Abstract
Neurodevelopmental disorders (NDDs), including intellectual disability (ID) and autism spectrum disorders (ASD), are a large group of disorders in which early insults during brain development result in a wide and heterogeneous spectrum of clinical diagnoses. Mutations in genes coding for chromatin remodelers are overrepresented in NDD cohorts, pointing towards epigenetics as a convergent pathogenic pathway between these disorders. In this review we detail the role of NDD-associated chromatin remodelers during the developmental continuum of progenitor expansion, differentiation, cell-type specification, migration and maturation. We discuss how defects in chromatin remodelling during these early developmental time points compound over time and result in impaired brain circuit establishment. In particular, we focus on their role in the three largest cell populations: glutamatergic neurons, GABAergic neurons, and glia cells. An in-depth understanding of the spatiotemporal role of chromatin remodelers during neurodevelopment can contribute to the identification of molecular targets for treatment strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A mature brain is the product of its development. Early developmental insults during the assembly of these neuronal circuits can severely impact how a person develops and behaves in their adult life. Across the ongoing developmental continuum of progenitor expansion, differentiation, cell-type specification, migration and maturation, early developmental insults will compound over time, leading to a circuit dysfunction.
Neurodevelopmental disorders (NDDs) pose such an example of disorders where early developmental insults from conception on result in a wide and heterogeneous spectrum of clinical diagnosis’s including intellectual disability (ID), autism spectrum disorder (ASD), attention deficit hyperactivity disorder (ADHD), schizophrenia (SCZ) and mood disorders (bipolar disorder (BD), major depressive disorder (MDD) [1]. These NDDs are often diagnosed during childhood, and overlap between diagnostic categories [2]. NDDs can be caused by both genetic and non-genetic sources. The most frequent non-genetic cause of NDDs is foetal alcohol syndrome disorder [3,4,5]. Additionally, the extreme genetic heterogeneity in NDDs is one of the major limiting factors in both diagnosis and treatment [6]. With the use of sophisticated diagnostic tools such as whole exome sequencing and whole genome sequencing, the number of genes and variants linked to the aetiology of NDDs is vastly increasing [7, 8]. By doing so, chromatin remodelling genes have been found enriched in large datasets of NDD patients, and thereby pointing towards epigenetics as a convergent pathogenic mechanism [8,9,10,11,12,13]. By altering the epigenetic state of genes or histones, chromatin remodelers play an integral part in the machinery that translate external signals into lasting changes in gene expression patterns [14]. Furthermore, chromatin remodelers are multifunctional proteins that can influence various processes across the developmental continuum, including neural progenitor generation and specification, cell-type differentiation and expansion, migration and circuit integration [15]. Hence, as a significant subset of NDDs are caused by a failure of chromatin remodelling, it is to be expected that deficient chromatin remodelling will have a compounding effect across the developmental continuum, ultimately causing circuit dysfunction in mature networks [16].
Several reviews have already focussed on the most recent findings regarding the epigenetic origin of NDDs [17,18,19], however comparatively little is known about the function of these chromatin remodelers during the different programs of progenitor expansion, cell-type specification, neuronal migration and circuit integration. Using examples from mouse models mimicking these NDDs by introducing mutations in chromatin remodelers (also called NDD-related chromatinopathies), this review will first discuss how altered chromatin remodelling affects the different processes of the ongoing developmental continuum from mouse embryonic day (E) 10–17. Here, we will specifically focus on the three most abundant cell types found in the neocortex: excitatory (glutamatergic) and inhibitory (GABAergic) neurons, as well as glia cells. Although various chromatin remodelers have been described to play a role at one of these developmental processes, we chose to elaborate only on specific well-studied examples that play a role at multiple of these developmental steps, stressing their importance in neurodevelopment (Table 1). Furthermore, we will only focus on chromatin remodeler proteins and protein complexes and thereby exclude chromatin remodelling by non-coding RNAs (for a good review the authors would like to refer the readers to [20,21,22]).
Chromatin remodelers
Chromatin was first described by Walther Flemming for the unique fibrous structures observed in cellular nuclei [23]. Chromatin is a highly dynamic structure that regulates the complex organization of the genome and thereby controls the gene expression underneath, and is composed of nucleosomes containing an octamer of histones (i.e. H2A, H2B, H3 and H4), wrapped by 147 base pairs of DNA and the linker histone H1 [24]. The distinction between condensed heterochromatin and open euchromatin structures were first reported by Emil Heitz [25], and can be altered by chromatin remodelers via three distinct mechanisms, including: (i) sliding of an octamer across the DNA (nucleosome sliding), (ii) changing the conformation of nucleosomal DNA, and (iii) altering the composition of the octamers (histone variant exchange). By doing so, chromatin remodelling facilitates downstream gene expression in a cell-type and cellular demand-specific way, stressing their important role during (neuro) development.
Based on their function, three categories of chromatin remodelers have been classified, which are (i) the enzymes that control histone post-translational modifications (PTM) [26], (ii) DNA modifications that can attract/repel chromatin remodelling proteins or complexes, or (iii) enzymes that alter histone-DNA contact within the nucleosome via ATP hydrolysis [27]. Furthermore, 3D genome architecture is increasingly considered an important epigenetic regulator of gene expression [28]. In the next section, we will briefly discuss the global function of these epigenetic regulators (Fig. 1).
Left panel: The genome is organised into euchromatic (accessible) or heterochromatic (inaccessible) chromosome territories. Within chromosome territories, large chromatin domains are organised into smaller and smaller sub-domains known as topologically associated domains (TADs). TADs are regions where DNA is highly organised in 3D space to enable long-range transcriptional regulation between non-linearly neighbouring strands. Right panel: DNA is wrapped around an octamer of histones, of which its accessibility is regulated by histone modifications like methylation (Me), Acetylation (Ac) or Ubiquitination (Ub) or by histone modifying enzymes. Finally, transcription can be regulated by direct DNA modifications, such as DNA methylation
Histone modifying enzymes
Over the last decade a major effort was put into the identification of enzymes that directly modify histones. So far, enzymes have been identified for methylation [29], acetylation [30], phosphorylation [31], ubiquitination [32], sumoylation [33], biotinylation [34], ADP-ribosylation [35], deamination [36, 37], proline isomerization [38], β-N-glycosylation [39], crotonylation [40], propionylation [41], butyrylation [41], serotonylation [42], dopaminylation [43], Glutarylation [44,45,46], Lactylation [47], Benzoylation [48], S-palmitoylation [49], O-palmitoylation [50] and 5-Hydroxylysine [51]. Also nonenzymatic histone PTMs have been identified including Glycation [52, 53], 4-Oxononanoylation [54, 55], Acrolein adduct [56, 57], Homocysteinylation [58, 59] nitrosylation [60,61,62,63], sulfe‐, sulfi‐, and sulfonylation [64, 65] and S-glutathionylation [66, 67]. For many of them, also enzymes have been identified that can remove the PTM, or ‘read’ the PTM and recruit other proteins to form a chromatin remodelling complex (for a detailed review of all histone modifying enzymes and their function see [26, 68]). Currently, two mechanisms are known by which histone PTMs can alter the state of chromatin. First, it is accepted that all histone modifications have the potential to affect higher order chromatin structure by neutralising the basic charge of the nucleosome, and therefore could loosen inter or intra-nucleosomal DNA-histone interactions [69,70,71,72]. A well-known example is acetylation of lysine residues, which removes the positive charge of lysine and therefore increases the probability to alter the structural state of chromatin [73]. Second, histone PTMs can recruit non-histone proteins to set in motion processes such as transcription, DNA repair and DNA replication [73]. Depending on which histone modification (or which sequence of histone modifications) are present at a given histone tail, different sets of proteins are encouraged to bind or prevent from binding to chromatin. This recruitment process is highly dynamic, as multi-step processes (e.g. transcription) require a distinct set of histone PTMs to recruit a distinct set of chromatin-remodelling proteins. Proteins that are recruited to PTMs contain specific PTM-reader domains (such as bromo-domains, chromo-(like) domains, PhD domains etc.). Many chromatin remodelers actually possess multiple reader domains, suggesting their ability for multivalent interactions that would increase both affinity and specificity [74, 75]. Functional interplay among writer-eraser PTM enzymes in the brain remains largely unknown. Recent reports, however, showed that knockout mice of the writer-eraser duo Kmt2a and Kdm5c, which are responsible for Wiedemann-Steiner Syndrome and Mental Retardation X-linked Syndromic Claes-Jensen, share similar brain transcriptomes, cellular- and behavioural deficits [76]. Double mutation of Kmt2a and Kdm5c however partly corrected H3K4 transcriptomes as well as their cellular and behavioural deficits, suggesting this balance is essential during development and might be an interesting therapeutic strategy for NDDs [76].
DNA methylation
In addition to methylation of histone tails, DNA can also be methylated to regulate chromatin state transitions (Fig. 1). The addition of a methyl group from S-adenosyl-L-methionine substrates only occurs on cytosines that are followed by guanines (called CpG sites), and is catalysed by DNA methyltransferases (DNMTs) leading to gene repression. There are two types of DNMT classes, namely either the de novo methyltransferases or maintenance methyltransferases [77]. DNMT3a and DNMT3b are classified as de novo methyltransferases as these can methylate previously unmethylated cytosine of CpG dinucleotides on both strands [78]. DNMT1 is classified as a maintenance methyltransferase as it has a substrate preference for hemimethylated DNA over unmethylated DNA. In contrast to its preference, DNMT1 can also display de novo methyltransferase activity in a specific cellular context-dependent manner [79]. DNA methylation can affect chromatin remodelling either by attracting transcriptional activators to the methylated cytosine [80], or it can attract transcriptional repressors that have methyl cytosine-binding domains. For example, DNA methylation can recruit histone deacetylases, which facilitate the formation of the silent chromatin state [81]. These methyl cytosine-binding proteins include methyl CpG-binding domains (MBDs) [82, 83] and methyl CpG-binding protein 2 (MeCP2) [84], which are known for their role in the aetiology of NDDs. During human development, genomic DNA methylation signatures are established in early development by two consecutive waves of nearly global demethylation, followed by targeted re-methylation [85, 86]. NDD mutations have often been found to underlie errors in methylation during these early time points [87, 88]. Consequently, altered methylation signatures during early development may be passed on across all cell lineages and can thus affect multiple tissues. When these epigenetic changes are maintained throughout development and across cell-types, these so called ‘episignatures’ can be used as biomarkers for the diagnosis of NDDs using easily accessible tissues such as peripheral blood [89,90,91,92]. Indeed, several very recent studies have already showed the potential for using disease-specific episignatures as diagnostic tool for NDDs, including patients with a known diagnosis as well as patients carrying variants of unknown significance [93,94,95].
ATP-dependent chromatin remodelers
The third class of chromatin modifying enzymes are the ATP-dependent chromatin remodelers including SWI/SNF (switch/sucrose-non-fermenting), ISWI (imitation switch), CHD (chromodomain-helicase-DNA binding) and INO80 (inositol requiring 80). In general, ATP-dependent chromatin remodelers hydrolyse ATP to generate enough energy to disrupt the interactions between histones and DNA. By doing so, ISWI remodelers can alter nucleosome positioning to promote heterochromatin formation and thus transcriptional repression. The SNF2/SNF chromatin remodelling family act as DNA translocases, by destroying histone-DNA bonds forming a DNA loop that propagates around the nucleosome until it reaches the exit site on the other side of the nucleosome [14]. Furthermore INO80 chromatin remodelers in vivo have been shown to play a role in nucleosome eviction, and histone variant exchange of the histone dimer H2A-H2B by the H2AZ-H2B dimer [96]. Finally, CHD family members exert a heterogeneous set of biological properties. One of the best studied examples is the NURD (nucleosome remodelling and deacetylase) complex, which contains lysine‐specific histone demethylase 1A (LSD1), Chromodomain Helicase DNA Binding Protein 3 (CHD3) or CHD4, histone deacetylases (HDAC1 or HDAC2) and methyl CpG-binding domain (MBD) proteins. The NURD complex has been shown to deacetylate specific gene sets during development leading to transcriptional repression [97].
ATP-dependent chromatin remodelers all share a conserved core ATPase domain, however all ATP-dependent chromatin remodelers harbour exclusive domains adjacent to the ATPase domain [14]. Each of these domains play a role in the recruitment of remodelers to chromatin, interaction with specific histone modifications and/or are involved in the regulation of the ATPase activity of the remodelers (see [14] for a detailed review on their function).
3D chromatin architecture
3D genome architecture is increasingly considered as an important epigenetic regulator of gene expression. On a coarse level, genomes are organised into structures known as chromosome territories (Fig. 1) [98]. These chromosome territories separate euchromatic from heterochromatic regions, and are termed ‘A’ and ‘B’ compartments, respectively [99]. Within the chromosome territories, megabase-sized chromatin domains are organised into smaller and smaller sub-domains known as topologically associated domains (TADs) [100]. TADs can be found in either ‘A’ or ‘B’ compartments, and are separated by sharp boundaries across which contacts are relatively infrequent. Interestingly, the boundaries between TADs are strikingly consistent across cell divisions and between cell types, as roughly 50%–90% of TAD boundaries have been shown to overlap in a pairwise comparisons between cell types [100]. In ‘A’ compartments, TADs are regions where DNA is highly organised in 3D space to enable “long-range” transcriptional regulation. This long-range transcriptional regulation is possible because enhancers are in close physical proximity to the promoters of their target genes in 3D space, despite long stretches of intervening nucleotides [101]. This physical proximity allows protein complexes bound at enhancers to interact with those bound at promoters (i.e. called enhancer-promotor loops), thereby influencing transcription of target genes. CCCTC-binding factor (CTCF) and Cohesin are such proteins that facilitate chromatin looping interactions [102]. CTCF-mediated loop formation requires one CTCF at each end of the chromatin loop, which dimerize if they are facing each other in the opposite orientation [103]. CTCF interacts with Cohesin via its C-terminal tail [104] and may thus allow Cohesin to locate on a particular side of the interaction to anchor and stabilize the chromatin loop [105]. In addition to Cohesin and CTCF, other proteins such as YY1 [106], ZNF143 and Polycomb group proteins [107], repetitive elements and PTMs are enriched at TAD boundaries to support transcription. These repetitive elements at TAD boundaries have been found to act as specific anchor points to spatially organize chromosomes [108], whereas enrichment for the transcription marks H3K4me3 and H3K36me3 in TAD boundaries show an association with highly expressed regions and suggests that transcription itself plays a role in TAD organisation [109]. By doing so, 3D chromatin structures play an essential role in orchestrating the lineage-specific gene expression programs that underlie cellular identity [110].
