Abstract
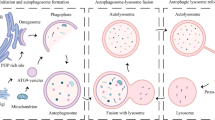
Autophagy is a lysosome-mediated degradative process that removes damaged proteins and organelles, during which autophagosome–lysosome fusion is a key step of the autophagic flux. Based on our observation that intermediate cytofilament keratin 8 (KRT8) enhances autophagic clearance in cells under oxidative stress condition, we investigated whether KRT8 supports the cytoplasmic architectural networks to facilitate the vesicular fusion entailing trafficking onto filamentous tracks. We found that KRT8 interacts with actin filaments via the cytolinker, plectin (PLEC) during trafficking of autophagosome. When PLEC was knocked down or KRT8 structure was collapsed by phosphorylation, autophagosome–lysosome fusion was attenuated. Inhibition of actin polymerization resulted in accumulation of autophagosomes owing to a decrease in autophagosome and lysosome fusion. Furthermore, myosin motor protein was found to be responsible for vesicular trafficking along the actin filaments to entail autolysosome formation. Thus, the autophagosome–lysosome fusion is aided by PLEC-stabilized actin filaments as well as intermediate cytofilament KRT8 that supports the structural integrity of actin filaments during macroautophagic process under oxidative stress condition.








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
References
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24(1):24–41. https://doi.org/10.1038/cr.2013.168
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132(1):27–42. https://doi.org/10.1016/j.cell.2007.12.018
Klionsky DJ (2005) The molecular machinery of autophagy: unanswered questions. J Cell Sci 118(Pt 1):7–18. https://doi.org/10.1242/jcs.01620
Klionsky DJ, Ohsumi Y (1999) Vacuolar import of proteins and organelles from the cytoplasm. Ann Rev Cell Dev Biol 15:1–32. https://doi.org/10.1146/annurev.cellbio.15.1.1
Abada A, Elazar Z (2014) Getting ready for building: signaling and autophagosome biogenesis. Embo Rep 15(8):839–852. https://doi.org/10.15252/embr.201439076
Fader CM, Sanchez D, Furlan M, Colombo MI (2008) Induction of autophagy promotes fusion of multivesicular bodies with autophagic vacuoles in k562 cells. Traffic 9(2):230–250. https://doi.org/10.1111/j.1600-0854.2007.00677.x
Berg TO, Fengsrud M, Stromhaug PE, Berg T, Seglen PO (1998) Isolation and characterization of rat liver amphisomes. Evidence for fusion of autophagosomes with both early and late endosomes. J Biol Chem 273(34):21883–21892. https://doi.org/10.1074/jbc.273.34.21883
Nakamura S, Yoshimori T (2017) New insights into autophagosome–lysosome fusion. J Cell Sci 130(7):1209–1216. https://doi.org/10.1242/jcs.196352
Mizushima N, Levine B, Cuervo AM, Klionsky DJ (2008) Autophagy fights disease through cellular self-digestion. Nature 451(7182):1069–1075. https://doi.org/10.1038/nature06639
Yu WH, Cuervo AM, Kumar A, Peterhoff CM, Schmidt SD, Lee JH, Mohan PS, Mercken M, Farmery MR, Tjernberg LO et al: Macroautophagy--a novel Beta-amyloid peptide-generating pathway activated in Alzheimer's disease. J Cell Biol (2005), 171(1):87–98. https://doi.org/10.1083/jcb.200505082
