of ATP, ROS levels, caspase-3/7 activation, and apoptosis. Transfection of a miR-4485 mimic downregulated the expression of regulatory glycolytic pathway genes and reduced the clonogenic ability of breast cancer cells. Ectopic expression of miR-4485 in MDA-MB-231 breast carcinoma cells decreased the tumorigenicity in a nude mouse xenograft model. Furthermore, levels of both precursor and mature miR-4485 are decreased in tumor tissue of breast cancer patients. We conclude that the mitochondria-targeted miR-4485 may act as a tumor suppressor in breast carcinoma cells by negatively regulating mitochondrial RNA processing and mitochondrial functions.





Similar content being viewed by others
References
Currais A (2015) Ageing and inflammation—a central role for mitochondria in brain health and disease. Ageing Res Rev 21:30–42
Friedman JR, Nunnari J (2014) Mitochondrial form and function. Nature 505:335–343. doi:10.1038/nature12985.
D’Aquila P, Bellizzi D, Passarino G (2015) Mitochondria in health, aging and diseases: the epigenetic perspective. Biogerontology 16:569–585
Iommarini L, Peralta S, Torraco A, Diaz F (2015) Mitochondrial diseases part II: mouse models of OXPHOS deficiencies caused by defects in regulatory factors and other components required for mitochondrial function. Mitochondrion 22:96–118
Pavlova NN, Thompson CB (2016) The emerging hallmarks of cancer metabolism. Cell Metab 23:27–47
Birsoy K, Possemato R, Lorbeer FK, Bayraktar EC, Thiru P, Yucel B, Wang T, Chen WW, Clish CB, Sabatini DM (2014) Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 508:108–112
Xu XD, Shao SX, Jiang HP, Cao YW, Wang YH, Yang XC, Wang YL, Wang XS, Niu HT (2015) Warburg effect or reverse Warburg effect? A review of cancer metabolism. Oncol Res Treat 38:117–122
Sullivan LB, Chandel NS (2014) Mitochondrial reactive oxygen species and cancer. Cancer Metab 2:17
Oronsky BT, Oronsky N, Fanger GR, Parker CW, Caroen SZ, Lybeck M, Scicinski JJ (2014) Follow the ATP: tumor energy production: a perspective. Anti Cancer Agents Med Chem 14:1187–1198
Dell’ Antone P (2012) Energy metabolism in cancer cells: how to explain the Warburg and Crabtree effects? Med Hypotheses 79:388–392
Weinberg SE, Chandel NS (2015) Targeting mitochondria metabolism for cancer therapy. Nat Chem Biol 11:9–15
Phan LM, Yeung SC, Lee M-HH (2014) Cancer metabolic reprogramming: importance, main features, and potentials for precise targeted anti-cancer therapies. Cancer Biol Med 11:1–19
Falk MJ, Shen L, Gonzalez M, Leipzig J, Lott MT, Stassen AP, Diroma MA, Navarro-Gomez D, Yeske P, Bai R et al (2015) Mitochondrial Disease Sequence Data Resource (MSeqDR): a global grass-roots consortium to facilitate deposition, curation, annotation, and integrated analysis of genomic data for the mitochondrial disease clinical and research communities. Mol Genet Metab 114:388–396
Schmidt O, Pfanner N, Meisinger C (2010) Mitochondrial protein import: from proteomics to functional mechanisms. Nat Rev Mol Cell Biol 11:655–667
Croft D, Mundo AF, Haw R, Milacic M, Weiser J, Wu G, Caudy M, Garapati P, Gillespie M, Kamdar MR et al (2014) The Reactome pathway Knowledgebase. Nucleic Acids Res 42:D472–D477
Calvo SE, Mootha VK (2010) The mitochondrial proteome and human disease. Annu Rev Genomics Hum Genet 11:25–44
Sripada L, Tomar D, Singh R (2012) Mitochondria: one of the destinations of miRNAs. Mitochondrion 12:593–599