In summary, the above examples show that cells possess a wide range of chromatin remodelers to translate external signalling cues into lasting changes in gene expression (Fig. 1). Interestingly, chromatin remodelers are especially strongly expressed in the brain [111], and it is therefore not surprising that impaired chromatin remodelling in any of the above described remodeler classes has been identified to a cause monogenic forms of NDDs [68, 112,113,114]. Chromatin remodelers are often multifunctional [15], and by doing so have the ability to play divergent roles in the multi-step continuum of brain development. This continuum encompasses neural progenitor generation and specification, cell-type differentiation and expansion, migration and circuit integration. Dysfunction of chromatin remodelers at any point during this developmental continuum will result in lasting changes on mature network function. In the next section we will discuss the different steps along this developmental continuum, and explain how altered chromatin remodelling at any of these steps ultimately affects the structure and function of mature neuronal networks.
Epigenetic modulation during neurodevelopment and disease
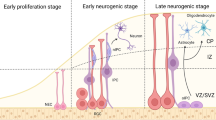
In the early stages of neocortical development, the telencephalic wall is composed neuroepithelial (NE) cells that will give rise to diverse pools of progenitor cells [115]. As these progenitors proliferate and expand in number, some begin to differentiate into radial glia cells (RGCs), establishing the ventricular zone (VZ). RGCs in turn begin to produce projection neurons and intermediate progenitors (IP) around E11.5 in mice which establish the subventricular zone (SVZ) [116]. In human development, RGCs not only produce projection neurons and IPs, but also the human specific outer radial glia (oRG) between gestational week (GW) 16–18, which will populate the outer SVZ (oSVZ) [117]. The RGCs in mice and oRG in humans act as transit-amplifying cells to increase the population of glutamatergic neurons until E18 in mice [118] or GW 21 in the human neocortex [119, 120], after which they switch to local glia production [121] (Fig. 2).
a Developmental timeline of the neocortex (top), with the generation times of the three most important cell classes indicated: Glutamatergic neuron generation in the cortical plate, glia production in the Medial Ganglionic Eminence (MGE) and cortical plate, and GABAergic neuron production in the Ganglionic Eminences (GEs). The timeline is based on mouse cortical development (indicated at the bottom as embryonal days (E), a mouse pregnancy lasts 19.5 days on average), with approximate human weeks post conception (W) displayed for reference. b The effect of chromatin remodelling defects on each cell population, with representative examples chosen from the NDD genes described in this review. Top four panels, defects in CHD family members (Chromodomain Helicase DNA Binding Protein), Setdb1/Ezh2, and BAF complex members on glutamatergic cell generation and maturation. Middle panel, comparable effects of mutations in two different chromatin remodelers on the generation of MGE-derived PV + and SST + neurons. Bottom panel, opposite effect of mutations in different chromatin remodelers on astrocyte generation timing
Epigenetic modulation during neocortical development
Histone PTMs
Mutations in the Polycomb repressive complex (PRC) are a prime example of how defective chromatin remodelling influences progenitor proliferation. The PRC consists of two complexes: PRC1 and PRC2. Although for PRC1 no dominant germline mutations have been described, autosomal recessive mutations in the PRC1 complex protein PHC1 have been shown to cause a form of microcephaly with short stature in two Saudi siblings [122]. Loss of PHC1 resulted of the inability of patient cells to ubiquitinate H2A, resulting in increased Geminin expression causing cell cycle abnormalities and impaired DNA damage response [122]. Additionally, mutations of interactors of PRC1 are mutually vulnerable to cause NDDs with brain volume abnormalities. For example, mutations in the PRC1 interactor AUTS2 have been shown to cause an autosomal dominant form of syndromic mental retardation, including comorbidities such as ID, ASD, microcephaly, short stature and cerebral palsy [123, 124]. Mouse embryonic stem cells (mESCs) carrying heterozygous mutations in Auts2 showed an increase in cell death during in vitro corticogenesis, which was rescued by overexpressing the human AUTS2 transcripts. Furthermore, mESCs harbouring a truncated AUTS2 protein (missing exons 12–20) showed premature neuronal differentiation, whereas cells overexpressing AUTS2 showed increase in expression of pluripotency markers and delayed differentiation.
The PRC2 complex consists of core subunits Enhancer of Zeste 2 (EZH2) and its homolog EZH1, which catalyse mono‐, di‐, and tri‐methylation of H3K27 resulting in heterochromatin formation and gene repression. EZH2 is mainly expressed during embryogenesis, while EZH1 is more ubiquitously expressed in adult and quiescent cells [125]. Loss-of-function mutations in EZH2 and thus reduced H3K27me3 levels in humans have been shown to cause Weaver syndrome, causing overgrowth and macrocephaly, accelerated bone maturation, ASD, developmental delay and characteristic facial features [126,127,128]. In accordance with the overgrowth phenotype in Weaver syndrome, conditional KO of Ezh2 in mice accelerated proliferation of neuronal precursors in the cerebral cortex at the expense of self-renewal of progenitors [129]. These results were strengthened by showing that fate-mapped E14-born neurons in a cKO for the PRC2 complex member Eed (which interacts with EZH2) mainly resided in the layer 2/3, in contrast to WT E14- born neurons, which resided in layer 4 [130]. These results confirmed PRC2 regulates the progression of apical progenitor’s temporal specification [130, 131].
Progenitors follow a specific pattern of cell divisions to initiate the build-up of the different layers in the cortex [132]. This is done in an ‘inside-out’ fashion, meaning the early-born neurons form the deep neocortical layers (i.e. 6 and 5), whereas late born neurons radially migrate through the deeper layers to create the more superficial layers (layer 4 and 2/3). Each RGC has been shown to consistently produce 8 to 9 glutamatergic neurons progressing from lower-layer excitatory neurons at mouse E12.5 to glutamatergic neurons that undergo radial migration towards the upper-layers at mouse embryonal day E15.5 [130]. Approximately 1 in 6 RGCs switches to gliogenesis, generating astrocytes and oligodendrocytes at the end of the cell cycle [133]. The process of neuroprogenitor specification during corticogenesis has been studied and reviewed in great detail elsewhere (see [134] or for a review see [116]) and will therefore not be discussed in this review. Numerous studies however have shown that defects in chromatin remodelers during RGC specification result in shifts in the neuron classes produced or in precocious cell cycle exit and gliogenesis, and thereby could contribute to the neuronal phenotypes found in NDD patients [135]. In mice, Ezh2 is highly expressed in RGCs up to E14.5 and has been proposed to regulate RGC identity and proliferation behaviour, as well as RGC‐to‐glial‐progenitor transition [129, 136, 137] by inhibiting Neurogenin 1 expression [136]. Ablation of Ezh2 in mouse RGCs correlates with premature RGC differentiation as described earlier, increased generation of lower‐layer neurons, decreased upper‐layer neuron production, and precocious astrocyte generation [129]. Furthermore, knockdown of Ezh2 in mice has been shown to affect the neuronal polarization and radial neuronal migration [138].
In addition to PRC2, deletion of the PRC1 component Ring1b at E13 prolongs the expression of Fez transcription factor family member zinc-finger 2 (Fezf2). The prolonged expression of Fezf2 results in the continuous expression of downstream target genes such as Ctip2, resulting in an increased production of deep-layer neurons [139]. Interestingly, when Ring1b is knocked out later in developmental time at E14.5, the number of upper-layer neurons is increased instead, due to an extended neurogenic period [136].
Similar to H3K27me, H3K9me2/3 is a repressive mark which is established by the methyltransferases SETDB1, (KMT1E), SUV39H1 (KMT1A), G9a (EHMT2, KMT1C) and G9a‐like protein (EHMT1, KMT1D). SETDB1 has been associated to several NDDs. In humans, two missense mutations [140] and a microdeletion [141] in SETDB1 were found in a large cohort of ASD patients. In addition, SETDB1 has been implicated to play a role in the aetiology of Schizophrenia by regulating GRIN2B expression [142]. Moreover, SETDB1 has been shown to influence chromatin 3D structure by binding to a non-coding element upstream of the Pcdh cluster [143, 144] which was a near-perfect match to a Schizophrenia risk haplotype (number 108 in [144]). Finally, SETDB1 is also described to play an indirect role in Prader-Willi syndrome, by contributing to the maternal silencing of the SNORD116 gene [145, 146]. Setdb1 is highly expressed early in corticogenesis (E9.5) in NE cells in the VZ, however its expression declines at E15.5 [135]. While deletion of Setdb1 does not affect RGC numbers, it has been shown to reduce layer V and VI Ctip2+ basal progenitors between E14.5 and E16.5 [147]. At the same time, an increase of the number of Brn2+ layer II and III neurons was found in the CP. This shift in the production of upper layer neurons at the expense of deep layer neurons remained after neurogenesis ceased at E18.5. All these events together lead to a reduced cortical volume in Setdb1 knockout mice at E18.5 and P7 [147]. Moreover, deletion of Setdb1 causes accelerated astrogliogenesis, demonstrating that Setdb1 does not only regulate the timing of late neurogenic events, but also the RGC‐to‐astrogenic‐progenitor transition [147]. As SETDB1 is a H3K9 methyltransferase, its general role is to repress gene transcription, or methylate DNA. SETDB1 does so by being involved in several complexes. SETDB1 interacts with The Krüppel-associated box (KRAB) domain-containing zinc-finger proteins (KRAB-Zfp) and KRAB domain-associated protein 1 (KAP-1) [148], which recruit the NuRD complex and HP1 to form a repressive complexes [149]. SETDB1 can also interact with MBD1 and ATF7IP [150], which has been suggested to play a role in X-inactivation [151]. To repress gene transcription via DNA methylation, SETDB1 interacts with DNMT3A/B in cancer cells, to represses the expression of p53BP2 and RASSF1A [152]. Through the interaction with these complexes, SETDB1 can thus target various genomic loci in different cell types, at different stages during brain development.
Like SETDB1, EHMT1 plays a major role in embryonic development, as full knockout of this gene leads to embryonic lethality [153]. In hematopoietic stem cells, EHMT1 activity is essential to facilitate the long-term silencing of pluripotency genes, and thus inhibition of EHMT1 is essential for maintaining pluripotency [154]. Furthermore, a reduction of large H3K9me2 chromosome territories was found in these stem cells [154], which are proposed to stimulate lineage specification by affecting the higher order chromatin structure [155]. Little is known about the role of EHMT1 during early neurodevelopment, however knockdown of EHMT1 in NPCs revealed differential expression of genes important in development, such as BMP7, WNT7A, CTNNB1, TGFB2 and CHD3 [156]. Furthermore, in neural progenitors EHMT1 has been found to interact with ZNF644 to silence multipotency and proliferation genes. Disruption of the ZNF644/EHMT1 resulted in maintenance of proliferative identity and delayed formation of differentiated retinal neurons [157]. Similar roles for EHMT1 in the regulation neural progenitor genes were found in a conditional knockout mice (Camk2a-Cre; GLPfl/fl) [158]. Interestingly, no brain volume abnormalities are described in animal models of EHMT1 haploinsufficiency [158, 159] whereas microcephaly was found in 20% of patients carrying intragenic EHMT1 mutations [160]. These patients are diagnosed with Kleefstra Syndrome, which is in addition to microcephaly characterised by mild to severe ID, ASD, developmental delay, speech problems, hypotonia, characteristic facial features, epileptic seizures, heart defects and various behavioural difficulties [160, 161].
Histone acetylation by HATs and removal by HDACs is another example where deficits in chromatin remodelling affect the cellular distribution during corticogenesis. For example, the gene encoding the HAT cAMP‐response element binding protein (CBP) is highly expressed in proliferating RGCs and post‐mitotic neurons during corticogenesis [162], and CBP null mice have been shown embryonically lethal due to failure of neural tube closure (E9-E12.5) [163], stressing their role in neurodevelopment [164]. In humans, heterozygous mutations in CBP are associated with Rubinstein–Taybi syndrome (OMIM# 180849), which is characterized by ID, postnatal growth deficiency, microcephaly, broad thumbs and halluces, and dysmorphic facial features [68]. In mice, similar brain volume abnormalities have been described [165]. CBP induces acetylation of H3K9, H3K14 and H3K27 within target gene promoters, such as α1‐tubulin (E13‐E16), Gfap (E16‐P3), S100β, Plp2 and Mbp (P14) [162]. Accordingly, heterozygous loss of Cbp in mice diminishes acetylation at these promoters and leads to decreased differentiation of progenitor towards astrocytes and neurons [162]. Consequently, a reduced number of neurons, astrocytes and oligodendrocytes were observed in the cortex of heterozygous Cbp mice around P14, whereas an increase in PAX6 expressing progenitors was found as compared to wild-type mice [162]. At later ages (P50), only a reduction in gliogenesis remained in the corpus callosum [162]. Another example of a HAT important in NPC development is Lysine acetyltransferase 8 (KAT8), a member of the Non-specific Lethal (NSL) complex which is responsible for acetylation of H4K16 and plays a role in H4K16 propionylation [166]. Cerebrum specific knockout of Kat8 in mice has recently been shown to cause severe cerebral hypoplasia in the neocortex and in the hippocampus, together with postnatal growth retardation and pre-weaning lethality [166]. Furthermore, these mice showed a loss of RGC proliferation and thus reduced progenitor pool at E13.5, massive apoptosis starting at E12.5 and increased numbers of Tuj1+ cells, indicating precocious neurogenesis at E13.5 [166]. Similarly, patients with KAT8 mutations presented with variable brain MRI abnormalities, epilepsy, global developmental delay, ID, facial dysmorphisms, variable language delay, and other developmental anomalies [166]. Interestingly, patients with epilepsy responded well to the histone deacetylase inhibitor Valproate [166], stressing the importance of KAT8 and other lysine acetyltransferases function in brain development.