De Leo MG, Staiano L, Vicinanza M, Luciani A, Carissimo A, Mutarelli M, Di Campli A, Polishchuk E, Di Tullio G, Morra V et al: Autophagosome–lysosome fusion triggers a lysosomal response mediated by TLR9 and controlled by OCRL. Nat Cell Biol (2016), 18(8):839–850. https://doi.org/10.1038/ncb3386
Aplin A, Jasionowski T, Tuttle DL, Lenk SE, Dunn WA Jr (1992) Cytoskeletal elements are required for the formation and maturation of autophagic vacuoles. J Cell Physiol 152(3):458–466. https://doi.org/10.1002/jcp.1041520304
Mackeh R, Perdiz D, Lorin S, Codogno P, Pous C (2013) Autophagy and microtubules—new story, old players. J Cell Sci 126(Pt 5):1071–1080. https://doi.org/10.1242/jcs.115626
Geeraert C, Ratier A, Pfisterer SG, Perdiz D, Cantaloube I, Rouault A, Pattingre S, Proikas-Cezanne T, Codogno P, Pous C (2010) Starvation-induced hyperacetylation of tubulin is required for the stimulation of autophagy by nutrient deprivation. J Biol Chem 285(31):24184–24194. https://doi.org/10.1074/jbc.M109.091553
Maday S, Holzbaur EL (2014) Autophagosome biogenesis in primary neurons follows an ordered and spatially regulated pathway. Dev Cell 30(1):71–85. https://doi.org/10.1016/j.devcel.2014.06.001
Du W, Su QP, Chen Y, Zhu Y, Jiang D, Rong Y, Zhang S, Zhang Y, Ren H, Zhang C et al (2016) Kinesin 1 drives autolysosome tubulation. Dev Cell 37(4):326–336. https://doi.org/10.1016/j.devcel.2016.04.014
Aguilera MO, Beron W, Colombo MI (2012) The actin cytoskeleton participates in the early events of autophagosome formation upon starvation induced autophagy. Autophagy 8(11):1590–1603. https://doi.org/10.4161/auto.21459
Reggiori F, Monastyrska I, Shintani T, Klionsky DJ (2005) The actin cytoskeleton is required for selective types of autophagy, but not nonspecific autophagy, in the yeast Saccharomyces cerevisiae. Mol Biol Cell 16(12):5843–5856. https://doi.org/10.1091/mbc.e05-07-0629
Zhuo C, Ji Y, Chen Z, Kitazato K, Xiang Y, Zhong M, Wang Q, Pei Y, Ju H, Wang Y (2013) Proteomics analysis of autophagy-deficient Atg7-/- MEFs reveals a close relationship between F-actin and autophagy. Biochem Biophys Res Commun 437(3):482–488. https://doi.org/10.1016/j.bbrc.2013.06.111
Kruppa AJ, Kendrick-Jones J, Buss F (2016) Myosins, actin and autophagy. Traffic 17(8):878–890. https://doi.org/10.1111/tra.12410
Brandstaetter H, Kishi-Itakura C, Tumbarello DA, Manstein DJ, Buss F (2014) Loss of functional MYO1C/myosin 1c, a motor protein involved in lipid raft trafficking, disrupts autophagosome–lysosome fusion. Autophagy 10(12):2310–2323. https://doi.org/10.4161/15548627.2014.984272
Tumbarello DA, Waxse BJ, Arden SD, Bright NA, Kendrick-Jones J, Buss F (2012) Autophagy receptors link myosin VI to autophagosomes to mediate Tom1-dependent autophagosome maturation and fusion with the lysosome. Nat Cell Biol 14(10):1024–1035. https://doi.org/10.1038/ncb2589
Herrmann H, Strelkov SV, Burkhard P, Aebi U (2009) Intermediate filaments: primary determinants of cell architecture and plasticity. J Clin Invest 119(7):1772–1783. https://doi.org/10.1172/JCI38214
Chang L, Goldman RD (2004) Intermediate filaments mediate cytoskeletal crosstalk. Nat Rev Mol Cell Biol 5(8):601–613. https://doi.org/10.1038/nrm1438
Huber F, Boire A, Lopez MP, Koenderink GH (2015) Cytoskeletal crosstalk: when three different personalities team up. Curr Opin Cell Biol 32:39–47. https://doi.org/10.1016/j.ceb.2014.10.005
