Abstract
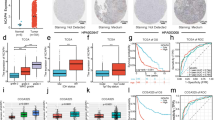
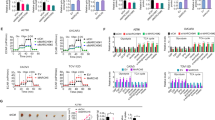
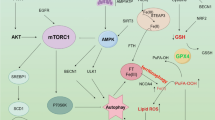
Mitochondrial glutaminase (GA) plays an essential role in cancer cell metabolism, contributing to biosynthesis, bioenergetics, and redox balance. Humans contain several GA isozymes encoded by the GLS and GLS2 genes, but the specific roles of each in cancer metabolism are still unclear. In this study, glioma SFxL and LN229 cells with silenced isoenzyme glutaminase KGA (encoded by GLS) showed lower survival ratios and a reduced GSH-dependent antioxidant capacity. These GLS-silenced cells also demonstrated induction of apoptosis indicated by enhanced annexin V binding capacity and caspase 3 activity. GLS silencing was associated with decreased mitochondrial membrane potential (ΔΨm) (JC-1 dye test), indicating that apoptosis was mediated by mitochondrial dysfunction. Similar observations were made in T98 glioma cells overexpressing glutaminase isoenzyme GAB, encoded by GLS2, though some characteristics (GSH/GSSG ratio) were different in the differently treated cell lines. Thus, control of GA isoenzyme expression may prove to be a key tool to alter both metabolic and oxidative stress in cancer therapy. Interestingly, reactive oxygen species (ROS) generation by treatment with oxidizing agents: arsenic trioxide or hydrogen peroxide, synergizes with either KGA silencing or GAB overexpression to suppress malignant properties of glioma cells, including the reduction of cellular motility. Of note, negative modulation of GLS isoforms or GAB overexpression evoked lower c-myc and bcl-2 expression, as well as higher pro-apoptotic bid expression. Combination of modulation of GA expression and treatment with oxidizing agents may become a therapeutic strategy for intractable cancers and provides a multi-angle evaluation system for anti-glioma pre-clinical investigations.
Key message
-
Silencing GLS or overexpressing GLS2 induces growth inhibition in glioma cell lines.
-
Inhibition is synergistically enhanced after arsenic trioxide (ATO) or H2O2 treatment.
-
Glutatione levels decrease in GLS-silenced cells but augment if GLS2 is overexpressed.
-
ROS synergistically inhibit cell migration by GLS silencing or GLS2 overexpression.
-
c-myc, bid, and bcl-2 mediate apoptosis resulting from GLS silencing or GLS2 overexpression.











Similar content being viewed by others
References
Márquez J, de la Oliva AR, Matés JM, Segura JA, Alonso FJ (2006) Glutaminase: a multifaceted protein not only involved in generating glutamate. Neurochem Int 48:465–471
DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB (2008) The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab 7:11–20
Olalla L, Gutiérrez A, Campos JA, Khan ZU, Alonso FJ, Segura JA, Márquez J, Aledo JC (2002) Nuclear localization of L-type glutaminase in mammalian brain. J Biol Chem 277:38939–38944
Márquez J, Tosina M, de la Rosa V, Segura JA, Alonso FJ, Matés JM, Campos-Sandoval JA (2009) New insights into brain glutaminases: beyond their role on glutamatergic transmission. Neurochem Int 55:64–70
Dang CV (2009) MYC, microRNAs and glutamine addiction in cancers. Cell Cycle 8:3243–3245
Wang JB, Erickson JW, Fuji R, Ramachandran S, Gao P, Dinavahi R, Wilson KF, Ambrosio AL, Dias SM, Dang CV et al (2010) Targeting mitochondrial glutaminase activity inhibits oncogenic transformation. Cancer Cell 18:207–219
Suzuki S, Tanaka T, Poyurovsky MV, Nagano H, Mayama T, Ohkubo S, Lokshin M, Hosokawa H, Nakayama T, Suzuki Y et al (2010) Phosphate-activated glutaminase (GLS2), a p53-inducible regulator of glutamine metabolism and reactive oxygen species. Proc Natl Acad Sci U S A 107:7461–7466
Matés JM, Segura JA, Martín-Rufián M, Campos-Sandoval JA, Alonso FJ, Márquez J (2013) Glutaminase isoenzymes as key regulators in metabolic and oxidative stress against cancer. Curr Mol Med 13:514–534