Within the NSL complex, KAT8 is regulated by the NSL regulatory subunits KANSL1 and KANSL2. Mutations in KANSL1 cause Koolen-de Vries Syndrome (OMIM# 610443), characterized by ID, distinctive facial features, and friendly demeanour [167]. Likewise, KANSL2 mutations have been identified in ID patients [8]. Interestingly, haploinsufficiency of Kansl1 in the mouse causes craniofacial abnormalities, reduced activity levels and impaired fear learning, as well as epigenetic dysregulation in genes linked to glutamatergic and GABAergic cells [168].
The balance between acetylation and deacetylation plays an important role in progenitor proliferation and differentiation, as inhibition of HDAC activity also results in progenitor proliferation/differentiation deficits [169]. Histone acetylation is removed by HDACs, resulting in chromatin condensation and transcriptional repression [170]. So far, over 18 HDACs are characterised in the mammalian genome, and they are expressed in a cell type- and developmental stage-dependent fashion [171]. For example, HDAC1 is highly expressed in neural stem cells/progenitors and glia, whereas HDAC2 expression is initiated in neural progenitors and is up-regulated in post-mitotic neuroblasts and neurons, but not in fully differentiated glia [172]. Conditional knockout of Hdac1 or Hdac2 in mice progenitors impairs neuronal differentiation [173]. Specifically, conditional knockout of Hdac1 and Hdac2 in Gfap-Cre mice resulted in major brain abnormalities and lethality at around P7 [173], whereas conditional knockout of Hdac1 and Hdac2 in Nestin-Cre mice resulted in reduced proliferation and premature differentiation of NPCs prior to abnormal cell death [174]. Moreover, inhibition of HDAC activity at the neurogenic stage decreases the production of deep-layer neurons and increases the production of superficial-layer neurons [175]. Conditional deletion of Hdac1 and Hdac2 in oligodendrocyte precursors results in severe defects in oligodendrocyte production and maturation [176]. Recently, the first patient carrying a mutation in HDAC2 has been characterised presenting with many clinical features consistent with Cornelia de Lange Syndrome (CdLS) including severe developmental delay, limb abnormalities, congenital heart defects, altered development of the reproductive system, growth retardation and characteristic craniofacial features [177]. No patients harbouring mutations in HDAC1 have been characterized as to date.
Next to HDAC2, missense mutations in HDAC8 are also linked to the aetiology of CdLS presenting with overlapping clinical features (OMIM# 300269) [178, 179]. HDAC8 plays a key role in regulating cohesion function by deacetylating one of the core cohesion proteins, SMC3, which affects mitosis as well as transcription through loss of TAD function [180, 181]. Loss of HDAC8 activity in SVZ progenitors from 4-month old mice has been shown to reduce progenitor proliferation and differentiation [182]. Moreover, knockdown of HDAC8 in the mice embryonic carcinoma cell line P19 cells permitted the formation of embryoid bodies [183]. Furthermore, loss of HDAC8 in zebrafish has been found to increase apoptosis in CNS progenitors [182]. Recently a novel intronic variant in HDAC8 was found in a large Dutch family with seven affected males presenting with X-linked ID, hypogonadism, gynaecomastia, truncal obesity, short stature and recognisable craniofacial manifestations resembling but not identical to Wilson-Turner syndrome (OMIM# 309585) [184]. This variant disturbs the normal splicing of exon 2 resulting in exon skipping, and introduces a premature stop at the beginning of the HDAC catalytic domain [184]. How this specific variant influences neurodevelopment remains elusive.
Chromatin 3D organisation
The 3D organization of chromatin is changing dynamically as the cell differentiates. Whereas the nuclei of embryonic stem cells have been shown to be relatively homogenous, heterochromatin foci are becoming more apparent during differentiation into progenitors. When these progenitors differentiate into neurons, the heterochromatin foci are becoming even larger, suggesting that heterochromatin regions are actively reorganized during differentiation [185]. Deficiencies of 3D chromatin organizers such as CTCF and Cohesin are associated with the aetiology of NDDs called CTCF-associated NDDs and cohesinopathies, respectively. Heterozygous mutations in CTCF (OMIM# 604167) have been shown to cause the NDD mental retardation, autosomal dominant 21 (MRD21), which is characterised by variable levels of ID, microcephaly, and growth retardation [186,187,188]. In mice, loss-of-function studies of Ctcf revealed an important role for this protein in cell fate specification and neural differentiation. Knockout of Ctcf at E8.5 resulted in upregulation of PUMA (p53 upregulated modulator of apoptosis), leading to high levels of apoptosis and loss of the telencephalic structure [189]. Inactivation of CTCF several days later (E11) also resulted in PUMA upregulation and increased apoptotic cell death, and again the CTCF-null forebrain was hypocellular and disorganized at birth [189]. In contrast, conditional knockout of CTCF in postmitotic projection neurons resulted in misexpression of clustered protocadherin (Pcdh) genes leading to altered functional neuronal development and neuronal diversity [190]. These results suggest that CTCF activity regulates the survival of neuroprogenitor cells, and the balance between neuroprogenitor cell proliferation and differentiation [189].
The two best-described cohesinopathies are CdLS (OMIM# 122470, 300590, 610759, 614701, 300882) and Roberts syndrome (and its variant SC Phocomelia, OMIM# 268300). As described above, CdLS can be caused by loss of function mutations in HDAC2 and HDAC8. Additionally, mutations in three Cohesin subunits (SMC1α, SMC3, RAD21) [191, 192] and in one Cohesin-interacting protein (NIPBL) [193] have been found causal for CdLS, whereas mutations in ESCO2 are responsible for Roberts syndrome/SC Phocomelia (OMIM# 269000) [194]. While a full knockout of Cohesin subunits in mice is lethal, mice carrying heterozygous mutations in these genes are viable and show altered gene expression in developmental programs, DNA repair and replication [195].
Chromatin remodelling complexes: CHD proteins and the NuRD complex
Altered chromatin remodelling by multi-subunit protein complexes has been shown to play a role in RGC differentiation and in the aetiology of NDDs. One of these complexes, called the NuRD complex, consists amongst other proteins of LSD1, HDAC1/2, and a Chromodomain Helicase DNA (CHD) Binding Protein (either CHD3, 4 or 5) [196]. These CHD proteins have been shown to play essential roles during neurodevelopment, as pathogenic variants in CHD1, CHD2, CHD3, CHD4, CHD6, CHD7 and CHD8 have been associated with a range of neurological phenotypes. Of the nine human CHD family members that have been characterized (CHD1-9), further subdivisions are being made into subgroups based on their function. Subfamily one consists of CHD1 and CHD2 because of their shared DNA binding domain that is not well-conserved in the other CHD proteins [197]. CHD1 has been found to play an essential role in early mice development, as Chd1−/− embryos show proliferation defects and increased apoptosis, are smaller than controls by E5.5 and fail to become patterned or to gastrulate [198]. Similar results in decreased self-renewal and pluripotency were found using knockdown of Chd1 in mESC cells [199]. Furthermore, Chd1−/− ESCs show deficits in self-renewal and a reduction in genome-wide transcriptional output by directly affecting ribosomal RNA synthesis and ribosomal assembly [198]. In contrast, mice lacking a single Chd1 allele (Chd1+/−) are healthy, fertile and phenotypically normal [198].
Recently the first patients with CHD1 missense mutations were identified, which presented with ID, ASD, developmental delay, speech apraxia, seizures, and dysmorphic features. Interestingly, also a patient with a microdeletion spanning RGMB and the last exons of CHD1 was characterised with no obvious NDD phenotype, suggesting that whereas deletions of CHD1 may not cause a consistent neurological phenotype, missense mutations in CHD1 may do so via a dominant negative mechanism [200].
Despite the fact that CHD family members are rather ubiquitously expressed, only CHD2 pathogenic variants cause a brain-restricted phenotype, suggesting a unique role for this gene in neurodevelopment. Based on loss-of-function studies, CHD2 has been shown to regulate self‐amplification of RGCs and prevents precocious cell-cycle exit. CHD2 is mainly expressed in RGCs between E12-E18 and rarely in IPs, however knockdown of Chd2 in utero resulted in a reduction of RGCs in the SVZ whereas an increase was found in the number of produced IPs and neurons (Fig. 2) [201, 202]. This premature differentiation in RGCs can lead to a depletion of the progenitor pool, resulting in a smaller overall brain volume as a consequence [203]. Indeed patients harbouring mutations in CHD2 present with a reduced head size and in 20% of the cases microcephaly [204, 205], developmental delay, ID, ASD, epilepsy and behavioural problems with phenotypic variability between individuals [206]. A subset of patients carrying CHD2 pathogenic variants present with developmental and epileptic encephalopathy (also called Dravet Syndrome), which is an early onset of epilepsy disorders characterized by refractory seizures and cognitive decline or regression associated with ongoing seizure activity [207].
CHD3, CHD4 and CHD5 are categorised in the second CHD family because they share dual plant homeodomain zinc finger domains [207]. Furthermore, these class 2 CHD remodelers exhibit subunit‐specific functions and display mutually exclusive occupancy within the NuRD complex at different stages of corticogenesis [208]. CHD3, has been shown to play an important role in the correct cortical layering and controls the timing of upper‐layer neuron specification [208]. In mice, CHD3 expression starts around E12.5 where it is still low expressed, and this increases from E15.5 to E18.5. At these later time points, CHD3 is mainly expressed in upper layer neurons in the cortical plate. Neurons lacking CHD3 (CHD3-knockdown) have been found more likely to express transcription factors that regulate laminar positioning and differentiation of deeper cortical layers (i.e. Tbr1 and Sox5), whereas a lower number of neurons expressed the upper layer markers Brn2 and Cux1, implicating that CHD3 may influence the expression of genes that couple radial migration with laminar identity (Fig. 2) [208]. Patients with CHD3 mutations are only very recently identified with Snijders Blok–Campeau syndrome, which is characterized by ID (with a wide range of severity), developmental delays, macrocephaly, impaired speech and language skills, and characteristic facial features [209, 210].
The second CHD protein that plays a role in the NuRD complex is CHD4. Mice lacking Chd4 almost always died at birth, however when examining brain volumes at E18.5 a significant reduction of the cortical thickness was found, caused by reduced NPC proliferation and premature cell cycle exit, followed by increased apoptosis of premature born neurons [208]. As a result, CHD4fl/fl/Nestin-Cre mice presented with lower numbers of IPs and late born upper layer neurons (Fig. 2) [208]. Interestingly, Chd4 appears to guide Polycomb repressor complex (PRC2) and especially Ezh2 to opposing effects early vs. late in corticogenesis, first interacting to repress the gliogenic gene Gfap, and later repressing the neurogenic Ngn1 after the neurogenic-to-gliogenic switch [137]. Interestingly, patients carrying mutations in CHD4 actually present with macrocephaly, amongst other characteristics like ID, hearing loss, ventriculomegaly, hypogonadism, palatal abnormalities and facial dysmorphisms that are diagnosed by Sifrim–Hitz–Weiss syndrome [211]. The opposing phenotypes found for brain volume between rodent models and patients might be explained by a gene dosage effect, as for some variants in CHD4 altered ATPase activity levels were found, suggesting a possible gain-of-function phenotype in certain patients [212, 213]. Another possibility might be that the NuRD complex function is differentially regulated in humans and mice, as was recently suggested in a study comparing mouse and human pluripotent stem cells [214].
Finally, CHD5 has only recently been characterized as one of the core subunits of the NuRD complex [215]. CHD5 is the only CHD member that is mainly expressed in the total brain, foetal brain and cerebellum [216]. Chd5 expression was mainly found in late-stage neuronal progenitors undergoing terminal differentiation, rather than in proliferating progenitors [215]. In utero knockdown of Chd5 furthermore resulted in an accumulation of undifferentiated progenitors, which were unable to exit the VZ, SVZ and intermediate zone (IZ; Fig. 2) [215]. Additionally, knockdown of Chd5 in mouse ESCs resulted in a failure to upregulate late stage neuronal genes [215]. Similar results for the role of Chd5 in neuronal gene regulation were found in primary rat cultures [217]. A de novo damaging missense variant in the CHD5 gene was identified in an ASD proband from the Autism Sequencing Consortium [9]. In accordance, knockout of Chd5 in mice indeed has been shown to cause ASD-like behaviour including increased anxiety and decreased social interaction [218].
The third subfamily consists of the remaining family members CHD6-9 [219]. CHD6 mutations have previously been described, including a large translocation in one Pitt-Hopkins patient [220], in a single case of mental retardation [221], in sporadic acute myeloid leukaemia incidences [222], and most recently for the very rare Hallermann–Streiff syndrome [223]. CHD6 is the least studied member of the CHD family, and little is known for its contribution during neurodevelopment.
Similar as to other members of the CHD family, loss of CHD7 resulted in impaired proliferation and self-renewal of RGCs in the SVZ. Consequently, a reduction of NE thickness in telencephalon and midbrain was shown in Chd7 homozygous gene-trap mutant embryo at E10.5 [224]. Similarly, mESCs from Chd7Gt/Gt mice showed a reduced potential to differentiate into neuronal and glial lineages, and presented with altered accessibility and expression of NPC genes. Furthermore, neurons generated from these Chd7Gt/Gt mESCs presented with a significant lower length and complexity [225]. Furthermore, CHD7 plays a key role in oligodendrocyte precursor survival and differentiation (Fig. 2) [226, 227] and has been shown to cooperate with Sox10 to regulate myelination and re-myelination [227]. Additionally, CHD7 has been shown to play an important role in cerebral granular cell differentiation and cell survival [228].