Wickstead B, Gull K (2011) The evolution of the cytoskeleton. J Cell Biol 194(4):513–525. https://doi.org/10.1083/jcb.201102065
Wiche G (1989) Plectin: general overview and appraisal of its potential role as a subunit protein of the cytomatrix. Critical Rev Biochem Mol Biol 24(1):41–67. https://doi.org/10.3109/10409238909082551
Svitkina TM, Verkhovsky AB, Borisy GG (1996) Plectin sidearms mediate interaction of intermediate filaments with microtubules and other components of the cytoskeleton. J Cell Biol 135(4):991–1007. https://doi.org/10.1083/jcb.135.4.991
Baek A, Yoon S, Kim J, Baek YM, Park H, Lim D, Chung H, Kim DE (2017) Autophagy and KRT8/keratin 8 protect degeneration of retinal pigment epithelium under oxidative stress. Autophagy 13(2):248–263. https://doi.org/10.1080/15548627.2016.1256932
Eriksson JE, Brautigan DL, Vallee R, Olmsted J, Fujiki H, Goldman RD (1992) Cytoskeletal integrity in interphase cells requires protein phosphatase activity. Proc Natl Acad Sci USA 89(22):11093–11097. https://doi.org/10.1073/pnas.89.22.11093
Goldman RD, Khuon S, Chou YH, Opal P, Steinert PM (1996) The function of intermediate filaments in cell shape and cytoskeletal integrity. J Cell Biol 134(4):971–983. https://doi.org/10.1083/jcb.134.4.971
Yatsunami J, Fujiki H, Suganuma M, Yoshizawa S, Eriksson JE, Olson MO, Goldman RD (1991) Vimentin is hyperphosphorylated in primary human fibroblasts treated with okadaic acid. Biochem Biophys Res Commun 177(3):1165–1170. https://doi.org/10.1016/0006-291x(91)90662-q
Tao GZ, Toivola DM, Zhou Q, Strnad P, Xu B, Michie SA, Omary MB (2006) Protein phosphatase-2A associates with and dephosphorylates keratin 8 after hyposmotic stress in a site- and cell-specific manner. J Cell Sci 119(Pt 7):1425–1432. https://doi.org/10.1242/jcs.02861
Wiche G, Winter L (2011) Plectin isoforms as organizers of intermediate filament cytoarchitecture. BioArchitecture 1(1):14–20. https://doi.org/10.4161/bioa.1.1.14630
Moch M, Windoffer R, Schwarz N, Pohl R, Omenzetter A, Schnakenberg U, Herb F, Chaisaowong K, Merhof D, Ramms L et al (2016) Effects of plectin depletion on keratin network dynamics and organization. PLoS ONE 11(3):e0149106. https://doi.org/10.1371/journal.pone.0149106
Casella JF, Flanagan MD, Lin S (1981) Cytochalasin D inhibits actin polymerization and induces depolymerization of actin filaments formed during platelet shape change. Nature 293(5830):302–305. https://doi.org/10.1038/293302a0
Schliwa M (1982) Action of cytochalasin D on cytoskeletal networks. J Cell Biol 92(1):79–91. https://doi.org/10.1083/jcb.92.1.79
Hammer JA 3rd, Sellers JR (2011) Walking to work: roles for class V myosins as cargo transporters. Nat Rev Mol Cell Biol 13(1):13–26. https://doi.org/10.1038/nrm3248
Kneussel M, Wagner W (2013) Myosin motors at neuronal synapses: drivers of membrane transport and actin dynamics. Nat Rev Neurosci 14(4):233–247. https://doi.org/10.1038/nrn3445
Mizushima N, Klionsky DJ (2007) Protein turnover via autophagy: implications for metabolism. Ann Rev Nutr 27:19–40. https://doi.org/10.1146/annurev.nutr.27.061406.093749
