Abstract
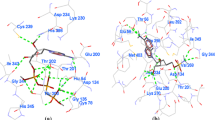
Farnesyltransferase (FTase) is one of the prenyltransferase family enzymes that catalyse the transfer of 15-membered isoprenoid (farnesyl) moiety to the cysteine of CAAX motif-containing proteins including Rho and Ras family of G proteins. Inhibitors of FTase act as drugs for cancer, malaria, progeria and other diseases. In the present investigation, we have developed two structure-based pharmacophore models from protein–ligand complex (3E33 and 3E37) obtained from the protein data bank. Molecular dynamics (MD) simulations were performed on the complexes, and different conformers of the same complex were generated. These conformers were undergone protein–ligand interaction fingerprint (PLIF) analysis, and the fingerprint bits have been used for structure-based pharmacophore model development. The PLIF results showed that Lys164, Tyr166, TrpB106 and TyrB361 are the major interacting residues in both the complexes. The RMSD and RMSF analyses on the MD-simulated systems showed that the absence of FPP in the complex 3E37 has significant effect in the conformational changes of the ligands. During this conformational change, some interactions between the protein and the ligands are lost, but regained after some simulations (after 2 ns). The structure-based pharmacophore models showed that the hydrophobic and acceptor contours are predominantly present in the models. The pharmacophore models were validated using reference compounds, which significantly identified as HITs with smaller RMSD values. The developed structure-based pharmacophore models are significant, and the methodology used in this study is novel from the existing methods (the original X-ray crystallographic coordination of the ligands is used for the model building). In our study, along with the original coordination of the ligand, different conformers of the same complex (protein–ligand) are used. It concluded that the developed methodology is significant for the virtual screening of novel molecules on different targets.











Similar content being viewed by others
References
Arooj M, Sakkiah S, Kim S, Arulalapperumal V, Lee KW (2013) A combination of receptor-based pharmacophore modeling & QM techniques for identification of human chymase inhibitors. PLoS One 8:e63030. doi:10.1371/journal.pone.0063030
Bellesia F, Choi SR, Felluga F, Fiscaletti G, Ghelfi F, Menziani MC, Parsons AF, Poulter CD, Roncaglia F, Sabbatini M, Spinelli D (2013) Novel route to chaetomellic acid A and analogues: serendipitous discovery of a more competent FTase inhibitor. Bioorg Med Chem 21:348–358
Buckner FS, Eastman RT, Nepomuceno-Silva JL, Speelmon EC, Myler PJ, Van Voorhis WC, Yokoyama K (2002) Cloning, heterologous expression, and substrate specificities of protein farnesyltransferases from Trypanosoma cruzi and Leishmania major. Mol Biochem Parasitol 122:181–188
Buckner FS, Eastman RT, Yokoyama K, Gelb MH, Van Voorhis WC (2005) Protein farnesyl transferase inhibitors for the treatment of malaria and African trypanosomiasis. Curr Opin Invest Drugs 6:791–797
Capell BC, Erdos MR, Madigan JP, Fiordalisi JJ, Varga R, Conneely KN, Gordon LB, Der CJ, Cox AD, Collins FC (2005) Inhibiting farnesylation of progerin prevents the characteristic nuclear blebbing of Hutchinson-Gilford progeria syndrome. Proc Natl Acad Sci USA 102:12879–12884
Case DA, Darden TA, Cheatham TE III, Simmerling CL, Wang J, Duke RE, Luo R, Walker RC, Zhang W, Merz KM, Roberts B, Hayik S, Roitberg A, Seabra G, Swails J, Götz AW, Kolossváry I, Wong KF, Paesani F, Vanicek J, Wolf RM, Liu J, Wu X, Brozell SR, Steinbrecher T, Gohlke H, Cai Q, Ye X, Wang J, Hsieh MJ, Cui G, Roe DR, Mathews DH, Seetin MG, Salomon-Ferrer R, Sagui C, Babin V, Luchko T, Gusarov S, Kovalenko A, Kollman PA (2012) AMBER 12. University of California, San Francisco
Casey PJ, Seabra MC (1996) Protein prenyltransferases. J Biol Chem 271:5289–5292
