Abstract
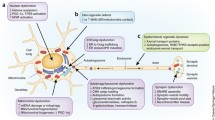
Parthanatos is a form of regulated cell death involved in the pathogenesis of many diseases, particularly neurodegenerative disorders, such as Parkinson's disease, Alzheimer's disease, Huntington’s disease, and amyotrophic lateral sclerosis. Parthanatos is a multistep cell death pathway cascade that involves poly (ADP-ribose) polymerase 1 (PARP-1) overactivation, PAR accumulation, PAR binding to apoptosis-inducing factor (AIF), AIF release from the mitochondria, nuclear translocation of the AIF/macrophage migration inhibitory factor (MIF) complex, and MIF-mediated large-scale DNA fragmentation. All the key players in the parthanatos pathway are pleiotropic proteins with diverse functions. An in-depth understanding of the structure-based activity of the key factors, and the biochemical mechanisms of parthanatos, is crucial for the development of drugs and therapeutic strategies. In this review, we delve into the key players of the parthanatos pathway and reveal the multiple levels of therapeutic opportunities for treating parthanatos-based pathogenesis.




Similar content being viewed by others
Availability of data and material
Not applicable.
References
Wang X, Ge P (2020) Parthanatos in the pathogenesis of nervous system diseases. Neuroscience 449:241–250. https://doi.org/10.1016/j.neuroscience.2020.09.049
Yu SW, Wang H, Poitras MF, Coombs C, Bowers WJ, Federoff HJ, Poirier GG, Dawson TM, Dawson VL (2002) Mediation of poly(ADP-ribose) polymerase-1-dependent cell death by apoptosis-inducing factor. Science 297(5579):259–263. https://doi.org/10.1126/science.1072221
Yu SW, Andrabi SA, Wang H, Kim NS, Poirier GG, Dawson TM, Dawson VL (2006) Apoptosis-inducing factor mediates poly(ADP-ribose) (PAR) polymer-induced cell death. Proc Natl Acad Sci U S A 103(48):18314–18319. https://doi.org/10.1073/pnas.0606528103
Andrabi SA, Kim NS, Yu SW, Wang H, Koh DW, Sasaki M, Klaus JA, Otsuka T, Zhang Z, Koehler RC et al (2006) Poly(ADP-ribose) (PAR) polymer is a death signal. Proc Natl Acad Sci U S A 103(48):18308–18313. https://doi.org/10.1073/pnas.0606526103
Galluzzi L, Vitale I, Abrams JM, Alnemri ES, Baehrecke EH, Blagosklonny MV, Dawson TM, Dawson VL, El-Deiry WS, Fulda S et al (2012) Molecular definitions of cell death subroutines: recommendations of the nomenclature committee on cell death 2012. Cell Death Differ 19(1):107–120. https://doi.org/10.1038/cdd.2011.96
Galluzzi L, Vitale I, Aaronson SA, Abrams JM, Adam D, Agostinis P, Alnemri ES, Altucci L, Amelio I, Andrews DW et al (2018) Molecular mechanisms of cell death: recommendations of the nomenclature committee on cell death 2018. Cell Death Differ 25(3):486–541. https://doi.org/10.1038/s41418-017-0012-4
Fatokun AA, Dawson VL, Dawson TM (2014) Parthanatos: mitochondrial-linked mechanisms and therapeutic opportunities. Br J Pharmacol 171(8):2000–2016. https://doi.org/10.1111/bph.12416
Pacher P, Szabo C (2008) Role of the peroxynitrite-poly(ADP-ribose) polymerase pathway in human disease. Am J Pathol 173(1):2–13. https://doi.org/10.2353/ajpath.2008.080019
Kam TI, Mao X, Park H, Chou SC, Karuppagounder SS, Umanah GE, Yun SP, Brahmachari S, Panicker N, Chen R et al (2018) Poly(ADP-ribose) drives pathologic alpha-synuclein neurodegeneration in Parkinson’s disease. Science. https://doi.org/10.1126/science.aat8407
Wang Y, An R, Umanah GK, Park H, Nambiar K, Eacker SM, Kim B, Bao L, Harraz MM, Chang C et al (2016) A nuclease that mediates cell death induced by DNA damage and poly(ADP-ribose) polymerase-1. Science. https://doi.org/10.1126/science.aad6872
David KK, Andrabi SA, Dawson TM, Dawson VL (2009) Parthanatos, a messenger of death. Front Biosci (Landmark Ed) 14:1116–1128. https://doi.org/10.2741/3297
Andrabi SA, Dawson TM, Dawson VL (2008) Mitochondrial and nuclear cross talk in cell death: parthanatos. Ann N Y Acad Sci 1147:233–241. https://doi.org/10.1196/annals.1427.014
Park H, Kam TI, Dawson TM, Dawson VL (2020) Poly (ADP-ribose) (PAR)-dependent cell death in neurodegenerative diseases. Int Rev Cell Mol Biol 353:1–29. https://doi.org/10.1016/bs.ircmb.2019.12.009
Wang H, Yu SW, Koh DW, Lew J, Coombs C, Bowers W, Federoff HJ, Poirier GG, Dawson TM, Dawson VL (2004) Apoptosis-inducing factor substitutes for caspase executioners in NMDA-triggered excitotoxic neuronal death. J Neurosci 24(48):10963–10973. https://doi.org/10.1523/JNEUROSCI.3461-04.2004