Zhang X, Zuo X, Yang B, Li Z, Xue Y, Zhou Y, Huang J, Zhao X, Zhou J, Yan Y et al (2014) MicroRNA directly enhances mitochondrial translation during muscle differentiation. Cell 158:607–619
Sripada L, Tomar D, Prajapati P, Singh R, Singh AK, Singh R (2012) Systematic analysis of small RNAs associated with human mitochondria by deep sequencing: detailed analysis of mitochondrial associated miRNA. PLoS One 7:e44873
Galimov ER, Chernyak BV, Sidorenko AS, Tereshkova AV, Chumakov PM (2014) Prooxidant properties of p66shc are mediated by mitochondria in human cells. PLoS One 9:e86521
Li J-HH, Liu S, Zhou H, Qu L-HH, Yang J-HH (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92–D97
Lal A, Thomas MP, Altschuler G, Navarro F, O’Day E, Li XL, Concepcion C, Han YC, Thiery J, Rajani DK et al (2011) Capture of microRNA-bound mRNAs identifies the tumor suppressor miR-34a as a regulator of growth factor signaling. PLoS Genet 7:e1002363
Feng Y, Zhang X, Graves P, Zeng Y (2012) A comprehensive analysis of precursor microRNA cleavage by human Dicer. RNA 18:2083–2092
Vargas JN, Kar AN, Kowalak JA, Gale JR, Aschrafi A, Chen CY, Gioio AE, Kaplan BB (2016) Axonal localization and mitochondrial association of precursor microRNA 338. Cell Mol Life Sci. doi:10.1007/s00018-016-2270-6
Barrey E, Saint-Auret G, Bonnamy B, Damas D, Boyer O, Gidrol X (2011) Pre-microRNA and mature microRNA in human mitochondria. PLoS One 6:e20220
Rusek AM, Abba M, Eljaszewicz A, Moniuszko M, Niklinski J, Allgayer H (2015) MicroRNA modulators of epigenetic regulation, the tumor microenvironment and the immune system in lung cancer. Mol Cancer 14:34
Kadamkode V, Banerjee G (2014) Micro RNA: an epigenetic regulator of type 2 diabetes. Microrna 3:86–97
Hommers LG, Domschke K, Deckert J (2015) Heterogeneity and individuality: microRNAs in mental disorders. J Neural Transm (Vienna) 122:79–97
Mendell JT, Olson EN (2012) MicroRNAs in stress signaling and human disease. Cell 148:1172–1187
Bianchessi V, Badi I, Bertolotti M, Nigro P, D’Alessandra Y, Capogrossi MC, Zanobini M, Pompilio G, Raucci A, Lauri A (2015) The mitochondrial lncRNA ASncmtRNA-2 is induced in aging and replicative senescence in endothelial cells. J Mol Cell Cardiol 81:62–70
King MP, Attardi G (1996) Isolation of human cell lines lacking mitochondrial DNA. Meth Enzymol 264:304–313
Castanotto D, Lingeman R, Riggs AD, Rossi JJ (2009) CRM1 mediates nuclear-cytoplasmic shuttling of mature microRNAs. Proc Natl Acad Sci U S A 106:21655–21659
Rhee WJ, Bao G (2010) Slow non-specific accumulation of 2′-deoxy and 2′-O-methyl oligonucleotide probes at mitochondria in live cells. Nucleic Acids Res 38:e109
Tan AS, Baty JW, Dong L-FF, Bezawork-Geleta A, Endaya B, Goodwin J, Bajzikova M, Kovarova J, Peterka M, Yan B et al (2015) Mitochondrial genome acquisition restores respiratory function and tumorigenic potential of cancer cells without mitochondrial DNA. Cell Metab 21:81–94
Murphy MP (2009) How mitochondria produce reactive oxygen species. Biochem J 417:1–13
Webster KA (2012) Mitochondrial membrane permeabilization and cell death during myocardial infarction: roles of calcium and reactive oxygen species. Futur Cardiol 8:863–884
Ueda S, Masutani H, Nakamura H, Tanaka T, Ueno M, Yodoi J (2002) Redox control of cell death. Antioxid Redox Signal 4:405–414
