Abstract
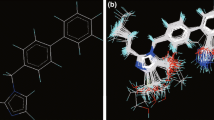
Partial least squares (PLS) method as ligand-based method was applied for building quantitative structure–activity relationships (QSAR) regression model to predict the inhibitory activity of some A2B antagonists. The accuracy and predictability of the developed model were evaluated by several validation methods using external and internal test sets and also the criteria recommended by Tropsha and Roy were met. The result of the PLS model had a high statistical quality (R 2 = 0.936 and Q 2 = 0.867) for predicting the activity of the compounds. Evaluation of a test set of seven compounds with the developed PLS model revealed that this model is reliable and has a good predictability. Because of high correlation between the predicted and experimental values of activity, PLS model proved to be a highly predictive QSAR approach. Also, the reliability of the model was assessed through docking for the selected antagonists as structure-based method. A potential binding site of A2BAR was verified according to the previous studies of site-directed mutagenesis. Phe173, Glu174, His251, Asn254, Lys269, Ile276, and His 280 were determined to be involved in ligand–receptor interactions.




Similar content being viewed by others
References
Bonet I, Franco-Montero P, Rivero V, Teijeira M, Borges F, Uriarte E, Helguera AM (2013) Classifier ensemble based on feature selection and diversity measures for predicting the affinity of A2B adenosine receptor antagonists. J Chem Inf Model 53:3140–3155
Cheng F, Xu Z, Liu G, Tang Y (2010) Insights into binding modes of adenosine A(2B) antagonists with ligand-based and receptor-based methods. Europ J Med Chem 45:3459–3471
Cheung AW, Brinkman J, Firooznia F, Flohr A, Grimsby J, Gubler ML, Guertin K, Hamid R, Marcopulos N, Norcross RD, Qi L, Ramsey G, Tan J, Wen Y, Sarabu R (2010) 4-Substituted-7-N-alkyl-N-acetyl 2-aminobenzothiazole amides: drug-like and non-xanthine based A2B adenosine receptor antagonists. Bioorg Med Chem Lett 20:4140–4146
Debnath AK (2001) Quantitative structure–activity relationship (QSAR): a versatile tool in drug design. In: Ghose AK, Viswanadhan VN (eds) Combinatorial library design and evaluation: principles, software tools, and applications in drug discovery. Marcel Dekker, New York, pp 73–129
Deeb O, Hemmateenejad B, Jaber A, Garduno-Juarez R, Miri R (2007) Effect of the electronic and physicochemical parameters on the carcinogenesis activity of some sulfa drugs using QSAR analysis based on genetic-MLR and genetic-PLS. Chemosphere 67:2122–2130
Dubey RK, Gillespie DG, Mi Z, Jackson EK (2005) Adenosine inhibits PDGF-induced growth of human glomerular mesangial cells via A2B receptors. Hypertension 46:628–634
Fernandez M, Caballero J (2006) Modeling of activity of cyclic urea HIV-1 protease inhibitors using regularized-artificial neural networks. Bioorg Med Chem 14:280–294
Fitzgerald GA (2005) Opinion: anticipating change in drug development: the emerging era of translational medicine and therapeutics. Nat Rev Drug Discov 4:815–818
Fredholm BB, IJzerman AP, Jacobson KA, Klotz KN, Linden J (2001) International union of pharmacology. XXV. Nomenclature and classification of adenosine receptors. Pharmacol Rev 53:527–552
Fredholm BB, IJzerman AP, Jacobson KA, Linden J, Müller CE (2011) Nomenclature and classification of adenosine receptors—an update. Pharmacol Rev 63:1–34
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Zakrzewski VG et al (1998) Gaussian 98, revision A7. Gaussian Inc., Pittsburgh
Geladi P, Kowalsky BR (1987) Partial least-squares regression: a tutorial. Anal Chim Acta 185:1–17
Golbraikh A, Tropsha A (2002) Beware of q2. J Mol Graph Model 20:269–276
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26:694–701
Hawkins DM (2004) The problem of overfitting. J Chem Inf Comput Sci 44:1–12
Hemmateenejad B, Miri R, Edraki N, Khoshneviszadeh M, Shafiee A (2007) Molecular modeling and QSAR analysis of some 4,5-dichloroimidazolyl-1,4-DHP based calcium channel blockers. J Iran Chem Soc 4:182–193
Ivanov AA, Wang B, Klutz AM, Chen VL, Gao ZG, Jacobson KA (2008) Probing distal regions of the A2B adenosine receptor by quantitative structure–activity relationship modeling of known and novel agonists. J Med Chem 51:2088–2099
Jaakola VP, Griffith MT, Hanson MA, Cherezov V, Chien EYT, Lane JR, Ijzerman AP, Stevens RC (2008) The 2.6 angstrom crystal structure of a human A(2A) adenosine receptor bound to an antagonist. Science 322:1211–1217
Jiang QL, Lee BX, Glashofer M, van Rhee AM, Jacobson KA (1997) Mutagenesis reveals structure–activity parallels between human A(2A) adenosine receptors and biogenic amine G protein coupled receptors. J Med Chem 40:2588–2595
Joseph TB, Kumar BVSS, Santhosh B, Kriti S, Pramod AB, Ravikumar M, Kishore M (2008) Quantitative structure activity relationship and pharmacophore studies of adenosine receptor A2B inhibitors. Chem Biol Drug Des 72:395–408
Kennard RW, Stone LA (1969) Computer aided design of experiments. Technometrics 11:137–149