Mice lacking one Chd7 copy (found in Chd7COA1/+ mice [229] and Chd7Gt/+ mice [230]) also often present with brain abnormalities including the absence or hypoplasia of olfactory bulb, cerebral hypoplasia, defects in the development of telencephalic midline and reduction of the cortical thickness [229, 230]. Patients with CHD7 mutations present with similar brain structure abnormalities including hypoplasia of olfactory bulb and cerebellum, agenesis of the corpus callosum, microcephaly and atrophy of the cerebral cortex [231,232,233]. Additionally, in relation to its role in oligodendrocyte differentiation and function, some patients have been characterised with white matter defects [234, 235]. Loss of CHD7 in patients is called CHARGE syndrome (OMIM# 214800), which is next to the brain malformations characterised by coloboma, heart defects, growth retardation, genital hypoplasia, and nose and ear abnormalities (including choanal atresia, deafness and vestibular disorders) [236].
Finally, CHD8 has been identified as a causal gene for ASD, presenting with common phenotypic features included macrocephaly, accompanied by rapid early postnatal growth, characteristic facial features, increased rates of gastrointestinal complaints and marked sleep dysfunction [237]. Chd8 is strongly expressed around the transition from symmetric proliferative to asymmetric neurogenic RGC division [238], and knockdown of Chd8 at E13 has been shown to prematurely deplete the neural progenitor pool in developing mice cortices (Fig. 2) [239]. Similarly, both heterozygous and homozygous knockout of Chd8 in mESCs resulted in an upregulation of neuronal genes upon differentiation into NPCs [240]. Indeed, CHD8 binds the promoters of cell cycle genes and serves as a transcriptional activator of for example PRC2 components Ezh2 and Suppressor of Zeste 12 [239], which allows for the repression of neural genes during this developmental period [129]. In human iPSC-derived CHD8+/− organoids, a number of ASD risk genes was upregulated [241]. Furthermore, CHD8 is identified as a negative regulator of the Wnt–β-catenin signalling pathway [242], as knockdown of Chd8 in non-neuronal cells lead to an enrichment of up-regulated Wnt–β-catenin signalling pathway genes [239]. Interestingly, knockdown of Chd8 in neuronal cells lead to an enrichment of down-regulated Wnt–β-catenin signalling pathway genes, indicating CHD8 plays cell-type specific roles [239]. Furthermore, conditional knockout of CHD8 in oligodendrocytes (Chd8flox/flox;Olig1-Cre+/−) has shown to impair oligodendrocyte differentiation and myelination in a cell-autonomous manner [243, 244]. Whereas homozygous deletion of Chd8 in mice results in early embryonic lethality [245], Chd8 heterozygous mice were viable and presented with similar phenotypes as patients, including macrocephaly, abnormal craniofacial features, and ASD like behaviour [246,247,248,249,250]. Moreover, introducing the human truncating variant N237K into the mouse Chd8 gene (Chd8+/N2373K) revealed ASD-like behaviour, aberrant vocalization, enhanced mother attachment behaviour and enhanced isolation-induced self-grooming specifically in males, but not females [251]. These phenotypes were also conserved in zebrafish, where Chd8 knockdown was found to cause macrocephaly and gastrointestinal phenotypes [237, 238].
Chromatin remodelling complexes: SWI/SNF
Also SWI/SNF chromatin remodelling complex subunits are expressed in a temporal and cell‐type specific manner [199, 252]. During differentiation from embryonic stem cells into neurons, the SWI/SNF complex begins to switch subunits to those unique to neural progenitors, followed by subunits specific to neurons [253, 254]. Neural progenitor proliferation requires a SWI/SNF complex containing PHF10 and ACTL6A subunit, which are replaced by the subunits DPF1, DPF3, and ACTL6B when neural progenitors exit the cell cycle to become post-mitotic neurons [255, 256]. Interestingly, the neural progenitor-specific SWI/SNF complex exclusively incorporates either SMARCC2 or SMARCC1 subunits at distinct developmental stages. In the mouse, neural progenitor SWI/SNF complexes harbours SMARCC2 until E14.5 to repress intermediate progenitor generation, whereas between E14.5 and E15.5, SMARCC2 is replaced by SMARCC1, activating intermediate progenitor generation in RGCs via the interaction with the H3K27 demethylases JMJD3 and UTX [257,258,259]. Double loss of Smarcc2 and Smarcc1 from as early as E10.5 (Smarcc1fl/fl:Smarcc2fl/fl, Emx-Cre) resulted in reduced numbers of proliferative progenitors, thinning of the cortical SVZ and loss of projection neurons [259]. In addition, loss of Smarcc2 and Smarcc1 in late NPCs (Smarcc1fl/fl:Smarcc2fl/fl, hGFAP-Cre) resulted in H3K27me3-mediated silencing of neuronal differentiation genes, causing delayed cortical and hippocampal neurogenesis [260]. Similarly, a loss of the catalytic subunit Smarca4 in late NPCs (Smarca4fl/fl::hGFAP-Cre) resulted in reduced cortical thickness, dendritic abnormalities, hippocampal underdevelopment and cerebellar disorganization [261]. Furthermore, Smarcc2 and Smarcc1 double knockouts showed an upregulation of Wnt signalling activity resulting in increased progenitor proliferation-related defects [260]. This Wnt /β-catenin pathway has indeed previously been shown to be a critical regulator of NPC proliferation and neurogenesis during cortical development [116, 262, 263]. Thus, timely expression of these SWI/SNF subunits [264] is essential for regulating cell fate during neurodevelopment, and can control this processes by regulating Wnt signalling activity [262].
Not only SMARCC1 and SMARCC2 expression is essential during brain development, also divergent patterns of expression of SMARCA1 and SMARCA5 were found in the mouse embryo. Whereas Smarca5 is mainly expressed in proliferating progenitors in the neocortex and the cerebellum, Smarca1 is predominantly expressed in terminally differentiated neurons after birth in the cerebellum and hippocampus of adult animals [265, 266]. Consequently, Smarca5-null mice die during early preimplantation due to hypoproliferation of the inner cell mass [267], whereas Smarca1-null mice develop normally, but show hyperproliferation of progenitors causing an enlarged brain size [268]. Both remodelers have been shown to play a role in the proliferation and differentiation of IPs by controlling FoxG1 expression. FoxG1 is a critical transcription factor for IP proliferation and control of the timing of neurogenesis [269, 270]. On the one hand, conditional knockout of Smarca5 in forebrain progenitors (Emx-cre) resulted in reduced FoxG1 expression, impaired cell cycle kinetics and increased cell death. This resulted in a reduced number of Tbr2+ and FoxG1+ intermediate progenitors and thus a reduced cortical size [271]. Similarly, conditional knockout of Smarca5 (Nestin-cre) caused reduced brain size and cerebral hypoplasia as a result of reduced granular progenitor proliferation [266]. On the other hand, loss of Smarca1 (Ex6DEL) has been shown result in an increased FoxG1 expression, a disruption of progenitor cell cycle kinetics, increased progenitor proliferation and increased neurogenesis [268]. Furthermore, Smarca1 has been shown to directly bind to the promotor of the FoxG1 gene, suggesting that timed chromatin remodelling by SMARCA1 is essential for controlling neuronal development and differentiation [268]. Taken together, these results confirm that timed chromatin remodelling of SWI/SNF-remodelers is essential during neurodevelopment [272]. It is therefore thus not surprising that misexpression of SWI/SNF complex subunits cause NDDs.
SWI/SNF-Related Intellectual Disability Disorders comprise a spectrum of disorders that includes the classic Coffin-Siris syndrome (CSS, OMIM# 135900) and Nicolaides–Baraitser syndrome (NCBRS, OMIM# 601358) [273]. These disorders differ amongst each other in a phenotypic spectrum ranging from syndromic ID over to classic and atypical/severe CSS to NCBRS. The core manifestations of CSS include ID, hypotonia, feeding problems, characteristic facial features, hypertrichosis, sparse scalp hair, visual problems, lax joints, short fifth finger, and one or more underdeveloped nails [274], and in a minority of the cases, microcephaly [275]. In contrast, NCBRS is defined by ID, short stature, microcephaly, typical face, sparse hair, brachydactyly, prominent interphalangeal joints, behavioural problems, and seizures [276]. Interestingly, structural brain midline defects such as corpus callosum malformations or even absence, are described in the majority of CSS patients [274, 277,278,279,280] and in some NCBRS patients [281], and animal models for these disorders [282].
The mild form of CSS is caused by either mutations in the ATPase subunit ARID1B (OMIM# 614556) [283], or by pathogenic changes in other chromatin remodelling proteins with no direct interaction with SWI/SNF complex, including SOX11 (OMIM# 600898) [256] and DPF2 (OMIM# 601671) [284]. Additionally, classic and more severe CSS are known to be caused by mutations in SMARCA4 (OMIM# 603254), the common core subunit SMARCB1 (OMIM# 601607), and accessory subunits such as SMARCE1 (OMIM# 603111), ARID1A (OMIM# 603024) and ARID2 (OMIM# 609539) [285]. NCBRS on the contrary has been shown to be caused by mutations in SMARCA2 (OMIM# 600014). Other patients found with mutations in SWI/SNF complex subunits are described for SMARCB1 (OMIM# 601607) have been shown to cause either CSS, but also DOORS syndrome (OMIM# 220500) or Kleefstra syndrome (OMIM# 610253) depending on the site/location of the mutation [286, 287]. Together, these studies on chromatin remodelling complexes support the notion that combinatorial assembly of subunits can instruct cell lineage specification by creating specific patterns of chromatin states at different developmental stages, are essential for normal neurodevelopment [288, 289], and will result in clinical overlapping phenotypes [279].
Chromatin remodelling complexes: histone variant remodelers
The third class of ATP dependent chromatin remodelers is the INO80 family. The INO80 subfamily is known for its role in histone variant exchange of canonical H2A or H3 histone variants, which is assisted by editing remodelers such as Swr1 complex (SWR1C) [290], mammalian Snf2-related CBP activator protein (SRCAP) [291] and p400. Recently, INO80 function in NPCs has been found essential in homologous recombination (HR) DNA repair in a p53-dependent manner [292]. Loss of Ino80 in NPCs (Neurod6Cre/+;Ino80fl/fl) impairs these processes, causing apoptosis and microcephaly in mice [292]. Interestingly, Ino80 deletion in mice leads to unrepaired DNA breaks and apoptosis in symmetric NPC-NPC divisions, but not in asymmetric neurogenic divisions [292]. In correspondence with these findings, INO80 was recently identified as a candidate gene for ID and microcephaly [293]. Among the key histone variants that can be incorporated by the INO80 family is the H2A variant H2A.Z. Specifically, it has been shown that SRCAP removes canonical H2A–H2B dimers and replaces them with H2A.Z–H2B dimers [294]. Little is known yet how mutations in SRCAP affect neurodevelopment. However, mutations in SRCAP have been shown to cause the NDD Floating Harbour Syndrome (OMIM# 136140), which is characterized by intellectual and learning disabilities, a short stature, delayed osseous maturation, language deficits, and distinctive facial features [291, 295,296,297,298].
In addition to the INO80 family, another mechanism for histone exchange is suggested for the α-thalassemia X-linked mental retardation (ATRX) protein. ATRX is an ATP-dependent DNA translocase belonging to the Swi/Snf family of chromatin remodelers [299]. ATRX forms a complex with death domain associated protein (DAXX) [299, 300], and plays a critical role in the replication-independent deposition of the histone variant H3.3, functioning as a histone chaperone at specific genomic regions, including the telomeric domains [301, 302]. Furthermore, ATRX is involved in the suppression of several imprinted genes in the neonatal brain by promoting 3D chromatin structures via CTCF and cohesion [303]. In mice, germline deletion of Atrx has been shown embryonic lethal [304], whereas conditional deletion of Atrx in NPCs (Foxg1KiCre/+) caused a widespread cellular reduction in both the neocortex and hippocampus resulting in a significant smaller forebrain size [305]. In addition, AtrxFoxg1Cre mice show excessive DNA damage caused by DNA replication stress and subsequent Tp53-dependent apoptosis [306, 307]. On a cellular level, these AtrxFoxg1Cre mice show a reduction in precursor cell number and abnormal migration of progenitors in the hippocampus and the upper layers of the cortex [305, 306]. Furthermore, fewer Neuropeptide Y (NPY), SST and cholecystokinin (CCK) expressing GABAergic neurons were generated in the ventral telencephalon [306]. In humans, mutations in ATRX cause the rare congenital X-linked disorder ATRX syndrome (OMIM# 301040), characterised by moderate to severe ID, DD, microcephaly, hypomyelination, and a mild form of a-thalassemia [308].
To summarize, chromatin remodelers are highly expressed in neural progenitors, and are essential to dynamically activate, repress, or poise gene expression during the transition from RGCs to glutamatergic neurons or glia (Fig. 2, Table 1). Epigenetic modulation in glutamatergic neuron maturation is reviewed in detail elsewhere [135], however in the next section we want to highlight the maturation a specialized glutamatergic subpopulation that is frequently impacted in NDDs, called callosal projection neurons.
Epigenetic modulation in callosal projection neuron development
The corpus callosum is a critical link between the two cortical hemispheres. The developmental mechanisms for callosal projections have been well researched in mice (reviewed in [309, 310]), and abnormalities of the corpus callosum often feature in human NDDs, especially ID and epilepsy [311] but also in Coffin-Siris Syndrome [312]. During mouse brain development, the cortical hemispheres fuse along the midline around E16 [313], aided by specialized glia populations called midline zipper glia and indusium griseum glia. Those establish the glial wedge to both sides, as well as the bridge-like subcallosal sling. Subsequently, callosal-projecting cortical pyramidal neurons start projecting axons across the midline and connect to their homotopic cortical area. Those projection neurons mostly reside in the upper cortical layers, and their callosal-projecting identity is under direct epigenetic control.