Chen J, Lai L (2006) Pocket v. 2: further developments on receptor-based pharmacophore modelling. J Chem Inf Model 46:2684–2691
Eastman RT, White J, Hucke O, Bauer K, Yokoyama K, Nallan L, Chakrabarti D, Verlinde CL, Gelb MH, Rathod PK, Van Voorhis WC (2005) Resistance to a protein farnesyltransferase inhibitor in Plasmodium falciparum. J Biol Chem 280:13554–13559
Esteva MI, Kettler K, Maidana C, Fichera L, Ruiz AM, Bontempi EJ, Andersson B, Dahse HM, Haebel P, Ortmann R, Klebe G, Schlitzer M (2005) Benzophenone-based farnesyltransferase inhibitors with high activity against Trypanosoma cruzi. J Med Chem 48:7186–7191
Guner OF (2002) History and evolution of the pharmacophore concept in computer-aided drug design. Curr Top Med Chem 2:1321–1332
Hast MA, Fletcher S, Cummings CG, Pusateri EE, Blaskovich MA, Rivas K, Gelb MH, Van Voorhis WC, Sebti SM, Hamilton AD, Beese LS (2009) Structural basis for binding and selectivity of antimalarial and anticancer ethylenediamine inhibitors to protein farnesyltransferase. Chem Biol 16:181–192
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65:712–725
Humphrey W, Dalke A, Schulten K (1996) VMD—visual molecular dynamics. J Mol Graph 14:33–38
Kurczab R, Bojarski AJ (2013) New strategy for receptor-based pharmacophore query construction: a case study for 5-HT7 receptor ligands. J Chem Inf Model 53:3233–3243
Long SB, Hancock PJ, Kral AM, Hellinga HW, Beese LS (2001) The crystal structure of human protein farnesyltransferase reveals the basis for inhibition by CaaX tetrapeptides and their mimetics. Proc Natl Acad Sci USA 98:12948–12953
Long SB, Casey PJ, Beese LS (2002) Reaction path of protein farnesyltransferase at atomic resolution. Nature 419:645–650
Loving K, Salam NK, Sherman W (2009) Energetic analysis of fragment docking and application to structure-based pharmacophore hypothesis generation. J Comput Aided Mol Des 23:541–554
Meslamani J, Li J, Sutter J, Stevens A, Bertrand HO, Rognan D (2012) Protein–ligand-based pharmacophores: generation and utility assessment in computational ligand profiling. J Chem Inf Model 52:943–955
MOE (Molecular Operating Environment) (2013) Chemical Computing Group Inc. Montreal, H3A 2R7 Canada
Moorthy NSHN, Sousa SF, Fernandes PA, Ramos MJ (2011a) Structural feature study of benzofuran derivatives as farnesyltransferase inhibitors. J Enz Inhibit Med Chem 26(6):777–791
Moorthy NSHN, Sousa SF, Ramos MJ, Fernandes PA (2011b) In silico based structural analysis of arylthiophene derivatives for FTase inhibitory activity, hERG and other toxic effects. J Biomol Screen 16:1037–1046
Moorthy NSHN, Sousa SF, Fernandes PA, Ramos MJ (2012) In silico based structural analysis of some piperidine analogs as farnesyltransferase inhibitors. Med Chem 8:853–864
Moorthy NSHN, Sousa SF, Ramos MJ, Fernandes PA (2013) Farnesyltransferease inhibitors: a comprehensive review based on quantitative structural analysis. Curr Med Chem 20:4888–4923
Moorthy NSHN, Cerquira NMFS, Fernandes PA, Ramos MJ (2015) Ligand based analysis on HMG-CoA reductase inhibitors. Chemom Intell Lab Syst 140:102–116
Ohkanda J, Buckner FS, Lockman JW, Yokoyama K, Carrico D, Eastman R, Luca-Fradley K, Davies W, Croft SL, Van Voorhis WC, Gelb MH, Sebti SM, Hamilton AD (2004) Design and synthesis of peptidomimetic protein farnesyltransferase inhibitors as anti-Trypanosoma brucei agents. J Med Chem 47:432–445
Park HW, Boduluri SR, Moomaw JF, Casey PJ, Beese LS (1997) Crystal structure of protein farnesyltransferase at 2.25 angstrom resolution. Science 275:1800–1804
Pickett JS, Bowers KE, Fierke CA (2003a) Mutagenesis studies of protein farnesyltransferase implicate aspartate {beta}352 as a magnesium ligand. J Biol Chem 278:51243–51250
Pickett JS, Bowers KE, Hartman HL, Fu HW, Embry AC, Casey PJ, Fierke CA (2003b) Kinetic studies of protein farnesyltransferase mutants establish active substrate conformation. Biochemistry 42:9741–9748
Reid TS, Long SB, Beese LS (2004a) Crystallographic analysis reveals that anticancer clinical candidate L-778,123 inhibits protein farnesyltransferase and geranylgeranyltransferase-I by different binding modes. Biochemistry 43:9000–9008