Wang R, Li C, Qiao P, Xue Y, Zheng X, Chen H, Zeng X, Liu W, Boldogh I, Ba X (2018) OGG1-initiated base excision repair exacerbates oxidative stress-induced parthanatos. Cell Death Dis 9(6):628. https://doi.org/10.1038/s41419-018-0680-0
Mashimo M, Kato J, Moss J (2013) ADP-ribosyl-acceptor hydrolase 3 regulates poly (ADP-ribose) degradation and cell death during oxidative stress. Proc Natl Acad Sci U S A 110(47):18964–18969. https://doi.org/10.1073/pnas.1312783110
Fan J, Dawson TM, Dawson VL (2017) Cell death mechanisms of neurodegeneration. Adv Neurobiol 15:403–425. https://doi.org/10.1007/978-3-319-57193-5_16
Jang KH, Do YJ, Son D, Son E, Choi JS, Kim E (2017) AIF-independent parthanatos in the pathogenesis of dry age-related macular degeneration. Cell Death Dis 8(1):e2526. https://doi.org/10.1038/cddis.2016.437
Regdon Z, Robaszkiewicz A, Kovacs K, Rygielska Z, Hegedus C, Bodoor K, Szabo E, Virag L (2019) LPS protects macrophages from AIF-independent parthanatos by downregulation of PARP1 expression, induction of SOD2 expression, and a metabolic shift to aerobic glycolysis. Free Radic Biol Med 131:184–196. https://doi.org/10.1016/j.freeradbiomed.2018.11.034
Erdelyi K, Bai P, Kovacs I, Szabo E, Mocsar G, Kakuk A, Szabo C, Gergely P, Virag L (2009) Dual role of poly(ADP-ribose) glycohydrolase in the regulation of cell death in oxidatively stressed A549 cells. FASEB J 23(10):3553–3563. https://doi.org/10.1096/fj.09-133264
Robaszkiewicz A, Erdelyi K, Kovacs K, Kovacs I, Bai P, Rajnavolgyi E, Virag L (2012) Hydrogen peroxide-induced poly(ADP-ribosyl)ation regulates osteogenic differentiation-associated cell death. Free Radic Biol Med 53(8):1552–1564. https://doi.org/10.1016/j.freeradbiomed.2012.08.567
Virag L, Salzman AL, Szabo C (1998) Poly(ADP-ribose) synthetase activation mediates mitochondrial injury during oxidant-induced cell death. J Immunol 161(7):3753–3759
Virag L, Scott GS, Cuzzocrea S, Marmer D, Salzman AL, Szabo C (1998) Peroxynitrite-induced thymocyte apoptosis: the role of caspases and poly (ADP-ribose) synthetase (PARS) activation. Immunology 94(3):345–355. https://doi.org/10.1046/j.1365-2567.1998.00534.x
Ha HC, Snyder SH (1999) Poly(ADP-ribose) polymerase is a mediator of necrotic cell death by ATP depletion. Proc Natl Acad Sci U S A 96(24):13978–13982. https://doi.org/10.1073/pnas.96.24.13978
Ba X, Garg NJ (2011) Signaling mechanism of poly(ADP-ribose) polymerase-1 (PARP-1) in inflammatory diseases. Am J Pathol 178(3):946–955. https://doi.org/10.1016/j.ajpath.2010.12.004
Belenky P, Bogan KL, Brenner C (2007) NAD+ metabolism in health and disease. Trends Biochem Sci 32(1):12–19. https://doi.org/10.1016/j.tibs.2006.11.006
Andrabi SA, Umanah GK, Chang C, Stevens DA, Karuppagounder SS, Gagne JP, Poirier GG, Dawson VL, Dawson TM (2014) Poly(ADP-ribose) polymerase-dependent energy depletion occurs through inhibition of glycolysis. Proc Natl Acad Sci U S A 111(28):10209–10214. https://doi.org/10.1073/pnas.1405158111
Fouquerel E, Goellner EM, Yu Z, Gagne JP, Barbi de Moura M, Feinstein T, Wheeler D, Redpath P, Li J, Romero G et al (2014) ARTD1/PARP1 negatively regulates glycolysis by inhibiting hexokinase 1 independent of NAD+ depletion. Cell Rep 8(6):1819–1831. https://doi.org/10.1016/j.celrep.2014.08.036
Luo X, Kraus WL (2012) On PAR with PARP: cellular stress signaling through poly(ADP-ribose) and PARP-1. Genes Dev 26(5):417–432. https://doi.org/10.1101/gad.183509.111
D’Amours D, Desnoyers S, D’Silva I, Poirier GG (1999) Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem J 342(Pt 2):249–268
Krishnakumar R, Kraus WL (2010) The PARP side of the nucleus: molecular actions, physiological outcomes, and clinical targets. Mol Cell 39(1):8–24. https://doi.org/10.1016/j.molcel.2010.06.017
Gibson BA, Kraus WL (2017) Identification of protein substrates of specific PARP enzymes using analog-sensitive PARP mutants and a “clickable” NAD(+) analog. Methods Mol Biol 1608:111–135. https://doi.org/10.1007/978-1-4939-6993-7_9
Li M, Lu LY, Yang CY, Wang S, Yu X (2013) The FHA and BRCT domains recognize ADP-ribosylation during DNA damage response. Genes Dev 27(16):1752–1768. https://doi.org/10.1101/gad.226357.113