In newly generated pyramidal cell precursors, the transcription factor Ctip2 specifies a subcortical-projecting fate, and is normally expressed in layer 5 neurons. In contrast, in future upper-layer callosal-projecting neurons Ctip2 is repressed by de-acetylation of H4K12 at its promoter region via NuRD complex and HDAC1 recruitment by the DNA-binding protein SATB2 [314,315,316]. After fate specification in the early postnatal period, HDAC1 is gradually removed from the Ctip2 promoter by the transcription factor LMO4, leading to re-establishment of H4K12ac and consequentially re-expression of Ctip2 in a subset of upper-layer neurons [317, 318].
Several other mouse models presented with deficits of callosal projections. For example in Cbp knockout mice (Rubinstein–Taybi Syndrome) a reduced size of the corpus callosum was found [319], similar to the phenotype of mutants for the chromatin remodelling complex members Prdm8 and HP1γ [320, 321]. Furthermore, knockdown of the chromatin remodeler Chd8 in the neocortex impaired dendrite and axonal growth and branching of upper-layer callosal projection neurons, and resulted in delayed migration of cortical neurons at E18.5, as the majority of labelled cells remained in the VZ/SVZ [322]. Moreover, mutations in the ISWI complex member Smarca5 cause partial agenesis of the corpus callosum, specifically due to reduced generation of viable upper-layer pyramidal neurons during mid-neurogenesis [271]. Lastly, a mouse model for Arid1b and Smarcb1 deficits (Coffin-Siris Syndrome) indeed mirror the human phenotype, as Arid1b+/− and Smarcb1+/inv NesCre+/− mice also have a significantly reduced corpus callosum thickness [277, 282]. Brain-specific Smarcb1+/− mice showed agenesis of the corpus callosum due to midline glia defects, similar to human CSS patients with mutations in SMARCB1, SMARCE1 and ARID1B [277]. In a human patient cohort with non-syndromic callosal abnormalities, mutations in ARID1B were found to be the most common cause, accounting for 10% of all cases [323].
During the axonal crossover, a multitude of axon guidance factors are required, and defects in the expression of those factors can also cause callosal projection deficits. A recent study described that chromatin remodelling of the axon guidance cue Sema6a caused the callosal defects observed in WAGR Syndrome (OMIM# 194072), a complex disorder including aniridia, kidney tumours, genital abnormalities, and ID [324]. Specifically, this study identified a novel protein, C11orf46/ADP ribosylation factor like GTPase 14 effector protein (ARL14EP), of which mutations were previously associated with ID [325]. C11orf46 is a member of the SETDB1-KRAB associated protein (KAP1)-MCAF1 chromatin repressor complex, and controls H3K9 methylation levels at the Sema6a promoter, cell-autonomously in projection neurons [324]. The callosal projection phenotype could be rescued by targeted H3K9 re-methylation at the Sema6a locus, indicating a direct epigenetic repressive control over axon guidance receptors in callosal-projecting neurons.
Epigenetic modulation in GABAergic neuron development
Besides glutamatergic excitatory neurons, the mammalian neocortex contains between 12 and 20% GABAergic inhibitory neurons [326, 327]. Defects in GABAergic neurons feature prominently in NDDs, in for example epilepsy, Schizophrenia, and ASD (reviewed in [328]). Broadly, GABAergic neurons can be subdivided according to the expression of marker genes Parvalbumin (PV +), Somatostatin (SST +), and Serotonin Receptor 3a (5-HT3aR +) [329, 330]. Within each of those three large groups, further subdivisions can be made according to gene expression [331, 332], morphology, and electrophysiological firing parameters [333, 334], with current estimates ranging from 20 to 60 subdivisions [335].
The mechanism of GABAergic neuron generation is comparatively well-conserved between mice and humans [336, 337]. In mice, GABAergic neurons are generated in subdivisions of the Ganglionic Eminences (GEs: Medial GE (MGE), Lateral GE (LGE), Caudal GE (CGE), and Preoptic Area (POA)), which are temporary proliferative zones at the site of the future Striatum. In contrast to the developing cortical plate, the precursors in the GE do form a SVZ, but are not all anchored to the basal membrane, and this population is massively expanded in the human GEs [336]. The precursors divide asymmetrically to produce future GABAergic cells, which gather in the mantle zone and migrate in two morphogen-directed streams towards the cortical plate [338, 339]. The MGE and POA express the transcription factor Nkx2-1 and produce the majority of PV + , and SST + neurons [338, 340] (Fig. 2). In contrast, VIP + neurons (the largest subset of 5-HT3aR + neurons) are produced in the CGE (see for reviews: [338, 341, 342]). Although the networks of transcription factors that define cellular identities during GABAergic neuron development and migration are comparatively well-researched [341], data on epigenetic regulation and especially chromatin remodelers is scarce and mostly has been inferred from other cell types including cancer biology and other neuron classes [343].
The first evidence for involvement of chromatin remodelers in GABAergic neuron production was reported in mice with a knockout for the histone acetyltransferase Kat6b/Querkopf, where a reduced density of GAD67 + (GABAergic) neurons in the cortex was found [344]. Years after the initial mouse study, mutations in human KAT6B were found to cause Ohdo/SBBYS syndrome (OMIM# 603736) [345, 346], however the initial findings regarding GABAergic neuron density have not been followed up to date. Mutations in the related KAT6A histone acetyltransferase were also found to cause ID and craniofacial dysmorphism [347, 348], recently described as KAT6A Syndrome [349]. Mouse studies have reproduced a craniofacial phenotype via Hox gene regulation [350], however neurodevelopmental phenotypes have not yet been studied. We do have a more complete picture for the ATP-dependent chromatin remodeler CHD2, which in humans is associated with epilepsy and broad-spectrum NDDs as described above [351]. Specifically, Chd2 transcription is found to be activated in MGE/POA progenitors by the transcription factor NKX2-1, and by doing so the CHD2 protein in turn colocalizes with NKX2-1 on its downstream targets, illustrating the feedback loops in which chromatin remodelers act [352]. Chd2+/− mice display a marked reduction in MGE-derived GABAergic neuron production, which results in a reduced PV + /SST + GABAergic neuron count in the cortex [202]. The functional consequences (defects in inhibitory synaptic transmission, altered excitatory/inhibitory balance, and behavioural abnormalities) were rescued by an embryonal transplantation of MGE-derived GABAergic neurons, which indicates that already a reduced GABAergic neuron production can produce profound circuit abnormalities [202].
Haploinsufficiency of the epigenetic regulator ARID1B, which we previously discussed as the causal gene for Coffin-Siris Syndrome, was found to cause premature apoptosis in MGE precursors in mice. As a result, Arid1b+/− mice show a reduced production of MGE-derived (PV + and SST +) GABAergic neurons, and altered laminar arrangement of PV + and SST + neurons in the cortex (Fig. 2) [353]. Mechanistically, the same study found a general reduction of the permissive histone mark H3K9ac3 at the Pvalb promoter in Arid1b+/− mice, resulting in reduced PV transcription throughout development into the adult cortex [353]. Conditional Arid1b knockout in specific GABAergic neuron population showed an interesting bifurcation of effects, as PV-specific Arid1b haploinsufficiency led to reduced mobility and social deficits, while SST-specific Arid1b haploinsufficiency led stereotyped behaviour such as excessive grooming [354].
Also mutations in the histone acetyltransferase CBP (Rubinstein–Taybi Syndrome), have been implicated in GABAergic precursor generation [355]. Constitutive heterozygous Cbp knockout mice show a transient impairment in GABAergic neuron formation in vivo [356]. Using a more direct approach, region-specific Cbp knockout in the developing MGE reduces the number of PV + and SST + neurons in the cortex and results in a prominent seizure phenotype [357], indicating that epigenetic regulation by CBP is directly required for proper cell-type specification of inhibitory GABAergic neurons. These results were also confirmed outside of the cortex in non-cortical areas, as conditional knockout of CBP in cerebellar progenitors lead to cerebellar hypoplasia and altered morphology of the cerebellum in both mice (hGFAPCre::CrebbpFl/Fl P25) [358] and patients [359]. On a cellular level, conditional knockout of CBP in granule cell progenitors altered cerebellar foliation as a result of loss of glial endfeet on the pial surface by Bergmann glia fibers and abnormal Purkinje cells arborisation [358].
After the formation of GABAergic precursors, the immature GABAergic neurons migrate tangentially following two morphogen-directed streams along the developing cortical plate, where they subsequently invade the cortex in the late stages of corticogenesis between E19 and P4 [341, 360, 361]. The exact place and time for programming the subdivisions within PV + , SST + and 5-HT3aR + is an area of active debate, with different hypotheses highlighting programming at the progenitor stage, during migration to the cortex, or only by local factors in the cortical plate. A recent study indicates that for MGE-derived neurons, the subtype is determined prior to migration, and instructs the migratory route and the place of integration in the cortex [362]. Specifically, the SST + subgroup of Martinotti cells and the PV + subgroup of translaminar PV + neurons preferentially migrate through the Marginal Zone, along the outer side of the developing cortical plate [362]. Migrating GABAergic neurons sense a multitude of environmental cues and integrate them to gene expression patterns. Similarly to GABAergic neuron progenitors, the cascades of transcription factors in migrating GABAergic neurons are comparatively well-characterized, but epigenetic modulations have only recently come into focus (see for review [343]). A recent series of studies investigated cortical GABAergic neurons derived from the POA, which produces subgroups of SST + , PV + and Reelin + GABAergic neurons [341]. Specifically, POA-derived GABAergic neurons suppress the expression of the transcription factor Pak6 during migration via a non-canonical recruitment of the PRC (specifically EZH2) by DNMT1 to the Pak6 promoter [363, 364]. In POA-specific Dnmt1-knockout mice, the repressive mark H3K27me3 is reduced around the Pak6 transcription start site, leading to precocious expression of Pak6 during migration and consequentially precocious activation of a post-migratory genetic program. As a result, a large portion of POA-derived neurons undergo apoptosis before reaching their cortical destination in POA-specific Dnmt1-knockout mice [363].
After migrating to the cortex, GABAergic neurons distribute throughout the layers, in a cell-type and area-specific pattern. Broadly speaking, PV + and SST + neurons predominate in the mid- to lower layers, whereas 5-HT3aR + are predominant in layer 1 [365] and (the VIP + subset) in layer 2/3 [366]. Primary sensory areas contain a higher density of PV + neurons, while the areas at the edge of the cortical plate contain a higher density of SST + neurons [366]. Once the GABAergic precursors are located at the appropriate cortical area and laminar location, they integrate into the local circuitry as it develops [367, 368]. The SST + GABAergic neurons mature relatively early, around the same time as the excitatory neurons in the same circuit [362, 369]. However, PV + GABAergic neurons mature much slower, and are dependent on external inputs that activate the local circuit for a successful maturation. The activity levels need to be translated to gene expression patterns, and while no complete mechanism is currently known, the high number of PV + neuron maturation dysfunctions caused by mutations in chromatin remodelers is indicative of a tight epigenetic control over this process [370]. One example is the maturation of PV + neurons in MeCP2+/− mice, which is the primary cause for Rett Syndrome (OMIM# 312750) in humans [371,372,373,374,375]. It is characterized by arrested development between 6 and 18 months of age, regression of acquired skills, loss of speech, stereotypic movements (classically of the hands), microcephaly, seizures, and mental retardation. In MeCP2+/− mice, the lack of MECP2 leads to a premature maturation of PV + cells including marker expression, morphology, and synaptic properties [372, 376]. MECP2 directly binds to the promoter regions of Pvalb and Gad1, the rate-limiting GABA synthesizing enzyme [377, 378]. Also at the adult level, neuronal activity regulates the expression of PV in a dynamic manner [379], a phenotype which is also impaired in PV + neurons of MeCP2+/− mice [380], indicating an epigenetic component to the integration of the activity-dependent signal. In contrast, haploinsufficiency of the histone methyltransferase Ehmt1 (Kleefstra Syndrome) causes delayed maturation of PV + neurons in the mouse sensory cortices, consisting of delayed PV expression and PNN generation, as well as reduced GABAergic neurotransmission [381]. Besides neuronal activity levels, PV + neurons also integrate morphogenic signals such as the released transcription factor Otx2. OTX2 is not produced in the cortex, but rather released by thalamic afferents and the Choroid Plexus [382,383,384,385]. OTX2 is taken up by the future PV + neurons, where it upregulates the expression of Gadd45b/g, two DNA demethylases which then mediate the up/downregulation of large sets of genes necessary for the maturation to full PV + cells, including Pvalb itself [382].
Epigenetic modulation during glia development and function
At the end of the neurogenic period, cortical RGCs cells switch to glial production and generate a vast number of astrocytes and oligodendrocytes [133]. In the mouse cortex, astrocytes are first detected around E16 and oligodendrocytes around birth; however, the vast majority of both cell types are produced during the first month of postnatal development. Cre-loxP lineage tracing showed that oligodendrocytes in the cerebral cortex are produced at different sites outside of the cortex depending on the developmental stage [386]. The first wave of production begins around E12.5 in the MGE and anterior entopeduncular area. The second wave begins around E15 from in the LGE and CGE, and finally, local production begins in the cortical SVZ around birth (Fig. 2) [387]. Similar to oligodendrocytes, astrocytes can be both generated from dividing RGCs [388], from the postnatal SVZ [389], or locally by self-amplification in the postnatal cortex [118] (for detailed information about the origin and specification of glia see [390,391,392,393]). Glia play crucial roles in CNS homeostasis [394], including synaptic glutamate uptake [395], synaptogenesis [396], maintenance of extracellular potassium [397], nutrient support for neurons [398], the formation of ECM molecules [399, 400] and many other processes.