Reid TS, Terry KL, Casey PJ, Beese LS (2004b) Crystallographic analysis of CaaX prenyltransferases complexed with substrates defines rules of protein substrate selectivity. J Mol Biol 343:417–433
Rella M, Rushworth CA, Guy JL, Turner AJ, Langer T, Jackson RM (2006) Structure-based pharmacophore design and virtual screening for novel angiotensin converting enzyme 2 inhibitors. J Chem Inf Model 46:708–716
Sanders MPA, Verhoeven S, de Graaf C, Roumen L, Vroling B, Nabuurs NB, de Vlieg J, Klomp JPG (2011) Snooker: a structure-based pharmacophore generation tool applied to class A GPCRs. J Chem Inf Model 51:2277–2292
Shen M, Pan P, Li Y, Li D, Yu H, Hou T (2015) Farnesyltransferase and geranylgeranyltransferase I: structures, mechanism, inhibitors and molecular modelling. Drug Discov Today 20(2):267–276. doi:10.1016/j.drudis.2014.10.002
Sousa SF, Fernandes PA, Ramos MJ (2005) Farnesyltransferase—new insights into the zinc-coordination sphere paradigm: evidence for a carboxylate-shift mechanism. Biophys J 88:483–494
Sousa SF, Fernandes PA, Ramos MJ (2007a) Effective tailor-made force field parameterization of the several Zn coordination environments in the puzzling FTase enzyme: opening the door to the full understanding of its elusive catalytic mechanism. Theor Chem Acc 117:171–181
Sousa SF, Fernandes PA, Ramos MJ (2007b) Theoretical studies on farnesyltransferase: the distances paradox explained. Proteins 66:205–218
Sousa SF, Fernandes PA, Ramos MJ (2007c) Theoretical studies on farnesyltransferase: evidence for thioether product coordination to the active-site zinc sphere. J Comput Chem 28:1160–1168
Sousa SF, Fernandes PA, Ramos MJ (2008) Enzyme flexibility and the catalytic mechanism of farnesyltransferase: targeting the relation. J Phys Chem B 112:8681–8691
Sousa SF, Fernandes PA, Ramos MJ (2009) Molecular dynamics simulations on the critical states of the farnesyltransferase enzyme. Bioorg Med Chem 17:3369–3378
Sousa SF, Coimbra JTS, Paramos D, Pinto R, Guimarães RS, Teixeira V, Fernandes PA, Ramos MJ (2013) Molecular dynamics analysis of a series of 22 potential farnesyltransferase substrates containing a CaaX-motif. J Mol Model 19:673–688
Taylor JS, Reid TS, Terry KL, Casey PJ, Beese LS (2003) Structure of mammalian protein geranylgeranyltransferase type-I. EMBO J 22:5963–5974
Tobin DA, Pickett JS, Hartman HL, Fierke CA, Penner-Hahn JE (2003) Structural characterization of the zinc site in protein farnesyltransferase. J Am Chem Soc 125:9962–9969
Troutman JM, Subramanian T, Andres DA, Spielmann HP (2007) Selective modification of CaaX peptides with ortho-substituted anilinogeranyl lipids by protein farnesyl transferase: competitive substrates and potent inhibitors from a library of farnesyl diphosphate analogues. Biochemistry 46:11310–11321
Vasanthanathan P, Moorthy NSHN, Kongsted J (2014) Dual mechanism of HIV-1 integrase and RNase H inhibition by diketo derivatives: a computational study. RSC Adv 4:38672–38681
Yokoyama K, Trobridge P, Buckner FS, Van Voorhis WC, Stuart KD, Gelb MH (1998) Protein farnesyltransferase from Trypanosoma brucei. A heterodimer of 61- and 65-kDa subunits as a new target for antiparasite therapeutics. J Biol Chem 273:26497–26505
Zhang FL, Casey PJ (1996) Protein prenylation: molecular mechanisms and functional consequences. Annu Rev Biochem 65:241–269
Acknowledgments
One of the authors (N.S.H.N. Moorthy) gratefully acknowledges the Fundaçao para a Ciencia e Technologia (FCT), Portugal, for a postdoctoral grant (SFRH/BPD/44469/2008).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Moorthy, N.S.H.N., Sousa, S.F., Ramos, M.J. et al. Binding mode of conformations and structure-based pharmacophore development for farnesyltransferase inhibitors. Med Chem Res 25, 1340–1357 (2016). https://doi.org/10.1007/s00044-016-1578-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-016-1578-y