Mitter SK, Song C, Qi X, Mao H, Rao H, Akin D, Lewin A, Grant M, Dunn W Jr, Ding J et al (2014) Dysregulated autophagy in the RPE is associated with increased susceptibility to oxidative stress and AMD. Autophagy 10(11):1989–2005. https://doi.org/10.4161/auto.36184
Wang AL, Lukas TJ, Yuan M, Du N, Tso MO, Neufeld AH (2009) Autophagy and exosomes in the aged retinal pigment epithelium: possible relevance to drusen formation and age-related macular degeneration. PLoS ONE 4(1):e4160. https://doi.org/10.1371/journal.pone.0004160
Yoon SY, Choi JE, Kweon HS, Choe H, Kim SW, Hwang O, Lee H, Lee JY, Kim DH (2008) Okadaic acid increases autophagosomes in rat neurons: implications for Alzheimer’s disease. J Neurosci Res 86(14):3230–3239. https://doi.org/10.1002/jnr.21760
Harada M, Hanada S, Toivola DM, Ghori N, Omary MB (2008) Autophagy activation by rapamycin eliminates mouse Mallory-Denk bodies and blocks their proteasome inhibitor-mediated formation. Hepatology 47(6):2026–2035. https://doi.org/10.1002/hep.22294
Kang GY, Bang JY, Choi AJ, Yoon J, Lee WC, Choi S, Yoon S, Kim HC, Baek JH, Park HS et al (2014) Exosomal proteins in the aqueous humor as novel biomarkers in patients with neovascular age-related macular degeneration. J Proteome Res 13(2):581–595. https://doi.org/10.1021/pr400751k
Fuchs E, Weber K (1994) Intermediate filaments: structure, dynamics, function, and disease. Annu Rev Biochem 63:345–382. https://doi.org/10.1146/annurev.bi.63.070194.002021
Lau AT, Chiu JF (2007) The possible role of cytokeratin 8 in cadmium-induced adaptation and carcinogenesis. Cancer Res 67(5):2107–2113. https://doi.org/10.1158/0008-5472.CAN-06-3771
Snider NT, Omary MB (2014) Post-translational modifications of intermediate filament proteins: mechanisms and functions. Nat Rev Mol Cell Biol 15(3):163–177. https://doi.org/10.1038/nrm3753
Tao GZ, Looi KS, Toivola DM, Strnad P, Zhou Q, Liao J, Wei Y, Habtezion A, Omary MB (2009) Keratins modulate the shape and function of hepatocyte mitochondria: a mechanism for protection from apoptosis. J Cell Sci 122(Pt 21):3851–3855. https://doi.org/10.1242/jcs.051862
Caulin C, Ware CF, Magin TM, Oshima RG (2000) Keratin-dependent, epithelial resistance to tumor necrosis factor-induced apoptosis. J Cell Biol 149(1):17–22. https://doi.org/10.1083/jcb.149.1.17
Gilbert S, Loranger A, Daigle N, Marceau N (2001) Simple epithelium keratins 8 and 18 provide resistance to Fas-mediated apoptosis. The protection occurs through a receptor-targeting modulation. J Cell Biol 154(4):763–773. https://doi.org/10.1083/jcb.200102130
Klionsky DJ, Emr SD (2000) Autophagy as a regulated pathway of cellular degradation. Science 290(5497):1717–1721. https://doi.org/10.1126/science.290.5497.1717
Shintani T, Klionsky DJ (2004) Autophagy in health and disease: a double-edged sword. Science 306(5698):990–995. https://doi.org/10.1126/science.1099993
Xie R, Nguyen S, McKeehan K, Wang F, McKeehan WL, Liu L (2011) Microtubule-associated protein 1S (MAP1S) bridges autophagic components with microtubules and mitochondria to affect autophagosomal biogenesis and degradation. J Biol Chem 286(12):10367–10377. https://doi.org/10.1074/jbc.M110.206532
Moreau K, Ravikumar B, Renna M, Puri C, Rubinsztein DC (2011) Autophagosome precursor maturation requires homotypic fusion. Cell 146(2):303–317. https://doi.org/10.1016/j.cell.2011.06.023