Studies investigating the role of chromatin remodelling in mouse models of NDDs have primarily focussed on the alteration of neuronal network function. Recent advances in our understanding of astrocyte function have led to the emerging concept that primary astrocyte dysfunction alone is sufficient to drive the complex behavioural phenotypes observed in some cases of NDDs. As described earlier, RGCs undergo chromatin remodelling in response to various extracellular cues to enable the accessibility of neurogenic or gliogenic genes. Loss of the H3K9 methylase Setdb1 in mice has been shown to reduce H3K9me3 occupancy at the Gfap promotor, resulting in enhanced astrogenesis and accelerated differentiation (Fig. 2) [147]. Furthermore, EHMT1 has been shown to play a role in DNMT1-mediated DNA methylation via UHRF1/LIG1 interaction [401], which implies that astrocytes might contribute to the neuronal phenotypes in SETDB1-associated disorders or Kleefstra syndrome. Both SETDB1 and EHMT1 have recently been described to coexist in the same complex together with EHMT2 and SUV39H1 [402], revealing the interesting hypothesis that this complex plays an important role in the neurogenic-to-gliogenic switch, and any dysfunction in any of these genes will lead to a convergent phenotypic outcome.
Epigenetic regulation by the Polycomb Repressor Complex (PRC) has also been described to play a role in the differentiation from NPCs to astrocytes and oligodendrocytes. Acute deletion of the PRC1 component Ring1B or Ezh2 at E12.5 in mice prolonged the neurogenic phase and delayed the astrogenic phase in cultures of neocortical NPCs [136]. In contrast, another report found that cerebral specific loss of Ezh2 in the Emx1-Cre mice accelerated gliogenesis and glial differentiation at P0 [129]. Furthermore, overexpression of Ezh2 in postmitotic astrocytes in turn lead to a downregulation of pro-astrocytic genes S100b and GFAP, whereas an increase in progenitor like genes like SOX2 and CD133 was found [403]. Similarly, a small population of specialized neurogenic astrocytes that resides in the SVZ and survives into adulthood expresses Ezh2, which is required for those astrocytes to keep their neurogenic potential [404]. These results indicate that the PRC associated proteins are essential for promoting the onset of the astrocytic differentiation of NPCs during neocortical development.
Mature glia function has been studied widely in models of NDD (including Noonan syndrome [405], Neurofibromatosis-1 [406], Costello syndrome [399, 407], Cardiofaciocutaneous syndrome [408], Fragile X syndrome [409], Alexander disease [410], and Tuberous Sclerosis Complex [411]) however only in few models of deficient chromatin remodelling. One example is a mouse model for MeCP2 deficiency. Aside from the clear neuronal phenotype found in these mouse models, co-culture studies showed that secreted factors by Mecp2−/− mouse astrocytes significantly affect the development of wild type hippocampal neurons in a non-cell autonomous manner, as was visualised by altered dendrite morphology [412]. Furthermore, neuronal phenotypes found in co-culture with Mecp2−/− astrocytes appear to be dependent upon the expression of astroglial gap-junction protein Connexin-43 (Cx-43), as blocking Cx-43 restored this phenotype [413]. Mecp2−/− mouse astrocytes also showed an increased expression of astroglial marker genes Gfap and S100β and abnormal glutamate clearance [414]. Interestingly, selective restoration of MECP2 in astrocytes in vivo using the Cre-loxP recombination system significantly improves locomotion and anxiety levels, and restores respiratory abnormalities to a normal pattern [415]. At the cellular level, re-expressed MECP2 in astrocytes also restores normal neuronal dendritic morphology [415]. Similar to these findings, an increased expression of GFAP and CX-43 proteins was found in the superior frontal cortex in a cohort of ASD patients [416]. Furthermore, increased levels of H3K9me3 occupancy at the promotor of the gap junction proteins Cx-30 and Cx-43 have been found in cortical and subcortical regions of patients with MDD [417]. This cohort consisted of patients expressing extremely low levels of pro-astrocytic genes GFAP, ALDH1L1, SOX9, GLUL, SCL1A3, GJA1, and GJB6 [418], proposing a possible role for the H3K9 methylases SETDB1 and SUV39H1 in mature astrocyte function and CX-43 expression [417].
CHD8 is another example of a chromatin remodeler that plays a role in glia function. Recent studies show cell-type specific Chd8 deletion in OPCs results in myelination defects in mice [243]. In addition to altered myelination, conditional knockout of Chd8 in OPCs (Olig1-Cre;Chd8fl/fl) has been shown to slow down action potential propagation as a result of impaired myelination, leading to deficits including increased social interaction and anxiety-like behaviour as similar to Chd8 heterozygous mutant mice [244] and behavioural phenotypes found in patients.
Heterozygous loss of Smarca4 (Brg1fl/fl, Nestin-cre) was furthermore shown to cause precocious neuronal differentiation before the onset of gliogenesis [419]. This resulted in a significant reduction of astrocyte and oligodendrocyte differentiation in these animals, suggesting Smarca4 controls the switch from neurogenesis to gliogenesis [419]. Furthermore, SMARCA4 is known as a mediator of long-range interactions of enhancer regions and TTSs [420], and by doing so is involved in the STAT3 dependent [421] inter-chromosomal gene clustering of Gfap and Osmr resulting in transcriptional enhancement of these genes [422]. Interestingly, loss of SMARCA2 in SMARCA2K755R/+, and SMARCA2R1159Q/+ NPCs resulted in a reduction of Smarca4 mRNA expression, together with an increased and functionally active binding to chromatin [423]. These results suggested that mutations in SMARCA2 result in global retargeting of SMARCA4, which was shown to drive de novo activation of enhancers and upregulation of astrocyte genes [423].
To summarize, current evidence shows that chromatin remodelers play a role in the development, migration and circuit integration of each of the major cortical cell classes: Glutamatergic and GABAergic neurons and glia. Consequentially, failures of chromatin remodelling can impact the development of each of those cell types, resulting in a lasting impairment in cellular function.
Future perspectives
In this review, we detailed the contribution of chromatin remodelers in different neural cell-classes during the multiple stages of the developmental continuum. Chromatin remodelers are crucial parts of a cell’s information processing machinery, by integrating external and internal signals into gene expression patterns. Developing neurons inhabit an extraordinarily complex epigenetic landscape, and events such as cell-type specification are under tight epigenetic control [424]. Consequentially, defects in chromatin remodelling will lead to a relaxation of that epigenetic control, causing for example premature neural differentiation at the expense of progenitor pool expansion [208].
As chromatin remodelers have such a variety of functions in different cell types, timepoints and at specific genetic loci, a full picture requires concurrent measurements at several levels simultaneously—a task that current technologies are only starting to address. We see the potential for progress in the following fields:
-
1)
Understanding chromatin remodeler locus specificity: Chromatin modifications are site-specific on the genome level, such as histone methylation at the activity-dependent Bdnf exon IV [425]. However, until recently, to study this site-specific targeting one had to rely on the cell’s innate targeting abilities. Coupling catalytic subunits to a precise targeting protein allows artificial induction of locus-specific chromatin modifications. One example is the dCas9-SunTAG method [143, 426], where a genetic locus is tagged via dCas9 and gRNAs. Subsequently, local chromatin is modified by a chromatin modifier’s catalytic subunit targeted towards the tag. The ability to induce chromatin modifications at specific genomic sites will improve our understanding of the regulatory networks in gene expression, for example during cell fate specification.
-
2)
Understanding the role of chromatin remodeler presence in complexes: Chromatin modifiers exist in complexes that dynamically assemble, disassemble, and bind to chromatin at different locations. Complexes are hypothesized to differ between different locations (or time points), however those have proven difficult to investigate with classic immunolabelling techniques. Recent advances in spatial proximity labelling, such as promiscuous biotinylation targeted via dCas9 [427], allow for a precise snapshot of protein complexes assembled in spatial proximity to a single genomic region. Importantly, this technique can be applied in living cells and in vivo in the developing brain [428], making it applicable to the neurodevelopmental questions that we have detailed here. This technique allows detailed insights site-specific complex dynamics, a largely unexplored feature of the genetic landscape.
-
3)
Chromatin remodelling temporal specificity: Neuronal specification is thought to be a series of tightly controlled gene expression (and hence epigenetic regulation) states. For glutamatergic neuron generation in the cortex, recent evidence points to a stochastic generation of different subtypes [121, 429], however it is currently unknown whether GABAergic neuron generation is controlled in a similar way [361]. Classic labelling techniques such as BrdU were only able to identify neurons born within approximately 12 h from each other, which is slower than the hypothesized changes in genetic expression state. Recently developed labelling techniques such as FlashTag selectively label neurons born in a 2-h window in vivo, leading to a more precise identification of the transcriptional program controlling glutamatergic neuron specification [424, 430]. Application of the same technique for GABAergic neurons might deliver interesting insights into subtype specification as well.
-
4)
Measuring cell-type specificity: The classification of the brain’s cells has been controversial since the start of neuroscience as a field. For example, GABAergic neurons and glia have long resisted simple classification [431, 432]. However, recent large-scale single-cell RNA sequencing studies [331, 332, 433,434,435] attempt to map the cellular diversity of brain from the bottom up. Furthermore, studies measuring multiple modalities on the same neurons promise a unification of classifications from single-cell electrophysiology, morphology and RNA sequencing (Patch-Seq), and have delivered insights in glutamatergic [436] and GABAergic neuron populations [333]. Especially when coupled with advanced analysis techniques [334], those large datasets might soon be available as a “reference classifier” that experimental data can be compared with, similarly to reference atlases in neuroanatomy or reference genomes in genomics.
-
5)
Identification of converging molecular pathways for therapeutic interventions: Functional interactions between several NDD related chromatin remodelers and their regulatory proteins has been shown to converge on a shared transcriptional axis [156, 287, 437,438,439]. One example is the H3K4 demethylase KDM5C, whose expression is controlled by three regulatory proteins: ARX, ZNF711 and PHF8. All four of those genes are located on the X chromosome, and consequentially mutations in any of these four genes are associated with X-linked NDDs [440,441,442]. Interestingly, loss of Arx caused a significant reduction of Kdm5c expression and neuronal maturation in C. Elegans and mice, which could be restored using the HDAC inhibitor SAHA [438]. These findings imply that chromatin remodelers function in closely coupled transcriptional networks, with mutations in genes in the same cluster producing overlapping NDD phenotypes [439, 443]. As demonstrated in the case of KDM5C, overlapping regulatory pathways might be used as drug targets, and mutations in shared pathways could prove to be relatively easily identifiable biomarkers [444, 445]. In a promising first step, several groups have shown that chemical inhibition of HDACs can successfully rescue behavioural phenotypes in mouse models of NDDs [116, 438, 444,445,446,447]. Furthermore, the fact that those pathways tend to be well-conserved might prove valuable in translation to clinical practice.
The studies summarized in this review were almost exclusively performed in animal models of NDDs. While mice have many advantages as model organisms and many features are conserved down to the cellular level [331], some features appear to be unique to the human lineage, for example specialized cell-types such as outer radial glia cells [448], subpial interlaminar astrocytes [449], or the recently described rosehip neurons [450]. Furthermore, some time periods in neurodevelopment are much longer in comparison, elongating the vulnerable periods for many regulatory processes in humans. Therefore, using mouse brains as the sole model we might overlook important human-specific aspects of brain development and NDD pathogenesis. Although the use of animal models will remain essential to study complex developmental processes like cortical layering or migration, human induced pluripotent stem cells (hiPSCs) offer a higher-throughput model to investigate the developmental continuum from the earliest point of progenitor specification until the formation of neuronal circuits in vitro. For this reason, the use hiPSCs has gained a lot of attention recent years in the field of NDD research. HiPSCs provide an unlimited pool of (patient) material, which can be differentiated into neuronal networks, and can be monitored over development in vitro. In addition, these cells are comparatively easy to manipulate using for example CRISPR-Cas9 genome editing, and therefore can be used as a high throughput tool to study genotype–phenotype correlations in a controlled environment [451, 452]. Moreover, patient specific hiPSCs carry the same genetic background as the patient, which allows the study of polygenic disorders like ASD or Schizophrenia that cannot be modelled using animal models.
Protocols for the differentiation of hiPSCs into 3D cerebral organoids are becoming increasingly popular as these models have been shown to resemble the complex developmental programs of early corticogenesis during the first and second trimester of human foetal development [453, 454]. Indeed, 3D cerebral organoids derived from patients with severe microcephaly as a result of CDK5RAP2 mutations showed reduced neuroepithelial differentiation, fewer and smaller progenitor regions, and premature neuronal differentiation [455]. Furthermore, 3D human organoids from idiopathic ASD patients showed reduced proliferation of progenitors, increased neurogenesis, and an imbalance between the production of glutamatergic and GABAergic neurons [456]. Moreover, organoids derived from CNTNAP2+/− hiPSCs showed increased organoid volumes as a result of increased proliferation of progenitors, which in turn expanded the neuronal population [457]. Recent work has shown that patient-derived iPSC organoids with copy number variants in the ASD risk locus 16p11.2 mirror the patient’s micro/macrocephaly phenotype [458]. Similarly, RAB39b loss in 3D organoids has recently been shown to cause hyperproliferation and enlarged organoid size [459]. Studies are currently exploring organoid vascularization to further extend the development and complexity of these organoids [460,461,462], which will allow in the future to study more complex brain phenotypes using these in vitro approaches.
In summary, despite lots of progress in the field, the full influence of chromatin remodelling on neurodevelopment is currently unknown. To fully understand chromatin remodelers’ influence throughout the developmental continuum and identify possible human-specific pathways, future studies should combine human-specific in vitro models such as 3D cerebral organoids and well-characterized developmental models such as mice.
Availability of data and material
Correspondence and should be addressed to n.nadif@donders.ru.nl.
