Abstract.
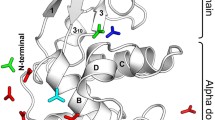
The three-dimensional structure of mouse lysozyme M, glycoside hydrolase, with 130 amino acids has been determined by heteronuclear NMR spectroscopy. We found that mouse lysozyme M had four α-helices, two 310 helices, and a double- and a triple-stranded anti-parallel β-sheet, and its structure was very similar to that of hen lysozyme in solution and in the crystalline state. The pH activity profile of p-nitrophenyl penta N-acetyl-β-D-chitopentaoside hydrolysis by mouse lysozyme M was similar to that of hen lysozyme, but the hydrolytic activity of mouse lysozyme M was lower. From analyses of binding affinities of lysozymes to a substrate analogue and internal motions of lysozymes, we suggest that the lower activity of mouse lysozyme M was due to the larger dissociation constant of its enzyme-substrate complex and the restricted internal backbone motions in the molecule.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received 18 September 2002; received after revision 25 October 2002; accepted 13 November 2002
RID="*"
ID="*"Corresponding author.
Rights and permissions
About this article
Cite this article
Obita, T., Ueda, T. & Imoto, T. Solution structure and activity of mouse lysozyme M. CMLS, Cell. Mol. Life Sci. 60, 176–184 (2003). https://doi.org/10.1007/s000180300012
Issue Date:
DOI: https://doi.org/10.1007/s000180300012