Wang Y, Kim NS, Haince JF, Kang HC, David KK, Andrabi SA, Poirier GG, Dawson VL, Dawson TM (2011) Poly(ADP-ribose) (PAR) binding to apoptosis-inducing factor is critical for PAR polymerase-1-dependent cell death (parthanatos). Sci Signal 4(167):ra20. https://doi.org/10.1126/scisignal.2000902
Wang Y, Luo W, Wang Y (2019) PARP-1 and its associated nucleases in DNA damage response. DNA Repair (Amst) 81:102651. https://doi.org/10.1016/j.dnarep.2019.102651
Lee Y, Karuppagounder SS, Shin JH, Lee YI, Ko HS, Swing D, Jiang H, Kang SU, Lee BD, Kang HC et al (2013) Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss. Nat Neurosci 16(10):1392–1400. https://doi.org/10.1038/nn.3500
Visochek L, Grigoryan G, Kalal A, Milshtein-Parush H, Gazit N, Slutsky I, Yeheskel A, Shainberg A, Castiel A, Seger R et al (2016) A PARP1-ERK2 synergism is required for the induction of LTP. Sci Rep 6:24950. https://doi.org/10.1038/srep24950
Gibson BA, Kraus WL (2012) New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol 13(7):411–424. https://doi.org/10.1038/nrm3376
Homburg S, Visochek L, Moran N, Dantzer F, Priel E, Asculai E, Schwartz D, Rotter V, Dekel N, Cohen-Armon M (2000) A fast signal-induced activation of Poly(ADP-ribose) polymerase: a novel downstream target of phospholipase c. J Cell Biol 150(2):293–307. https://doi.org/10.1083/jcb.150.2.293
Ju BG, Solum D, Song EJ, Lee KJ, Rose DW, Glass CK, Rosenfeld MG (2004) Activating the PARP-1 sensor component of the groucho/ TLE1 corepressor complex mediates a CaMKinase IIdelta-dependent neurogenic gene activation pathway. Cell 119(6):815–829. https://doi.org/10.1016/j.cell.2004.11.017
Cohen-Armon M, Visochek L, Rozensal D, Kalal A, Geistrikh I, Klein R, Bendetz-Nezer S, Yao Z, Seger R (2007) DNA-independent PARP-1 activation by phosphorylated ERK2 increases Elk1 activity: a link to histone acetylation. Mol Cell 25(2):297–308. https://doi.org/10.1016/j.molcel.2006.12.012
Teloni F, Altmeyer M (2016) Readers of poly(ADP-ribose): designed to be fit for purpose. Nucleic Acids Res 44(3):993–1006. https://doi.org/10.1093/nar/gkv1383
Cohen MS, Chang P (2018) Insights into the biogenesis, function, and regulation of ADP-ribosylation. Nat Chem Biol 14(3):236–243. https://doi.org/10.1038/nchembio.2568
Kamaletdinova T, Fanaei-Kahrani Z, Wang ZQ (2019) The enigmatic function of PARP1: from PARylation activity to PAR readers. Cells. https://doi.org/10.3390/cells8121625
Leung AK (2014) Poly(ADP-ribose): an organizer of cellular architecture. J Cell Biol 205(5):613–619. https://doi.org/10.1083/jcb.201402114
Kunze FA, Hottiger MO (2019) Regulating immunity via ADP-ribosylation: therapeutic implications and beyond. Trends Immunol 40(2):159–173. https://doi.org/10.1016/j.it.2018.12.006
Hassa PO, Haenni SS, Elser M, Hottiger MO (2006) Nuclear ADP-ribosylation reactions in mammalian cells: where are we today and where are we going? Microbiol Mol Biol Rev 70(3):789–829. https://doi.org/10.1128/MMBR.00040-05
Liu L, Ke Y, Jiang X, He F, Pan L, Xu L, Zeng X, Ba X (2012) Lipopolysaccharide activates ERK-PARP-1-RelA pathway and promotes nuclear factor-kappaB transcription in murine macrophages. Hum Immunol 73(5):439–447. https://doi.org/10.1016/j.humimm.2012.02.002
Ke Y, Han Y, Guo X, Wen J, Wang K, Jiang X, Tian X, Ba X, Boldogh I, Zeng X (2017) PARP1 promotes gene expression at the post-transcriptiona level by modulating the RNA-binding protein HuR. Nat Commun 8:14632. https://doi.org/10.1038/ncomms14632
Ray Chaudhuri A, Nussenzweig A (2017) The multifaceted roles of PARP1 in DNA repair and chromatin remodelling. Nat Rev Mol Cell Biol 18(10):610–621. https://doi.org/10.1038/nrm.2017.53
Zhou Y, Feng X, Koh DW (2011) Activation of cell death mediated by apoptosis-inducing factor due to the absence of poly(ADP-ribose) glycohydrolase. Biochemistry 50(14):2850–2859. https://doi.org/10.1021/bi101829r
Koh DW, Lawler AM, Poitras MF, Sasaki M, Wattler S, Nehls MC, Stoger T, Poirier GG, Dawson VL, Dawson TM (2004) Failure to degrade poly(ADP-ribose) causes increased sensitivity to cytotoxicity and early embryonic lethality. Proc Natl Acad Sci U S A 101(51):17699–17704. https://doi.org/10.1073/pnas.0406182101
Wang Y, Dawson VL, Dawson TM (2009) Poly(ADP-ribose) signals to mitochondrial AIF: a key event in parthanatos. Exp Neurol 218(2):193–202. https://doi.org/10.1016/j.expneurol.2009.03.020
