Abstract
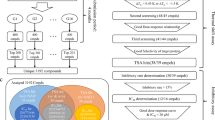
Sirt1, namely silent information regulator 1, belongs to a highly conserved family of NAD+-dependent deacetylase which is involved in innumerable human disorders such as obesity, type 2 diabetes, cancer, and aging. Combined high-throughput virtual screening, molecular dynamics simulation, MM/GBSA free energy calculation, and MM/GBSA free energy decomposition analysis approaches were utilized for identification of sirt1 activators. Four compounds with diverse chemical scaffold were retrieved as hits based on docking score and clustering analysis. Our simulations indicated that compound y040-6677 had the highest binding free energies, which could form one hydrogen bond with the residue Asn226. Compound y040-6677 could tightly plug into the hydrophobic allosteric site of sirt1 via strong interaction with the residues Leu215, Thr219, Gln222, Ile223, and Asn226, which were obtained from the MM/GBSA free energy decomposition. These simulation results were consistent with the in vivo enzymatic assay, which implied that compound y040-6677 had a comparable sirt1 activation compared with the reference molecule SRT1720. We hope that compound y040-6677 might represent a promising chemical scaffold for further development of novel sirt1 activators.





Similar content being viewed by others
References
Afshar G, Murnane JP (1999) Characterization of a human gene with sequence homology to Saccharomyces cerevisiae SIR2. Gene 234:161–168
Blander G, Guarente L (2004) The Sir2 family of protein deacetylases. Annu Rev Biochem 73:417–435
Cao D, Wang M, Qiu X, Liu D, Jiang H, Yang N, Xu RM (2015) Structural basis for allosteric, substrate-dependent stimulation of SIRT1 activity by resveratrol. Genes Dev 29:1316–1325
Case DA, Cheatham 3rd TE, Darden T, Gohlke H, Luo R, Merz Jr. KM, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688
Cerqueira NM, Gesto D, Oliveira EF, Santos-Martins D, Bras NF, Sousa SF, Fernandes PA, Ramos MJ (2015) Receptor-based virtual screening protocol for drug discovery. Arch Biochem Biophys 582:56–67
Chen F, Liu H, Sun H, Pan P, Li Y, Li D, Hou T (2016) Assessing the performance of the MM/PBSA and MM/GBSA methods. 6. Capability to predict protein-protein binding free energies and re-rank binding poses generated by protein-protein docking. Phys Chem Chem Phys 18:22129–22139
Cheng W, Wang JF, Yang CX, Wu L, Yin Q, Liu H, Fu ZJ (2016) Intrathecal injection of resveratrol attenuates burn injury pain by activating spinal sirtuin 1. Pharmacogn Mag 12:S201–S205
Dai H, Case AW, Riera TV, Considine T, Lee JE, Hamuro Y, Zhao H, Jiang Y, Sweitzer SM, Pietrak B, Schwartz B, Blum CA, Disch JS, Caldwell R, Szczepankiewicz B, Oalmann C, Yee Ng P, White BH, Casaubon R, Narayan R, Koppetsch K, Bourbonais F, Wu B, Wang J, Qian D, Jiang F, Mao C, Wang M, Hu E, Wu JC, Perni RB, Vlasuk GP, Ellis JL (2015) Crystallographic structure of a small molecule SIRT1 activator-enzyme complex. Nat Commun 6:7645
Gao J, Liang L, Chen Q, Zhang L, Huang T (2018) Insight into the molecular mechanism of yeast acetyl-coenzyme A carboxylase mutants F510I, N485G, I69E, E477R, and K73R resistant to soraphen A. J Comput Aided Mol Des 32:547–557
Gao J, Zhang Y, Chen H, Chen Q, Feng D, Zhang L, Li C (2019) Computational insights into the interaction mechanism of transcription cofactor vestigial-like protein4 binding to TEA domain transcription factor 4 by molecular dynamics simulation and molecular mechanics generalized Born/surface area) calculation. J Biomol Struct Dyn 37:2538–2545
Ge LQ, Li CZ, Wang ZR, Zhang Y, Chen L (2017) Suppression of oxidative stress and apoptosis in electrically stimulated neonatal rat cardiomyocytes by resveratrol and underlying mechanisms. J Cardiovasc Pharmacol 70:396–404