Korolchuk VI, Saiki S, Lichtenberg M, Siddiqi FH, Roberts EA, Imarisio S, Jahreiss L, Sarkar S, Futter M, Menzies FM et al (2011) Lysosomal positioning coordinates cellular nutrient responses. Nat Cell Biol 13(4):453–460. https://doi.org/10.1038/ncb2204
Pous C, Codogno P (2011) Lysosome positioning coordinates mTORC1 activity and autophagy. Nat Cell Biol 13(4):342–344. https://doi.org/10.1038/ncb0411-342
Fass E, Shvets E, Degani I, Hirschberg K, Elazar Z (2006) Microtubules support production of starvation-induced autophagosomes but not their targeting and fusion with lysosomes. J Biol Chem 281(47):36303–36316. https://doi.org/10.1074/jbc.M607031200
Scheel J, Matteoni R, Ludwig T, Hoflack B, Kreis TE (1990) Microtubule depolymerization inhibits transport of cathepsin D from the Golgi apparatus to lysosomes. J Cell Sci 96(Pt 4):711–720
Veeranna YDS, Lee JH, Vinod KY, Stavrides P, Amin ND, Pant HC, Nixon RA (2011) Declining phosphatases underlie aging-related hyperphosphorylation of neurofilaments. Neurobiol Aging 32(11):2016–2029. https://doi.org/10.1016/j.neurobiolaging.2009.12.001
Sontag E, Luangpirom A, Hladik C, Mudrak I, Ogris E, Speciale S, White CL 3rd (2004) Altered expression levels of the protein phosphatase 2A ABalphaC enzyme are associated with Alzheimer disease pathology. J Neuropathol Exp Neurol 63(4):287–301. https://doi.org/10.1093/jnen/63.4.287
Strnad P, Windoffer R, Leube RE (2001) In vivo detection of cytokeratin filament network breakdown in cells treated with the phosphatase inhibitor okadaic acid. Cell Tissue Res 306(2):277–293. https://doi.org/10.1007/s004410100455
Liu R, Wang JZ (2009) Protein phosphatase 2A in Alzheimer’s disease. Pathophysiology 16(4):273–277. https://doi.org/10.1016/j.pathophys.2009.02.008
Sontag JM, Sontag E (2014) Protein phosphatase 2A dysfunction in Alzheimer’s disease. Front Mol Neurosci 7:16. https://doi.org/10.3389/fnmol.2014.00016
Samari HR, Moller MT, Holden L, Asmyhr T, Seglen PO (2005) Stimulation of hepatocytic AMP-activated protein kinase by okadaic acid and other autophagy-suppressive toxins. Biochem J 386(Pt 2):237–244. https://doi.org/10.1042/BJ20040609
Cullis DN, Philip B, Baleja JD, Feig LA (2002) Rab11-FIP2, an adaptor protein connecting cellular components involved in internalization and recycling of epidermal growth factor receptors. J Biol Chem 277(51):49158–49166. https://doi.org/10.1074/jbc.M206316200
Funding
This research was supported by the grant from the National Research Foundation of Korea (2017R1E1A1A01074656) funded by the Korean government.
Author information
Authors and Affiliations
Contributions
D-EK and AB formulated the hypothesis, and initiated and organized the study. SS and AB designed and performed experiments with data analysis. SS, AB, JHL and D-EK wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared that no competing interests exist.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Son, S., Baek, A., Lee, J.H. et al. Autophagosome–lysosome fusion is facilitated by plectin-stabilized actin and keratin 8 during macroautophagic process. Cell. Mol. Life Sci. 79, 95 (2022). https://doi.org/10.1007/s00018-022-04144-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04144-1