Harrision D, Gravells P, Thompson R, Bryant HE (2020) Poly(ADP-ribose) glycohydrolase (PARG) vs. poly(ADP-ribose) polymerase (PARP) - function in genome maintenance and relevance of inhibitors for anti-cancer therapy. Front Mol Biosci 7:191. https://doi.org/10.3389/fmolb.2020.00191
Min W, Wang ZQ (2009) Poly (ADP-ribose) glycohydrolase (PARG) and its therapeutic potential. Front Biosci (Landmark Ed) 14:1619–1626. https://doi.org/10.2741/3329
Cozzi A, Cipriani G, Fossati S, Faraco G, Formentini L, Min W, Cortes U, Wang ZQ, Moroni F, Chiarugi A (2006) Poly(ADP-ribose) accumulation and enhancement of postischemic brain damage in 110-kDa poly(ADP-ribose) glycohydrolase null mice. J Cereb Blood Flow Metab 26(5):684–695. https://doi.org/10.1038/sj.jcbfm.9600222
Oka S, Kato J, Moss J (2006) Identification and characterization of a mammalian 39-kDa poly(ADP-ribose) glycohydrolase. J Biol Chem 281(2):705–713. https://doi.org/10.1074/jbc.M510290200
Mashimo M, Bu X, Aoyama K, Kato J, Ishiwata-Endo H, Stevens LA, Kasamatsu A, Wolfe LA, Toro C, Adams D et al (2019) PARP1 inhibition alleviates injury in ARH3-deficient mice and human cells. JCI Insight. https://doi.org/10.1172/jci.insight.124519
Susin SA, Zamzami N, Castedo M, Hirsch T, Marchetti P, Macho A, Daugas E, Geuskens M, Kroemer G (1996) Bcl-2 inhibits the mitochondrial release of an apoptogenic protease. J Exp Med 184(4):1331–1341. https://doi.org/10.1084/jem.184.4.1331
Otera H, Ohsakaya S, Nagaura Z, Ishihara N, Mihara K (2005) Export of mitochondrial AIF in response to proapoptotic stimuli depends on processing at the intermembrane space. EMBO J 24(7):1375–1386. https://doi.org/10.1038/sj.emboj.7600614
Polster BM, Basanez G, Etxebarria A, Hardwick JM, Nicholls DG (2005) Calpain I induces cleavage and release of apoptosis-inducing factor from isolated mitochondria. J Biol Chem 280(8):6447–6454. https://doi.org/10.1074/jbc.M413269200
Yu SW, Wang Y, Frydenlund DS, Ottersen OP, Dawson VL, Dawson TM (2009) Outer mitochondrial membrane localization of apoptosis-inducing factor: mechanistic implications for release. ASN Neuro. https://doi.org/10.1042/AN20090046
Mate MJ, Ortiz-Lombardia M, Boitel B, Haouz A, Tello D, Susin SA, Penninger J, Kroemer G, Alzari PM (2002) The crystal structure of the mouse apoptosis-inducing factor AIF. Nat Struct Biol 9(6):442–446. https://doi.org/10.1038/nsb793
Churbanova IY, Sevrioukova IF (2008) Redox-dependent changes in molecular properties of mitochondrial apoptosis-inducing factor. J Biol Chem 283(9):5622–5631. https://doi.org/10.1074/jbc.M709147200
Sevrioukova IF (2009) Redox-linked conformational dynamics in apoptosis-inducing factor. J Mol Biol 390(5):924–938. https://doi.org/10.1016/j.jmb.2009.05.013
Joza N, Oudit GY, Brown D, Benit P, Kassiri Z, Vahsen N, Benoit L, Patel MM, Nowikovsky K, Vassault A et al (2005) Muscle-specific loss of apoptosis-inducing factor leads to mitochondrial dysfunction, skeletal muscle atrophy, and dilated cardiomyopathy. Mol Cell Biol 25(23):10261–10272. https://doi.org/10.1128/MCB.25.23.10261-10272.2005
Cheung EC, Joza N, Steenaart NA, McClellan KA, Neuspiel M, McNamara S, MacLaurin JG, Rippstein P, Park DS, Shore GC et al (2006) Dissociating the dual roles of apoptosis-inducing factor in maintaining mitochondrial structure and apoptosis. EMBO J 25(17):4061–4073. https://doi.org/10.1038/sj.emboj.7601276
Klein JA, Longo-Guess CM, Rossmann MP, Seburn KL, Hurd RE, Frankel WN, Bronson RT, Ackerman SL (2002) The harlequin mouse mutation downregulates apoptosis-inducing factor. Nature 419(6905):367–374. https://doi.org/10.1038/nature01034
Lipton SA, Bossy-Wetzel E (2002) Dueling activities of AIF in cell death versus survival: DNA binding and redox activity. Cell 111(2):147–150. https://doi.org/10.1016/s0092-8674(02)01046-2
Moubarak RS, Yuste VJ, Artus C, Bouharrour A, Greer PA, Menissier-de Murcia J, Susin SA (2007) Sequential activation of poly(ADP-ribose) polymerase 1, calpains, and Bax is essential in apoptosis-inducing factor-mediated programmed necrosis. Mol Cell Biol 27(13):4844–4862. https://doi.org/10.1128/MCB.02141-06
Cabon L, Galan-Malo P, Bouharrour A, Delavallee L, Brunelle-Navas MN, Lorenzo HK, Gross A, Susin SA (2012) BID regulates AIF-mediated caspase-independent necroptosis by promoting BAX activation. Cell Death Differ 19(2):245–256. https://doi.org/10.1038/cdd.2011.91