References
Ernst C (2016) Proliferation and differentiation deficits are a major convergence point for neurodevelopmental disorders. Trends Neurosci 39(5):290–299
Geschwind DH, Flint J (2015) Genetics and genomics of psychiatric disease. Science 349(6255):1489–1494
May PA et al (2018) Prevalence of Fetal Alcohol Spectrum Disorders in 4 US Communities. JAMA 319(5):474–482
Lange S et al (2017) Global Prevalence of Fetal Alcohol Spectrum Disorder Among Children and Youth: A Systematic Review and Meta-analysis. JAMA Pediatrics 171(10):948–956
Kaminen-Ahola N (2020) Fetal alcohol spectrum disorders: Genetic and epigenetic mechanisms. Prenat Diagn 40(9):1185–1192
An JY, Claudianos C (2016) Genetic heterogeneity in autism: From single gene to a pathway perspective. Neurosci Biobehav Rev 68:442–453
Wright CF, FitzPatrick DR, Firth HV (2018) Paediatric genomics: diagnosing rare disease in children. Nat Rev Genet 19(5):253–268
Gilissen C et al (2014) Genome sequencing identifies major causes of severe intellectual disability. Nature 511(7509):344–347
De Rubeis S et al (2014) Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515(7526):209–215
Iossifov I et al (2014) The contribution of de novo coding mutations to autism spectrum disorder. Nature 515(7526):216–221
Pinto D et al (2014) Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am J Hum Genet 94(5):677–694
Satterstrom FK et al (2020) Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180(3):568–584 (e23)
Ciptasari U, van Bokhoven H (2020) The phenomenal epigenome in neurodevelopmental disorders. Hum Mol Genet 29(R1):R42–R50
Tyagi M et al (2016) Chromatin remodelers: we are the drivers!! Nucleus 7(4):388–404
Hsieh J, Gage FH (2005) Chromatin remodeling in neural development and plasticity. Curr Opin Cell Biol 17(6):664–671
Lomvardas S, Maniatis T (2016) Histone and DNA modifications as regulators of neuronal development and function. Cold Spring Harb Perspect Biol 8(7):a024208
Gabriele M et al (2018) The chromatin basis of neurodevelopmental disorders: rethinking dysfunction along the molecular and temporal axes. Prog Neuropsychopharmacol Biol Psychiatry 84:306–327
Parenti I et al (2020) Neurodevelopmental disorders: from genetics to functional pathways. Trends Neurosci 43(8):608–621
Iwase S et al (2017) Epigenetic etiology of intellectual disability. J Neurosci 37(45):10773–10782
De Majo F, Calore M (2018) Chromatin remodelling and epigenetic state regulation by non-coding RNAs in the diseased heart. Non-coding RNA Res 3(1):20–28
Böhmdorfer G, Wierzbicki AT (2015) Control of chromatin structure by long noncoding RNA. Trends Cell Biol 25(10):623–632
Wei JW et al (2017) Non-coding RNAs as regulators in epigenetics (Review). Oncol Rep 37(1):3–9
Flemming W (1882) Zellsubstanz, Kern und Zelltheilung, ed. F.C.W. Vogel. Leipzig.
Luger K et al (1997) Characterization of nucleosome core particles containing histone proteins made in bacteria. J Mol Biol 272(3):301–311
Heitz E (1928) Das Heterochromatin der Moose. Bornträger.
Sadakierska-Chudy A, Filip M (2015) A comprehensive view of the epigenetic landscape Part II: Histone post-translational modification, nucleosome level, and chromatin regulation by ncRNAs. Neurotox Res 27(2):172–197
Clapier CR, Cairns BR (2009) The biology of chromatin remodeling complexes. Annu Rev Biochem 78:273–304
Davis L, Onn I, Elliott E (2018) The emerging roles for the chromatin structure regulators CTCF and cohesin in neurodevelopment and behavior. Cell Mol Life Sci 75(7):1205–1214
Zhang Y, Reinberg D (2001) Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev 15(18):2343–2360
Sterner DE, Berger SL (2000) Acetylation of histones and transcription-related factors. Microbiol Mol Biol Rev 64(2):435–459
Nowak SJ, Corces VG (2004) Phosphorylation of histone H3: a balancing act between chromosome condensation and transcriptional activation. Trends Genet 20(4):214–220
Shilatifard A (2006) Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu Rev Biochem 75:243–269
Nathan D et al (2006) Histone sumoylation is a negative regulator in Saccharomyces cerevisiae and shows dynamic interplay with positive-acting histone modifications. Genes Dev 20(8):966–976
Hymes J, Fleischhauer K, Wolf B (1995) Biotinylation of biotinidase following incubation with biocytin. Clin Chim Acta 233(1–2):39–45
Hassa PO et al (2006) Nuclear ADP-ribosylation reactions in mammalian cells: where are we today and where are we going? Microbiol Mol Biol Rev 70(3):789–829
Cuthbert GL et al (2004) Histone deimination antagonizes arginine methylation. Cell 118(5):545–553
Wang Y et al (2004) Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 306(5694):279–283
Nelson CJ, Santos-Rosa H, Kouzarides T (2006) Proline isomerization of histone H3 regulates lysine methylation and gene expression. Cell 126(5):905–916
Kim E et al (1997) Deficiency of a protein-repair enzyme results in the accumulation of altered proteins, retardation of growth, and fatal seizures in mice. Proc Natl Acad Sci USA 94(12):6132–6137
Tan M et al (2011) Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146(6):1016–1028
Chen Y et al (2007) Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol Cell Proteomics 6(5):812–819
Farrelly LA et al (2019) Histone serotonylation is a permissive modification that enhances TFIID binding to H3K4me3. Nature 567(7749):535–539
Lepack AE et al (2020) Dopaminylation of histone H3 in ventral tegmental area regulates cocaine seeking. Science 368(6487):197–201
Tan M et al (2014) Lysine glutarylation is a protein posttranslational modification regulated by SIRT5. Cell Metab 19(4):605–617
Wagner GR et al (2017) A class of reactive Acyl-CoA species reveals the non-enzymatic origins of protein acylation. Cell Metab 25(4):823-837.e8
Bao X et al (2019) Glutarylation of histone H4 Lysine 91 regulates chromatin dynamics. Mol Cell 76(4):660-675.e9
Zhang D et al (2019) Metabolic regulation of gene expression by histone lactylation. Nature 574(7779):575–580
Huang H et al (2018) Lysine benzoylation is a histone mark regulated by SIRT2. Nat Commun 9(1):3374
Wilson JP et al (2011) Proteomic analysis of fatty-acylated proteins in mammalian cells with chemical reporters reveals S-acylation of histone H3 variants. Mol Cell Proteom 10(3):110001198
Zou C et al (2011) Acyl-CoA:lysophosphatidylcholine acyltransferase I (Lpcat1) catalyzes histone protein O-palmitoylation to regulate mRNA synthesis. J Biol Chem 286(32):28019–28025
Webby CJ et al (2009) Jmjd6 catalyses lysyl-hydroxylation of U2AF65, a protein associated with RNA splicing. Science 325(5936):90–93
Zheng Q et al (2019) Reversible histone glycation is associated with disease-related changes in chromatin architecture. Nat Commun 10(1):1289
Galligan JJ et al (2018) Methylglyoxal-derived posttranslational arginine modifications are abundant histone marks. Proc Natl Acad Sci U S A 115(37):9228–9233
Galligan JJ et al (2014) Stable histone adduction by 4-oxo-2-nonenal: a potential link between oxidative stress and epigenetics. J Am Chem Soc 136(34):11864–11866
Jin J et al (2016) SIRT2 Reverses 4-Oxononanoyl Lysine Modification on Histones. J Am Chem Soc 138(38):12304–12307
Chen D et al (2013) Cigarette smoke component acrolein modulates chromatin assembly by inhibiting histone acetylation. J Biol Chem 288(30):21678–21687
Fang L et al (2016) Mechanisms underlying acrolein-mediated inhibition of chromatin assembly. Mol Cell Biol 36(23):2995–3008
Xu L et al (2015) Crosstalk of homocysteinylation, methylation and acetylation on histone H3. Analyst 140(9):3057–3063
Zhang Q et al (2018) Elevated H3K79 homocysteinylation causes abnormal gene expression during neural development and subsequent neural tube defects. Nat Commun 9(1):3436
Bustin M (1971) Nitration of the tyrosine in histone F1 in salt solutions and in F1-polyanion complexes. Biochim Biophys Acta 251(2):172–180
Prütz WA et al (1985) Reactions of nitrogen dioxide in aqueous model systems: oxidation of tyrosine units in peptides and proteins. Arch Biochem Biophys 243(1):125–134
Haqqani AS, Kelly JF, Birnboim HC (2002) Selective nitration of histone tyrosine residues in vivo in mutatect tumors. J Biol Chem 277(5):3614–3621
Gould N et al (2013) Regulation of protein function and signaling by reversible cysteine S-nitrosylation. J Biol Chem 288(37):26473–26479
Lee CF, Paull TT, Person MD (2013) Proteome-wide detection and quantitative analysis of irreversible cysteine oxidation using long column UPLC-pSRM. J Proteome Res 12(10):4302–4315
Akter S et al (2018) Chemical proteomics reveals new targets of cysteine sulfinic acid reductase. Nat Chem Biol 14(11):995–1004
García-Giménez JL et al (2013) Histone h3 glutathionylation in proliferating mammalian cells destabilizes nucleosomal structure. Antioxid Redox Signal 19(12):1305–1320
Zhou M, Paša-Tolić L, Stenoien DL (2017) Profiling of histone post-translational modifications in mouse brain with high-resolution top-down mass spectrometry. J Proteome Res 16(2):599–608
Bjornsson HT (2015a) The Mendelian disorders of the epigenetic machinery. Genome Res 25(10):1473–1481
Chandy M et al (2006) SWI/SNF displaces SAGA-acetylated nucleosomes. Eukaryot Cell 5(10):1738–1747
Hassan YI, Zempleni J (2006) Epigenetic regulation of chromatin structure and gene function by biotin. J Nutr 136(7):1763–1765
Ito T et al (2000) p300-mediated acetylation facilitates the transfer of histone H2A–H2B dimers from nucleosomes to a histone chaperone. Genes Dev 14(15):1899–1907
Reinke H, Horz W (2003) Histones are first hyperacetylated and then lose contact with the activated PHO5 promoter. Mol Cell 11(6):1599–1607
Bowman GD, Poirier MG (2015) Post-translational modifications of histones that influence nucleosome dynamics. Chem Rev 115(6):2274–2295
Ruthenburg AJ et al (2007) Multivalent engagement of chromatin modifications by linked binding modules. Nat Rev Mol Cell Biol 8(12):983–994
Wang Z, Patel DJ (2011) Combinatorial readout of dual histone modifications by paired chromatin-associated modules. J Biol Chem 286(21):18363–18368
Vallianatos CN et al (2019) Amelioration of brain histone methylopathies by balancing a Writer-Eraser Duo KMT2A-KDM5C. bioRxiv 2019:567917
Yu N-K, Baek SH, Kaang B-K (2011) DNA methylation-mediated control of learning and memory. Mol Brain 4(1):5
Okano M et al (1999) DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99(3):247–257
Bestor TH (2000) The DNA methyltransferases of mammals. Hum Mol Genet 9(16):2395–2402
Jones PL et al (1998) Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat Genet 19(2):187–191
Nan X et al (1998) Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393(6683):386–389
Williams SR et al (2010) Haploinsufficiency of MBD5 associated with a syndrome involving microcephaly, intellectual disabilities, severe speech impairment, and seizures. Eur J Hum Genet 18(4):436–441
Talkowski ME et al (2011) Assessment of 2q23.1 microdeletion syndrome implicates MBD5 as a single causal locus of intellectual disability, epilepsy, and autism spectrum disorder. Am J Human Genet 89(4):551–563
Bird A (2008) The methyl-CpG-binding protein MeCP2 and neurological disease. Biochem Soc Trans 36(Pt 4):575–583
Zeng Y, Chen T (2019) DNA methylation reprogramming during mammalian development. Genes 10(4):257
Lee TW, Katz DJ (2020) Hansel, gretel, and the consequences of failing to remove histone methylation breadcrumbs. Trends Genet 36(3):160–176
Greenberg MVC, Bourc’his D (2019) The diverse roles of DNA methylation in mammalian development and disease. Nat Rev Mol Cell Biol 20(10):590–607
Cerrato F et al (2020) DNA methylation in the diagnosis of monogenic diseases. Genes (Basel) 11(4):355
Sadikovic B et al (2019) DNA methylation signatures in mendelian developmental disorders as a diagnostic bridge between genotype and phenotype. Epigenomics 11(5):563–575
Schenkel LC et al (2016) DNA methylation analysis in constitutional disorders: clinical implications of the epigenome. Crit Rev Clin Lab Sci 53(3):147–165
Sadikovic B, Levy MA, Aref-Eshghi E (2020) Functional annotation of genomic variation: DNA methylation episignatures in neurodevelopmental Mendelian disorders. Hum Mol Genet 29(R1):R27–R32
Reilly J, Kerkhof J, Sadikovic B (2020) EpiSigns: DNA methylation signatures in mendelian neurodevelopmental disorders as a diagnostic link between a genotype and phenotype. Adv Mol Pathol 3:29–39
Aref-Eshghi E et al (2020) Evaluation of DNA methylation episignatures for diagnosis and phenotype correlations in 42 Mendelian neurodevelopmental disorders. Am J Human Genet 106(3):356–370
Aref-Eshghi E et al (2019) Diagnostic utility of genome-wide dna methylation testing in genetically unsolved individuals with suspected hereditary conditions. Am J Human Genet 104(4):685–700
Aref-Eshghi E et al (2018) Genomic DNA methylation signatures enable concurrent diagnosis and clinical genetic variant classification in neurodevelopmental syndromes. Am J Hum Genet 102(1):156–174
Krogan NJ et al (2003) A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol Cell 12(6):1565–1576
Murawska M, Brehm A (2011) CHD chromatin remodelers and the transcription cycle. Transcription 2(6):244–253
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat Rev Genet 2(4):292–301
Lieberman-Aiden E et al (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326(5950):289–293