Richter-Dennerlein R, Oeljeklaus S, Lorenzi I, Ronsör C, Bareth B, Schendzielorz AB, Wang C, Warscheid B, Rehling P, Dennerlein S (2016) Mitochondrial protein synthesis adapts to influx of nuclear-encoded protein. Cell. doi:10.1016/j.cell.2016.09.003
Wolf AR, Mootha VK (2014) Functional genomic analysis of human mitochondrial RNA processing. Cell Rep 7:918–931
Jourdain AA, Boehm E, Maundrell K, Martinou J-CC (2016) Mitochondrial RNA granules: compartmentalizing mitochondrial gene expression. J Cell Biol 212:611–614
Popow J, Alleaume A-MM, Curk T, Schwarzl T, Sauer S, Hentze MW (2015) FASTKD2 is an RNA-binding protein required for mitochondrial RNA processing and translation. RNA 21:1873–1884
Antonicka H, Shoubridge EA (2015) Mitochondrial RNA granules are centers for posttranscriptional RNA processing and ribosome biogenesis. Cell Rep. doi:10.1016/j.celrep.2015.01.030
Stroud DA, Surgenor EE, Formosa LE, Reljic B, Frazier AE, Dibley MG, Osellame LD, Stait T, Beilharz TH, Thorburn DR et al (2016) Accessory subunits are integral for assembly and function of human mitochondrial complex I. Nature 538:123–126
Halestrap AP (1982) The pathway of electron flow through ubiquinol:cytochrome c oxidoreductase in the respiratory chain. Evidence from inhibition studies for a modified “Q cycle”. Biochem J 204:49–59
Cheung EC, Lee P, Ceteci F, Nixon C, Blyth K, Sansom OJ, Vousden KH et al (2016) Opposing effects of TIGAR- and RAC1-derived ROS on Wnt-driven proliferation in the mouse intestine. Genes Dev 30:52–63
Weinberg F, Hamanaka R, Wheaton WW, Weinberg S, Joseph J, Lopez M, Kalyanaraman B, Mutlu GM, Budinger GR, Chandel NS (2010) Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc Natl Acad Sci U S A 107:8788–8793
Chandel NS, Maltepe E, Goldwasser E, Mathieu CE, Simon MC, Schumacker PT (1998) Mitochondrial reactive oxygen species trigger hypoxia-induced transcription. Proc Natl Acad Sci U S A 95:11715–11720
Acknowledgements
This work is being supported by EMR/2014/000219, DST, Government of India to RS. This work is lead obtained from previous DBT grant BT/PR12906/AGR/36/638/2009, DBT, Government of India. A.V.L. and P.M.C. were supported by the grant 14-50-00060 from the Russian Science Foundation. The authors acknowledge the instrumentation facility by DBT MSUB ILSPARE. We thank Prof. Narry Kim, Institute for Basic Science and Seoul National University, Korea for giving us plasmids of pCK-Flag-Dicer as a gift. L.S., A.S., and K.S. received their Senior Research fellowship from University Grants Commission (UGC), Government of India. K.B. received her Senior Research fellowship from the Council of Scientific and Industrial Research (CSIR), Government of India. Dr. Rochika Singh received her fellowship as DST Young Scientist from Department of Science and Technology (DST), Government of India. This work consists of the PhD thesis of L.S.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest related to the study.
Electronic supplementary material
ESM 1
(PDF 794 kb)
Rights and permissions
About this article
Cite this article
Sripada, L., Singh, K., Lipatova, A.V. et al. hsa-miR-4485 regulates mitochondrial functions and inhibits the tumorigenicity of breast cancer cells. J Mol Med 95, 641–651 (2017). https://doi.org/10.1007/s00109-017-1517-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-017-1517-5