Culmsee C, Zhu C, Landshamer S, Becattini B, Wagner E, Pellecchia M, Blomgren K, Plesnila N (2005) Apoptosis-inducing factor triggered by poly(ADP-ribose) polymerase and bid mediates neuronal cell death after oxygen-glucose deprivation and focal cerebral ischemia. J Neurosci 25(44):10262–10272. https://doi.org/10.1523/JNEUROSCI.2818-05.2005
Cheung EC, Melanson-Drapeau L, Cregan SP, Vanderluit JL, Ferguson KL, McIntosh WC, Park DS, Bennett SA, Slack RS (2005) Apoptosis-inducing factor is a key factor in neuronal cell death propagated by BAX-dependent and BAX-independent mechanisms. J Neurosci 25(6):1324–1334. https://doi.org/10.1523/JNEUROSCI.4261-04.2005
Daugas E, Susin SA, Zamzami N, Ferri KF, Irinopoulou T, Larochette N, Prevost MC, Leber B, Andrews D, Penninger J et al (2000) Mitochondrio-nuclear translocation of AIF in apoptosis and necrosis. FASEB J 14(5):729–739
Ame JC, Spenlehauer C, de Murcia G (2004) The PARP superfamily. BioEssays 26(8):882–893. https://doi.org/10.1002/bies.20085
Mashimo M, Onishi M, Uno A, Tanimichi A, Nobeyama A, Mori M, Yamada S, Negi S, Bu X, Kato J et al (2021) The 89-kDa PARP1 cleavage fragment serves as a cytoplasmic PAR carrier to induce AIF-mediated apoptosis. J Biol Chem 296:100046. https://doi.org/10.1074/jbc.RA120.014479
Mashimo M, Moss J (2016) Functional role of ADP-ribosyl-acceptor hydrolase 3 in poly(ADP-ribose) polymerase-1 response to oxidative stress. Curr Protein Pept Sci 17(7):633–640. https://doi.org/10.2174/1389203717666160419144603
Wang X, Yang C, Chai J, Shi Y, Xue D (2002) Mechanisms of AIF-mediated apoptotic DNA degradation in Caenorhabditis elegans. Science 298(5598):1587–1592. https://doi.org/10.1126/science.1076194
Xu Z, Zhang J, David KK, Yang ZJ, Li X, Dawson TM, Dawson VL, Koehler RC (2010) Endonuclease G does not play an obligatory role in poly(ADP-ribose) polymerase-dependent cell death after transient focal cerebral ischemia. Am J Physiol Regul Integr Comp Physiol 299(1):R215-221. https://doi.org/10.1152/ajpregu.00747.2009
Cande C, Vahsen N, Kouranti I, Schmitt E, Daugas E, Spahr C, Luban J, Kroemer RT, Giordanetto F, Garrido C et al (2004) AIF and cyclophilin A cooperate in apoptosis-associated chromatinolysis. Oncogene 23(8):1514–1521. https://doi.org/10.1038/sj.onc.1207279
Zhu C, Wang X, Deinum J, Huang Z, Gao J, Modjtahedi N, Neagu MR, Nilsson M, Eriksson PS, Hagberg H et al (2007) Cyclophilin A participates in the nuclear translocation of apoptosis-inducing factor in neurons after cerebral hypoxia-ischemia. J Exp Med 204(8):1741–1748. https://doi.org/10.1084/jem.20070193
Artus C, Boujrad H, Bouharrour A, Brunelle MN, Hoos S, Yuste VJ, Lenormand P, Rousselle JC, Namane A, England P et al (2010) AIF promotes chromatinolysis and caspase-independent programmed necrosis by interacting with histone H2AX. EMBO J 29(9):1585–1599. https://doi.org/10.1038/emboj.2010.43
Baritaud M, Boujrad H, Lorenzo HK, Krantic S, Susin SA (2010) Histone H2AX: the missing link in AIF-mediated caspase-independent programmed necrosis. Cell Cycle 9(16):3166–3173. https://doi.org/10.4161/cc.9.16.12887
Doti N, Reuther C, Scognamiglio PL, Dolga AM, Plesnila N, Ruvo M, Culmsee C (2014) Inhibition of the AIF/CypA complex protects against intrinsic death pathways induced by oxidative stress. Cell Death Dis 5:e993. https://doi.org/10.1038/cddis.2013.518
Rodriguez J, Xie C, Li T, Sun Y, Wang Y, Xu Y, Li K, Zhang S, Zhou K, Wang Y et al (2020) Inhibiting the interaction between apoptosis-inducing factor and cyclophilin A prevents brain injury in neonatal mice after hypoxia-ischemia. Neuropharmacology 171:108088. https://doi.org/10.1016/j.neuropharm.2020.108088
Bloom BR, Bennett B (1966) Mechanism of a reaction in vitro associated with delayed-type hypersensitivity. Science 153(3731):80–82. https://doi.org/10.1126/science.153.3731.80
Bloom J, Sun S, Al-Abed Y (2016) MIF, a controversial cytokine: a review of structural features, challenges, and opportunities for drug development. Expert Opin Ther Targets 20(12):1463–1475. https://doi.org/10.1080/14728222.2016.1251582
Bernhagen J, Mitchell RA, Calandra T, Voelter W, Cerami A, Bucala R (1994) Purification, bioactivity, and secondary structure analysis of mouse and human macrophage migration inhibitory factor (MIF). Biochemistry 33(47):14144–14155. https://doi.org/10.1021/bi00251a025
Florez-Sampedro L, Soto-Gamez A, Poelarends GJ, Melgert BN (2020) The role of MIF in chronic lung diseases: looking beyond inflammation. Am J Physiol Lung Cell Mol Physiol 318(6):L1183–L1197. https://doi.org/10.1152/ajplung.00521.2019