Kim JH, Wess J, Vanrhee AM, Schoneberg T, Jacobson KA (1995) Site-directed mutagenesis identifies residues involved in ligand recognition in the human A2A adenosine receptor. J Biol Chem 270:13987–13997
Kim JH, Jiang QL, Glashofer M, Yehle S, Wess J, Jacobson KA (1996) Glutamate residues in the second extracellular loop of the human A(2a) adenosine receptor are required for ligand recognition. Mol Pharmacol 49:683–691
Kim SK, Gao ZG, Van Rompaey P, Gross AS, Chen A, Van Calenbergh S, Jacobson KA (2003) Modeling the adenosine receptors: comparison of the binding domains of A(2A) agonists and antagonists. J Med Chem 46:4847–4859
Kubinyi H, Hamprecht F, Mietzner AT (1998) Three-dimensional quantitative similarity–activity relationships (3D QSiAR) from SEAL similarity matrices. J Med Chem 41:2553–2564
Landells LJ, Jensen MW, Orr LM, Spina D, O’Connor BJ, Page CP (2000) The role of adenosine receptors in the action of theophylline on human peripheral blood mononuclear cells from healthy and asthmatic subjects. Br J Pharmacol 129:1140–1144
Mansourian M, Madadkar-Sobhani A, Mahnam K, Fassihi A, Saghaie L (2012) Characterization of adenosine receptor in its native environment: insights from molecular dynamics simulations of palmitoylated/glycosylated, membrane-integrated human A2B adenosine receptor. J Mol Model 18:4309–4324
Marshall GR (1994) Binding site modeling of unknown receptors. In: Kubinyi H (ed) 3D QSAR in drug design: theory, methods and applications. ESCOM, Leiden, pp 117–133
Mohajeri A, Hemmateenejad B, Mehdipour AR, Miri R (2008) Modeling calcium channel antagonistic activity of dihydropyridine derivatives using QTMS indices analyzed by GA-PLS and PC-GA-PLS. J Mol Graph Mod 26:1057–1065
Mohsenin A, Blackburn MR (2006) Adenosine signaling in asthma and chronic obstructive pulmonary disease. Curr Opin Pulm Med 12:54–59
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 19:1639–1662
Mustafa SJ, Nadeem A, Fan M, Zhong H, Belardinelli L, Zeng D (2007) Effect of a specific and selective A2B adenosine receptor antagonist on adenosine agonist AMP and allergen-induced airway responsiveness and cellular influx in a mouse model of asthma. J Pharmacol Exp Ther 320:1246–1251
Naes T, Isaksson T, Fearn T, Davies T (2004) A user-friendly guide to multivariate calibration and classification. NIR Publications, Chichester
Rosi S, McGann K, Hauss-Wegrzyniak B, Wenk GL (2003) The influence of brain inflammation upon neuronal adenosine A2B receptors. J Neurochem 86:220–227
Roy P, Roy K (2008) On some aspects of variable selection for partial least squares regression models. QSAR Comb Sci 27:302–313
Shahlaei M, Fassihi A, Saghaie L, Arkan E, Pourhossein A (2011) A modeling study of aldehyde inhibitors of human cathepsin K using partial least squares method. Res Pharm Sci 6:71–80
Sharma BK, Sarbhai K, Singh P (2010) A rationale for the activity profile of arylpiperazinylthioalkyls as 5-HT1A-serotonin and a1-adrenergic receptor ligands. Eur J Med Chem 45:1927–1934
Shen M, Beguin C, Golbraikh A, Stables J, Kohn H, Tropsha A (2004) Application of predictive QSAR models to database mining: identification and experimental validation of novel anticonvulsant compounds. J Med Chem 47:2356–2364
Sherbiny FF, Schiedel AC, Maass A, Muller C (2009) Homology modelling of the human adenosine A(2B) receptor based on X-ray structures of bovine rhodopsin, the beta(2)-adrenergic receptor and the human adenosine A(2A) receptor. J Comput Aided Mol Des 23:807–828
Snedecor GW, Cochran WG (1967) Statistical methods. Oxford and IBH Publishing Co. Pvt. Ltd., New Delhi, pp 381–418
Song Y, Coupar IM, Iskander MN (2001) Structural predictions of adenosine 2B antagonist affinity using molecular field analysis. Quant Struct Act Relat 20:23–30
Thanikaivelan P, Subramanian V, Rao JR, Nair BU (2000) Application of quantum chemical descriptor in quantitative structure activity and structure property relationship. Chem Phys Lett 323:59–70
Todeschini R, Consonni V, Mauri A, Pavan M (2002) DRAGON software, version 2.1. Milano, Italy. http://disat.unimib.it/chm/Dragon.htm/
Tropsha A, Gramatica P, Gombar V (2003) The importance of being Eearnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb Sci 22:69–77
Vandeginste BGM, Massart DL, Buydens LMC, De Jong S, Lewi PJ, Smeyers-Verbeke J (1998) Handbook of chemometrics and qualimetrics-part B. Elsevier Science, Amsterdam
Volpini R, Costanzi S, Vittori S, Cristalli G, Klotz KN (2003) Medicinal chemistry and pharmacology of A2B adenosine receptors. Curr Top Med Chem 3:427–443
Wallace AC, Laskowski RA, Thornton JM (1995) LIGPLOT—a program to generate schematic diagrams of protein ligand interactions. Protein Eng 8:127–134
Wold S, Sjostrom M, Eriksson L (2001) PLS-regression: a basic tool of chemometrics, Chemometr. Intell Lab Syst 58:109–130
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Mansourian, M., Fassihi, A., Saghaie, L. et al. QSAR and docking analysis of A2B adenosine receptor antagonists based on non-xanthine scaffold. Med Chem Res 24, 394–407 (2015). https://doi.org/10.1007/s00044-014-1133-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-014-1133-7