Gao P, Tchernyshyov I, Chang TC, Lee YS, Kita K, Ochi T, Zeller KI, De Marzo AM, Van Eyk JE, Mendell JT et al (2009) c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458:762–765
Martín-Rufián M, Tosina M, Campos-Sandoval JA, Manzanares E, Lobo C, Segura JA, Alonso FJ, Matés JM, Márquez J (2012) Mammalian glutaminase Gls2 gene encodes two functional alternative transcripts by a surrogate promoter usage mechanism. PLoS One 7:e38380
de la Rosa V, Campos-Sandoval JA, Martín-Rufián M, Cardona C, Matés JM, Segura JA, Alonso FJ, Márquez J (2009) A novel glutaminase isoform in mammalian tissues. Neurochem Int 55:76–84
Szeliga M, Obara-Michlewska M, Matyja E, Łazarczyk M, Lobo C, Hilgier W, Alonso FJ, Marquez J, Albrecht J (2009) Transfection with liver-type glutaminase cDNA alters gene expression and reduces survival, migration and proliferation of T98G glioma cells. Glia 57:1014–1023
Cheng T, Sudderth J, Yang C, Mullen AR, Jin ES, Matés JM, DeBerardinis RJ (2011) Pyruvate carboxylase is required for glutamine-independent growth of tumor cells. Proc Natl Acad Sci U S A 108:8674–8679
Wise DR, DeBerardinis RJ, Mancuso A, Sayed N, Zhang XY, Pfeiffer HK, Nissim I, Daikhin E, Yudkoff M, McMahon SB et al (2008) Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc Natl Acad Sci U S A 105:18782–18787
Hu W, Zhang C, Wu R, Sun Y, Levine A, Feng Z (2010) Glutaminase 2, a novel p53 target gene regulating energy metabolism and antioxidant function. Proc Natl Acad Sci U S A 107:7455–7460
Vander Heiden MG, Cantley LC, Thompson CB (2009) Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324:1029–1033
Kung HN, Marks JR, Chi JT (2011) Glutamine synthetase is a genetic determinant of cell type-specific glutamine independence in breast epithelia. PLoS Genet 7:e1002229
Seltzer MJ, Bennett BD, Joshi AD, Gao P, Thomas AG, Ferraris DV, Tsukamoto T, Rojas CJ, Slusher BS, Rabinowitz JD et al (2010) Inhibition of glutaminase preferentially slows growth of glioma cells with mutant IDH1. Cancer Res 70:8981–8987
Yuneva M, Zamboni N, Oefner P, Sachidanandam R, Lazebnik Y (2007) Deficiency in glutamine but not glucose induces MYC-dependent apoptosis in human cells. J Cell Biol 178:93–105
Yuneva MO, Fan TW, Allen TD, Higashi RM, Ferraris DV, Tsukamoto T, Matés JM, Alonso FJ, Wang C, Seo Y et al (2012) The metabolic profile of tumors depends on both the responsible genetic lesion and tissue type. Cell Metab 15:157–170
Marin-Valencia I, Yang C, Mashimo T, Cho S, Baek H, Yang XL, Rajagopalan KN, Maddie M, Vemireddy V, Zhao Z et al (2012) Analysis of tumor metabolism reveals mitochondrial glucose oxidation in genetically diverse human glioblastomas in the mouse brain in vivo. Cell Metab 15:827–837. Erratum in. Cell Metab 16:686
Maher EA, Marin-Valencia I, Bachoo RM, Mashimo T, Raisanen J, Hatanpaa KJ, Jindal A, Jeffrey FM, Choi C, Madden C et al (2012) Metabolism of [U-13 C]glucose in human brain tumors in vivo. NMR Biomed 25:1234–1244
Weinberg F, Hamanaka R, Wheaton WW, Weinberg S, Joseph J, Lopez M, Kalyanaraman B, Mutlu GM, Budinger GR, Chandel NS (2010) Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc Natl Acad Sci U S A 107:8788–8793
Sun RC, Board PG, Blackburn AC (2011) Targeting metabolism with arsenic trioxide and dichloroacetate in breast cancer cells. Mol Cancer 10:142
Matés JM, Segura JA, Alonso FJ, Márquez J (2012) Oxidative stress in apoptosis and cancer: an update. Arch Toxicol 86:1649–1665
Rahman I, Kode A, Biswas SK (2006) Assay for quantitative determination of glutathione and glutathione disulfide levels using enzymatic recycling method. Nat Protoc 1:3159–3165
Thangavelu K, Pan CQ, Karlberg T, Balaji G, Uttamchandani M, Suresh V, Schüler H, Low BC, Sivaraman J (2012) Structural basis for the allosteric inhibitory mechanism of human kidney-type glutaminase (KGA) and its regulation by Raf-Mek-Erk signaling in cancer cell metabolism. Proc Natl Acad Sci U S A 109:7705–7710