Calandra T, Roger T (2003) Macrophage migration inhibitory factor: a regulator of innate immunity. Nat Rev Immunol 3(10):791–800. https://doi.org/10.1038/nri1200
Onodera S, Nishihira J, Koyama Y, Majima T, Aoki Y, Ichiyama H, Ishibashi T, Minami A (2004) Macrophage migration inhibitory factor up-regulates the expression of interleukin-8 messenger RNA in synovial fibroblasts of rheumatoid arthritis patients: common transcriptional regulatory mechanism between interleukin-8 and interleukin-1beta. Arthritis Rheum 50(5):1437–1447. https://doi.org/10.1002/art.20190
Schindler L, Dickerhof N, Hampton MB, Bernhagen J (2018) Post-translational regulation of macrophage migration inhibitory factor: basis for functional fine-tuning. Redox Biol 15:135–142. https://doi.org/10.1016/j.redox.2017.11.028
Tarnowski M, Grymula K, Liu R, Tarnowska J, Drukala J, Ratajczak J, Mitchell RA, Ratajczak MZ, Kucia M (2010) Macrophage migration inhibitory factor is secreted by rhabdomyosarcoma cells, modulates tumor metastasis by binding to CXCR4 and CXCR7 receptors and inhibits recruitment of cancer-associated fibroblasts. Mol Cancer Res 8(10):1328–1343. https://doi.org/10.1158/1541-7786.MCR-10-0288
Bernhagen J, Krohn R, Lue H, Gregory JL, Zernecke A, Koenen RR, Dewor M, Georgiev I, Schober A, Leng L et al (2007) MIF is a noncognate ligand of CXC chemokine receptors in inflammatory and atherogenic cell recruitment. Nat Med 13(5):587–596. https://doi.org/10.1038/nm1567
Leng L, Metz CN, Fang Y, Xu J, Donnelly S, Baugh J, Delohery T, Chen Y, Mitchell RA, Bucala R (2003) MIF signal transduction initiated by binding to CD74. J Exp Med 197(11):1467–1476. https://doi.org/10.1084/jem.20030286
Shi X, Leng L, Wang T, Wang W, Du X, Li J, McDonald C, Chen Z, Murphy JW, Lolis E et al (2006) CD44 is the signaling component of the macrophage migration inhibitory factor-CD74 receptor complex. Immunity 25(4):595–606. https://doi.org/10.1016/j.immuni.2006.08.020
Binsky I, Haran M, Starlets D, Gore Y, Lantner F, Harpaz N, Leng L, Goldenberg DM, Shvidel L, Berrebi A et al (2007) IL-8 secreted in a macrophage migration-inhibitory factor- and CD74-dependent manner regulates B cell chronic lymphocytic leukemia survival. Proc Natl Acad Sci U S A 104(33):13408–13413. https://doi.org/10.1073/pnas.0701553104
Leyton-Jaimes MF, Kahn J, Israelson A (2018) Macrophage migration inhibitory factor: a multifaceted cytokine implicated in multiple neurological diseases. Exp Neurol 301(Pt B):83–91. https://doi.org/10.1016/j.expneurol.2017.06.021
Kleemann R, Kapurniotu A, Frank RW, Gessner A, Mischke R, Flieger O, Juttner S, Brunner H, Bernhagen J (1998) Disulfide analysis reveals a role for macrophage migration inhibitory factor (MIF) as thiol-protein oxidoreductase. J Mol Biol 280(1):85–102. https://doi.org/10.1006/jmbi.1998.1864
Thiele M, Bernhagen J (2005) Link between macrophage migration inhibitory factor and cellular redox regulation. Antioxid Redox Signal 7(9–10):1234–1248. https://doi.org/10.1089/ars.2005.7.1234
Jonas E (2016) The MIFstep in parthanatos. Science 354(6308):36–37. https://doi.org/10.1126/science.aai8756
Kosinski J, Feder M, Bujnicki JM (2005) The PD-(D/E)XK superfamily revisited: identification of new members among proteins involved in DNA metabolism and functional predictions for domains of (hitherto) unknown function. BMC Bioinformatics 6:172. https://doi.org/10.1186/1471-2105-6-172
MacKay C, Declais AC, Lundin C, Agostinho A, Deans AJ, MacArtney TJ, Hofmann K, Gartner A, West SC, Helleday T et al (2010) Identification of KIAA1018/FAN1, a DNA repair nuclease recruited to DNA damage by monoubiquitinated FANCD2. Cell 142(1):65–76. https://doi.org/10.1016/j.cell.2010.06.021
Kratz K, Schopf B, Kaden S, Sendoel A, Eberhard R, Lademann C, Cannavo E, Sartori AA, Hengartner MO, Jiricny J (2010) Deficiency of FANCD2-associated nuclease KIAA1018/FAN1 sensitizes cells to interstrand crosslinking agents. Cell 142(1):77–88. https://doi.org/10.1016/j.cell.2010.06.022
Laganeckas M, Margelevicius M, Venclovas C (2011) Identification of new homologs of PD-(D/E)XK nucleases by support vector machines trained on data derived from profile-profile alignments. Nucleic Acids Res 39(4):1187–1196. https://doi.org/10.1093/nar/gkq958