He WJ, Wang YY, Zhang MZ, You L, Davis LS, Fan H, Yang HC, Fogo AB, Zent R, Harris RC, Breyer MD, Hao CM (2010) Sirt1 activation protects the mouse renal medulla from oxidative injury. J Clin Investig 120:1056–1068
Hubbard BP, Gomes AP, Dai H, Li J, Case AW, Considine T, Riera TV, Lee JE, E SY, Lamming DW, Pentelute BL, Schuman ER, Stevens LA, Ling AJY, Armour SM, Michan S, Zhao H, Jiang Y, Sweitzer SM, Blum CA, Disch JS, Ng PY, Howitz KT, Rolo AP, Hamuro Y, Moss J, Perni RB, Ellis JL, Vlasuk GP, Sinclair DA (2013) Evidence for a common mechanism of SIRT1 regulation by allosteric activators. Science 339:1216–1219
Kitada M, Kume S, Takeda-Watanabe A, Kanasaki K, Koya D (2013) Sirtuins and renal diseases: relationship with aging and diabetic nephropathy. Clin Sci 124:153–164
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W, Donini O, Cieplak P, Srinivasan J, Case DA, Cheatham 3rd TE (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33:889–897
Kumar A, Chauhan S (2016) How much successful are the medicinal chemists in modulation of SIRT1: a critical review. Eur J Med Chem 119:45–69
Liu YW, Hao YC, Chen YJ, Yin SY, Zhang MY, Kong L, Wang TY (2018) Protective effects of sarsasapogenin against early stage of diabetic nephropathy in rats. Phytother Res 32:1574–1582
Longo VD, Kennedy BK (2006) Sirtuins in aging and age-related disease. Cell 126:257–268
Lu Q, Ji XJ, Zhou YX, Yao XQ, Liu YQ, Zhang F, Yin XX (2015) Quercetin inhibits the mTORC1/p70S6K signaling-mediated renal tubular epithelial-mesenchymal transition and renal fibrosis in diabetic nephropathy. Pharmacol Res 99:237–247
Sacconnay L, Carrupt PA, Nurisso A (2016) Human sirtuins: structures and flexibility. J Struct Biol 196:534–542
Sun H, Li Y, Tian S, Xu L, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set. Phys Chem Chem Phys 16:16719–16729
Villalba JM, Alcain FJ (2012) Sirtuin activators and inhibitors. Biofactors 38:349–359
Vyas VK, Goel A, Ghate M, Patel P (2015) Ligand and structure-based approaches for the identification of SIRT1 activators. Chem Biol Interact 228:9–17
Wakino S, Hasegawa K, Itoh H (2015) Sirtuin and metabolic kidney disease. Kidney Int 88:691–698
Weiser J, Shenkin PS, Still WC (1999) Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J Comput Chem 20:217–230
Xie J, Zhang XM, Zhang L (2013) Negative regulation of inflammation by SIRT1. Pharmacol Res 67:60–67
Yorimitsu T, Nair U, Yang ZF, Klionsky DJ (2006) Endoplasmic reticulum stress triggers autophagy. J Biol Chem 281:30299–30304
Zhou L, Fu L, Lv N, Chen XS, Liu J, Li Y, Xu QY, Huang S, Zhang XD, Dou LP, Wang LL, Li YH, Yu L (2017) A minicircuitry comprised of microRNA-9 and SIRT1 contributes to leukemogenesis in t(8;21) acute myeloid leukemia. Eur Rev Med Pharmacol Sci 21:786–794
Zhu SP, Liu G, Wu XT, Chen FX, Liu JQ, Zhou ZH, Zhang JF, Fei SJ (2013) The effect of phloretin on human gammadelta T cells killing colon cancer SW-1116 cells. Int Immunopharmacol 15:6–14
Acknowledgements
This work was supported by National Natural Science Foundation of China (NSFC No. 21708033), Six Talent Peaks Project in Jiangsu Province (grant number YY-046), and the Qinglan Project of Jiangsu Province of China.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
An, Y., Meng, C., Chen, Q. et al. Discovery of small molecule sirt1 activator using high-throughput virtual screening, molecular dynamics simulation, molecular mechanics generalized born/surface area (MM/GBSA) calculation, and biological evaluation. Med Chem Res 29, 255–261 (2020). https://doi.org/10.1007/s00044-019-02479-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-019-02479-2