Donadio AC, Lobo C, Tosina M, de la Rosa V, Martín-Rufián M, Campos-Sandoval JA, Matés JM, Márquez J, Alonso FJ, Segura JA (2008) Antisense glutaminase inhibition modifies the O-GlcNAc pattern and flux through the hexosamine pathway in breast cancer cells. J Cell Biochem 103:800–811
Yi W, Clark PM, Mason DE, Keenan MC, Hill C, Goddard WA 3rd, Peters EC, Driggers EM, Hsieh-Wilson LC (2012) Phosphofructokinase 1 glycosylation regulates cell growth and metabolism. Science 337:975–980
Souers AJ, Leverson JD, Boghaert ER, Ackler SL, Catron ND, Chen J, Dayton BD, Ding H, Enschede SH, Fairbrother WJ et al (2013) ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat Med 19:202–208
Liu J, Albrecht AM, Ni X, Yang J, Li M (2013) Glioblastoma tumor initiating cells: therapeutic strategies targeting apoptosis and MicroRNA pathways. Curr Mol Med 13:352–357
Szeliga M, Bogacińska-Karaś M, Różycka A, Hilgier W, Marquez J, Albrecht J (2013) Silencing of GLS and overexpression of GLS2 genes cooperate in decreasing the proliferation and viability of glioma cells. Tumor Biol. doi:10.1007/s13277-013-1247-4
Lee MJ, Ye AS, Gardino AK, Heijink AM, Sorger PK, MacBeath G, Yaffe MB (2012) Sequential application of anticancer drugs enhances cell death by rewiring apoptotic signaling networks. Cell 149:780–794
Matés JM, Segura JA, Alonso FJ, Márquez J (2011) Anticancer antioxidant regulatory functions of phytochemicals. Curr Med Chem 18:2315–2338
Katt WP, Ramachandran S, Erickson JW, Cerione RA (2012) Dibenzophenanthridines as inhibitors of glutaminase C and cancer cell proliferation. Mol Cancer Ther 11:1269–1278
Szeliga M, Zgrzywa A, Obara-Michlewska M, Albrecht J (2012) Transfection of a human glioblastoma cell line with liver-type glutaminase (LGA) down-regulates the expression of DNA-repair gene MGMT and sensitizes the cells to alkylating agents. J Neurochem 123:428–436
Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M, Perera RM, Ferrone CR, Mullarky E, Shyh-Chang N et al (2013) Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496:101–105
Pérez-Gómez C, Matés JM, Gómez-Fabre PM, del Castillo-Olivares A, Alonso FJ, Márquez J (2003) Genomic organization and transcriptional analysis of the human L-glutaminase gene. Biochem J 370:771–784
Soga T (2013) Cancer metabolism: key players in metabolic reprogramming. Cancer Sci 104:275–281
Acknowledgements
We apologize for omissions of original references because of space limitations. Thanks are due to Rosa M. Moreno and Juan I. Rodríguez-Arranz for their valuable help in experimental tasks. This work was supported by Ministerio de Educación of Spain, PHB2010-0014-PC; Ministerio de Ciencia y Tecnología of Spain, SAF2010-17573; Junta de Andalucía, CVI-6656, Spain; PI-0825-2010, Junta de Andalucía, Spain; grant RD06/1012 of the RTA RETICS network from the Spanish Health Institute Carlos III, Spain. R.J.D. is supported by grants from the NIH (R01 CA157996), the Cancer Prevention and Research Institute of Texas (HIRP100437 and RP101243), the Robert A. Welch Foundation (I-1733), and the Damon-Runyon Cancer Research Foundation. M.S. and J.A. received support from the Ministry of Science and Higher Education (National Science Centre), grant NN401 039238. Thanks are also due to CAPES/DGU 250/11, Brazil.
Conflict of interest
The authors declare no conflict of interests related to this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Martín-Rufián, M., Nascimento-Gomes, R., Higuero, A. et al. Both GLS silencing and GLS2 overexpression synergize with oxidative stress against proliferation of glioma cells. J Mol Med 92, 277–290 (2014). https://doi.org/10.1007/s00109-013-1105-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-013-1105-2