Steczkiewicz K, Muszewska A, Knizewski L, Rychlewski L, Ginalski K (2012) Sequence, structure and functional diversity of PD-(D/E)XK phosphodiesterase superfamily. Nucleic Acids Res 40(15):7016–7045. https://doi.org/10.1093/nar/gks382
Dawson TM, Dawson VL (2017) Mitochondrial mechanisms of neuronal cell death: potential therapeutics. Annu Rev Pharmacol Toxicol 57:437–454. https://doi.org/10.1146/annurev-pharmtox-010716-105001
McGurk L, Mojsilovic-Petrovic J, Van Deerlin VM, Shorter J, Kalb RG, Lee VM, Trojanowski JQ, Lee EB, Bonini NM (2018) Nuclear poly(ADP-ribose) activity is a therapeutic target in amyotrophic lateral sclerosis. Acta Neuropathol Commun 6(1):84. https://doi.org/10.1186/s40478-018-0586-1
Zhou Y, Liu L, Tao S, Yao Y, Wang Y, Wei Q, Shao A, Deng Y (2021) Parthanatos and its associated components: promising therapeutic targets for cancer. Pharmacol Res 163:105299. https://doi.org/10.1016/j.phrs.2020.105299
Matulonis UA, Harter P, Gourley C, Friedlander M, Vergote I, Rustin G, Scott C, Meier W, Shapira-Frommer R, Safra T et al (2016) Olaparib maintenance therapy in patients with platinum-sensitive, relapsed serous ovarian cancer and a BRCA mutation: overall survival adjusted for postprogression poly(adenosine diphosphate ribose) polymerase inhibitor therapy. Cancer 122(12):1844–1852. https://doi.org/10.1002/cncr.29995
Mirza MR, Monk BJ, Herrstedt J, Oza AM, Mahner S, Redondo A, Fabbro M, Ledermann JA, Lorusso D, Vergote I et al (2016) Niraparib maintenance therapy in platinum-sensitive, recurrent ovarian cancer. N Engl J Med 375(22):2154–2164. https://doi.org/10.1056/NEJMoa1611310
Swisher EM, Lin KK, Oza AM, Scott CL, Giordano H, Sun J, Konecny GE, Coleman RL, Tinker AV, O’Malley DM et al (2017) Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 part 1): an international, multicentre, open-label, phase 2 trial. Lancet Oncol 18(1):75–87. https://doi.org/10.1016/S1470-2045(16)30559-9
Litton JK, Rugo HS, Ettl J, Hurvitz SA, Goncalves A, Lee KH, Fehrenbacher L, Yerushalmi R, Mina LA, Martin M et al (2018) Talazoparib in patients with advanced breast cancer and a germline BRCA mutation. N Engl J Med 379(8):753–763. https://doi.org/10.1056/NEJMoa1802905
Singh N, Pay SL, Bhandare SB, Arimpur U, Motea EA (2020) Therapeutic strategies and biomarkers to modulate PARP activity for targeted cancer therapy. Cancers (Basel). https://doi.org/10.3390/cancers12040972
Beitz JM (2014) Parkinson’s disease: a review. Front Biosci (Schol Ed) 6:65–74. https://doi.org/10.2741/s415
Oh YK, Shin KS, Kang SJ (2006) AIF translocates to the nucleus in the spinal motor neurons in a mouse model of ALS. Neurosci Lett 406(3):205–210. https://doi.org/10.1016/j.neulet.2006.07.044
Shibata N, Kakita A, Takahashi H, Ihara Y, Nobukuni K, Fujimura H, Sakoda S, Sasaki S, Yamamoto T, Kobayashi M (2009) Persistent cleavage and nuclear translocation of apoptosis-inducing factor in motor neurons in the spinal cord of sporadic amyotrophic lateral sclerosis patients. Acta Neuropathol 118(6):755–762. https://doi.org/10.1007/s00401-009-0580-6
Duan Y, Du A, Gu J, Duan G, Wang C, Gui X, Ma Z, Qian B, Deng X, Zhang K et al (2019) PARylation regulates stress granule dynamics, phase separation, and neurotoxicity of disease-related RNA-binding proteins. Cell Res 29(3):233–247. https://doi.org/10.1038/s41422-019-0141-z
Soria Lopez JA, Gonzalez HM, Leger GC (2019) Alzheimer’s disease. Handb Clin Neurol 167:231–255. https://doi.org/10.1016/B978-0-12-804766-8.00013-3
Angelova PR, Abramov AY (2014) Interaction of neurons and astrocytes underlies the mechanism of Abeta-induced neurotoxicity. Biochem Soc Trans 42(5):1286–1290. https://doi.org/10.1042/BST20140153
Abeti R, Abramov AY, Duchen MR (2011) Beta-amyloid activates PARP causing astrocytic metabolic failure and neuronal death. Brain 134(Pt 6):1658–1672. https://doi.org/10.1093/brain/awr104
Kauppinen TM, Suh SW, Higashi Y, Berman AE, Escartin C, Won SJ, Wang C, Cho SH, Gan L, Swanson RA (2011) Poly(ADP-ribose)polymerase-1 modulates microglial responses to amyloid beta. J Neuroinflammation 8:152. https://doi.org/10.1186/1742-2094-8-152
Labbadia J, Morimoto RI (2013) Huntington’s disease: underlying molecular mechanisms and emerging concepts. Trends Biochem Sci 38(8):378–385. https://doi.org/10.1016/j.tibs.2013.05.003
Sun Y, Savanenin A, Reddy PH, Liu YF (2001) Polyglutamine-expanded huntingtin promotes sensitization of N-methyl-D-aspartate receptors via post-synaptic density 95. J Biol Chem 276(27):24713–24718. https://doi.org/10.1074/jbc.M103501200
Cardinale A, Paldino E, Giampa C, Bernardi G, Fusco FR (2015) PARP-1 inhibition is neuroprotective in the R6/2 mouse model of Huntington’s disease. PLoS ONE 10(8):e0134482. https://doi.org/10.1371/journal.pone.0134482
Visnes T, Cazares-Korner A, Hao W, Wallner O, Masuyer G, Loseva O, Mortusewicz O, Wiita E, Sarno A, Manoilov A et al (2018) Small-molecule inhibitor of OGG1 suppresses proinflammatory gene expression and inflammation. Science 362(6416):834–839. https://doi.org/10.1126/science.aar8048
Donley N, Jaruga P, Coskun E, Dizdaroglu M, McCullough AK, Lloyd RS (2015) Small molecule inhibitors of 8-oxoguanine DNA glycosylase-1 (OGG1). ACS Chem Biol 10(10):2334–2343. https://doi.org/10.1021/acschembio.5b00452
Kang HC, Lee YI, Shin JH, Andrabi SA, Chi Z, Gagne JP, Lee Y, Ko HS, Lee BD, Poirier GG et al (2011) Iduna is a poly(ADP-ribose) (PAR)-dependent E3 ubiquitin ligase that regulates DNA damage. Proc Natl Acad Sci U S A 108(34):14103–14108. https://doi.org/10.1073/pnas.1108799108
Andrabi SA, Kang HC, Haince JF, Lee YI, Zhang J, Chi Z, West AB, Koehler RC, Poirier GG, Dawson TM et al (2011) Iduna protects the brain from glutamate excitotoxicity and stroke by interfering with poly(ADP-ribose) polymer-induced cell death. Nat Med 17(6):692–699. https://doi.org/10.1038/nm.2387
Park YH, Seo JH, Park JH, Lee HS, Kim KW (2017) Hsp70 acetylation prevents caspase-dependent/independent apoptosis and autophagic cell death in cancer cells. Int J Oncol 51(2):573–578. https://doi.org/10.3892/ijo.2017.4039
Shelar SB, Kaminska KK, Reddy SA, Kumar D, Tan CT, Yu VC, Lu J, Holmgren A, Hagen T, Chew EH (2015) Thioredoxin-dependent regulation of AIF-mediated DNA damage. Free Radic Biol Med 87:125–136. https://doi.org/10.1016/j.freeradbiomed.2015.06.029
Russo L, Mascanzoni F, Farina B, Dolga AM, Monti A, Caporale A, Culmsee C, Fattorusso R, Ruvo M, Doti N (2021) Design, optimization, and structural characterization of an apoptosis-inducing factor peptide targeting human cyclophilin a to inhibit apoptosis inducing factor-mediated cell death. J Med Chem 64(15):11445–11459. https://doi.org/10.1021/acs.jmedchem.1c00777
Monti A, Sturlese M, Caporale A, Roger JA, Mascanzoni F, Ruvo M, Doti N (1864) Design, synthesis, structural analysis and biochemical studies of stapled AIF(370–394) analogues as ligand of CypA. Biochim Biophys Acta Gen Subj 12:129717. https://doi.org/10.1016/j.bbagen.2020.129717
Wang M, Zhang L, Han X, Yang J, Qian J, Hong S, Samaniego F, Romaguera J, Yi Q (2007) Atiprimod inhibits the growth of mantle cell lymphoma in vitro and in vivo and induces apoptosis via activating the mitochondrial pathways. Blood 109(12):5455–5462. https://doi.org/10.1182/blood-2006-12-063958
Kroemer G, El-Deiry WS, Golstein P, Peter ME, Vaux D, Vandenabeele P, Zhivotovsky B, Blagosklonny MV, Malorni W, Knight RA et al (2005) Classification of cell death: recommendations of the nomenclature committee on cell death. Cell Death Differ 12(Suppl 2):1463–1467. https://doi.org/10.1038/sj.cdd.4401724
Feng X, Zhou Y, Proctor AM, Hopkins MM, Liu M, Koh DW (2012) Silencing of apoptosis-inducing factor and poly(ADP-ribose) glycohydrolase reveals novel roles in breast cancer cell death after chemotherapy. Mol Cancer 11:48. https://doi.org/10.1186/1476-4598-11-48
Funding
This work was supported by grants from the National Natural Science Foundation of China (grant number: 31900557 to R.W., 31970686 to X.B. and 31801182 to Y.K.), China Postdoctoral Science Foundation Grant (grant number: 2019M662431 to R.W.).
Author information
Authors and Affiliations
Contributions
RW and XB had the idea for the article; RW, LL and JL performed the literature search; RW, XB, LL, JL, JG, YK and XZ drafted and critically revised work; RW designed the figures. LL and RW produced the figures.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors consent the publication of the manuscript in CMLS.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, L., Li, J., Ke, Y. et al. The key players of parthanatos: opportunities for targeting multiple levels in the therapy of parthanatos-based pathogenesis. Cell. Mol. Life Sci. 79, 60 (2022). https://doi.org/10.1007/s00018-021-04109-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-021-04109-w