Dixon JR et al (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485(7398):376–380
de Laat W, Duboule D (2013) Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502(7472):499–506
Phillips-Cremins JE et al (2013) Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153(6):1281–1295
Suhas Rao SP et al (2014) A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159(7):1665–1680
Xiao T, Wallace J, Felsenfeld G (2011) Specific sites in the C terminus of CTCF interact with the SA2 subunit of the cohesin complex and are required for cohesin-dependent insulation activity. Mol Cell Biol 31(11):2174–2183
Vietri Rudan M et al (2015) Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Reports 10(8):1297–1309
Schwalie PC et al (2013) Co-binding by YY1 identifies the transcriptionally active, highly conserved set of CTCF-bound regions in primate genomes. Genome Biol 14(12):R148
Mourad R, Cuvier O (2016) Computational identification of genomic features that influence 3D chromatin domain formation. PLoS Comput Biol 12(5):e1004908
Cournac A, Koszul R, Mozziconacci J (2015) The 3D folding of metazoan genomes correlates with the association of similar repetitive elements. Nucleic Acids Res 44(1):245–255
Melé M, John L (2016) Rinn, “Cat’s cradling” the 3D genome by the Act of LncRNA transcription. Mol Cell 62(5):657–664
Gorkin DU, Leung D, Ren B (2014) The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell 14(6):762–775
Bastle RM, Maze I (2019) Chromatin regulation in complex brain disorders. Curr Opin Behav Sci 25:57–65
Boukas L et al (2019) Coexpression patterns define epigenetic regulators associated with neurological dysfunction. Genome Res 29(4):532–542
Fahrner JA, Bjornsson HT (2019) Mendelian disorders of the epigenetic machinery: postnatal malleability and therapeutic prospects. Hum Mol Genet 28:R254–R264
Iwase S, Martin DM (2018) Chromatin in nervous system development and disease. Mol Cell Neurosci 87:1–3
Woodworth MB et al (2012) SnapShot: cortical development. Cell 151(4):918-918.e1
Greig LC et al (2013) Molecular logic of neocortical projection neuron specification, development and diversity. Nat Rev Neurosci 14(11):755–769
Nowakowski TJ et al (2016) Transformation of the radial glia scaffold demarcates two stages of human cerebral cortex development. Neuron 91(6):1219–1227
Ge W-P et al (2012) Local generation of glia is a major astrocyte source in postnatal cortex. Nature 484(7394):376–380
deAzevedo LC et al (2003) Cortical radial glial cells in human fetuses: Depth-correlated transformation into astrocytes. J Neurobiol 55(3):288–298
Kadhim HJ, Gadisseux JF, Evrard P (1988) Topographical and cytological evolution of the glial phase during prenatal development of the human brain: histochemical and electron microscopic study. J Neuropathol Exp Neurol 47(2):166–188
Llorca A et al (2019) A stochastic framework of neurogenesis underlies the assembly of neocortical cytoarchitecture. eLife 8:e51381
Awad S et al (2013) Mutation in PHC1 implicates chromatin remodeling in primary microcephaly pathogenesis. Hum Mol Genet 22(11):2200–2213
Beunders G et al (2013) Exonic deletions in AUTS2 cause a syndromic form of intellectual disability and suggest a critical role for the C terminus. Am J Hum Genet 92(2):210–220
Nagamani SCS et al (2013) Detection of copy-number variation in AUTS2 gene by targeted exonic array CGH in patients with developmental delay and autistic spectrum disorders. Eur J Hum Genet 21(3):343–346
Margueron R et al (2008) Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol Cell 32(4):503–518
Weaver DD et al (1974) A new overgrowth syndrome with accelerated skeletal maturation, unusual facies, and camptodactyly. J Pediatr 84(4):547–552
Gibson WT et al (2012) Mutations in EZH2 cause weaver syndrome. Am J Hum Genet 90(1):110–118
Choufani S et al (2020) DNA methylation signature for EZH2 functionally classifies sequence variants in three PRC2 complex genes. Am J Hum Genet 106(5):596–610
Pereira JD et al (2010) Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc Natl Acad Sci 107(36):15957–15962
Telley L et al (2019) Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364(6440):eaav522
Oberst P et al (2019) Temporal plasticity of apical progenitors in the developing mouse neocortex. Nature 573(7774):370–374
Molyneaux BJ et al (2007) Neuronal subtype specification in the cerebral cortex. Nat Rev Neurosci 8(6):427–437
Gao P et al (2014) Deterministic progenitor behavior and unitary production of neurons in the neocortex. Cell 159(4):775–788
Donega V et al (2018) Transcriptional dysregulation in postnatal glutamatergic progenitors contributes to closure of the cortical neurogenic period. Cell Rep 22(10):2567–2574
Amberg N, Laukoter S, Hippenmeyer S (2019) Epigenetic cues modulating the generation of cell-type diversity in the cerebral cortex. J Neurochem 149(1):12–26
Hirabayashi Y et al (2009) Polycomb limits the neurogenic competence of neural precursor cells to promote astrogenic fate transition. Neuron 63(5):600–613
Sparmann A et al (2013) The chromodomain helicase Chd4 is required for Polycomb-mediated inhibition of astroglial differentiation. Embo j 32(11):1598–1612
Zhao L et al (2015) Ezh2 is involved in radial neuronal migration through regulating Reelin expression in cerebral cortex. Sci Rep 5(1):15484
Morimoto-Suzki N et al (2014) The polycomb component Ring1B regulates the timed termination of subcerebral projection neuron production during mouse neocortical development. Development 141(22):4343–4353
Cukier HN et al (2012) The expanding role of MBD genes in autism: identification of a MECP2 duplication and novel alterations in MBD5, MBD6, and SETDB1. Autism Res 5(6):385–397
Xu Q et al (2016) Chromosomal microarray analysis in clinical evaluation of neurodevelopmental disorders-reporting a novel deletion of SETDB1 and illustration of counseling challenge. Pediatr Res 80(3):371–381
Jiang Y et al (2010) Setdb1 histone methyltransferase regulates mood-related behaviors and expression of the NMDA receptor subunit NR2B. J Neurosci 30(21):7152–7167
Jiang Y et al (2017) The methyltransferase SETDB1 regulates a large neuron-specific topological chromatin domain. Nat Genet 49(8):1239–1250
Ripke S et al (2014) Biological insights from 108 schizophrenia-associated genetic loci. Nature 511(7510):421–427
Cruvinel E et al (2014) Reactivation of maternal SNORD116 cluster via SETDB1 knockdown in Prader-Willi syndrome iPSCs. Hum Mol Genet 23(17):4674–4685
Zhu Y et al (2020) Epigenetic mechanism of SETDB1 in brain: implications for neuropsychiatric disorders. Transl Psychiatry 10(1):115
Ling BM et al (2012) Lysine methyltransferase G9a methylates the transcription factor MyoD and regulates skeletal muscle differentiation. Proc Natl Acad Sci USA 109(3):841–846
Sripathy SP, Stevens J, Schultz DC (2006) The KAP1 corepressor functions to coordinate the assembly of De Novo HP1-demarcated microenvironments of heterochromatin required for KRAB zinc finger protein-mediated transcriptional repression. Mol Cell Biol 26(22):8623–8638
Schultz DC, Friedman JR, Rauscher FJ 3rd (2001) Targeting histone deacetylase complexes via KRAB-zinc finger proteins: the PHD and bromodomains of KAP-1 form a cooperative unit that recruits a novel isoform of the Mi-2alpha subunit of NuRD. Genes Dev 15(4):428–443
Timms RT et al (2016) ATF7IP-mediated stabilization of the histone methyltransferase SETDB1 is essential for heterochromatin formation by the HUSH complex. Cell Rep 17(3):653–659
Minkovsky A et al (2014) The Mbd1-Atf7ip-Setdb1 pathway contributes to the maintenance of X chromosome inactivation. Epigenet Chromatin 7(1):12
Tian Z et al (2006) Expression of DNA methyltransferases in salivary adenoid cystic carcinoma and its association with the CpG islands methylation of tumor suppressor genes. Zhonghua Kou Qiang Yi Xue Za Zhi 41(7):411–415
Tachibana M et al (2002) G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev 16(14):1779–1791
Chen X et al (2012) G9a/GLP-dependent histone H3K9me2 patterning during human hematopoietic stem cell lineage commitment. Genes Dev 26(22):2499–2511
Wen B et al (2009) Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat Genet 41(2):246–250
Chen ES et al (2014) Molecular convergence of neurodevelopmental disorders. Am J Hum Genet 95(5):490–508
Olsen JB et al (2016) G9a and ZNF644 physically associate to suppress progenitor gene expression during neurogenesis. Stem Cell Rep 7(3):454–470
Schaefer A et al (2009) Control of cognition and adaptive behavior by the GLP/G9a epigenetic suppressor complex. Neuron 64(5):678–691
Balemans MCM et al (2014) Reduced euchromatin histone methyltransferase 1 causes developmental delay, hypotonia, and cranial abnormalities associated with increased bone gene expression in Kleefstra syndrome mice. Dev Biol 386(2):395–407
Willemsen MH et al (2012) Update on kleefstra syndrome. Mol Syndromol 2(3–5):202–212
Vermeulen K et al (2017) Adaptive and maladaptive functioning in Kleefstra syndrome compared to other rare genetic disorders with intellectual disabilities. Am J Med Genet A 173(7):1821–1830
Wang J et al (2010) CBP histone acetyltransferase activity regulates embryonic neural differentiation in the normal and Rubinstein-Taybi syndrome brain. Dev Cell 18(1):114–125
Yao T-P et al (1998) Gene dosage-dependent embryonic development and proliferation defects in mice lacking the transcriptional integrator p300. Cell 93(3):361–372
Lipinski M, Del Blanco B, Barco A (2019) CBP/p300 in brain development and plasticity: disentangling the KAT’s cradle. Curr Opin Neurobiol 59:1–8
Ateca-Cabarga JC et al (2015) Brain size regulations by cbp haploinsufficiency evaluated by in-vivo MRI based volumetry. Sci Rep 5:16256
Li L et al (2020) Lysine acetyltransferase 8 is involved in cerebral development and syndromic intellectual disability. J Clin Invest 130(3):1431–1445
Koolen DA et al (2012) Mutations in the chromatin modifier gene KANSL1 cause the 17q21.31 microdeletion syndrome. Nat Genet 44(6):639–641
Arbogast T et al (2017) Mouse models of 17q21.31 microdeletion and microduplication syndromes highlight the importance of Kansl1 for cognition. PLoS Genet 13(7):1006886
D’Mello SR (2019) Regulation of central nervous system development by class I histone deacetylases. Dev Neurosci 41(3):149–165
Hsieh J, Gage FH (2004) Epigenetic control of neural stem cell fate. Curr Opin Genet Dev 14(5):461–469
de Ruijter AJ et al (2003) Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J 370(Pt 3):737–749
MacDonald JL, Roskams AJ (2008) Histone deacetylases 1 and 2 are expressed at distinct stages of neuro-glial development. Dev Dyn 237(8):2256–2267
Montgomery RL et al (2009) Histone deacetylases 1 and 2 control the progression of neural precursors to neurons during brain development. Proc Natl Acad Sci USA 106(19):7876–7881
Hagelkruys A et al (2014) A single allele of Hdac2 but not Hdac1 is sufficient for normal mouse brain development in the absence of its paralog. Development 141(3):604–616
Yuniarti N et al (2013) Prenatal exposure to suberoylanilide hydroxamic acid perturbs corticogenesis. Neurosci Res 77(1–2):42–49
Ye F et al (2009) HDAC1 and HDAC2 regulate oligodendrocyte differentiation by disrupting the beta-catenin-TCF interaction. Nat Neurosci 12(7):829–838
Wagner VF et al (2019) A De novo HDAC2 variant in a patient with features consistent with Cornelia de Lange syndrome phenotype. Am J Med Genet A 179(5):852–856
Gao X et al (2018) A functional mutation in HDAC8 gene as novel diagnostic marker for cornelia de lange syndrome. Cell Physiol Biochem 47(6):2388–2395
Helgeson M et al (2018) Molecular characterization of HDAC8 deletions in individuals with atypical Cornelia de Lange syndrome. J Hum Genet 63(3):349–356
Deardorff MA, Porter NJ, Christianson DW (2016) Structural aspects of HDAC8 mechanism and dysfunction in Cornelia de Lange syndrome spectrum disorders. Protein Sci 25(11):1965–1976
Deardorff MA et al (2012a) HDAC8 mutations in cornelia de lange syndrome affect the cohesin acetylation cycle. Nature 489(7415):313–317
Bottai D et al (2018) Modeling Cornelia de Lange syndrome in vitro and in vivo reveals a role for cohesin complex in neuronal survival and differentiation. Hum Mol Genet 28(1):64–73
Katayama S et al (2018) HDAC8 regulates neural differentiation through embryoid body formation in P19cells. Biochem Biophys Res Commun 498(1):45–51
Harakalova M et al (2012) X-exome sequencing identifies a HDAC8 variant in a large pedigree with X-linked intellectual disability, truncal obesity, gynaecomastia, hypogonadism and unusual face. J Med Genet 49(8):539–543
Aoto T et al (2006) Nuclear and chromatin reorganization in the MHC-Oct3/4 locus at developmental phases of embryonic stem cell differentiation. Dev Biol 298(2):354–367
Gregor A et al (2013) De novo mutations in the genome organizer CTCF cause intellectual disability. Am J Hum Genet 93(1):124–131
Bastaki F et al (2017) Identification of a novel CTCF mutation responsible for syndromic intellectual disability—a case report. BMC Med Genet 18(1):68–68
Konrad EDH et al (2019) CTCF variants in 39individuals with a variable neurodevelopmental disorder broaden the mutational andclinical spectrum. Genet Med 21(12):2723–2733
Watson LA et al (2014) Dual effect of CTCF loss on neuroprogenitor differentiation and survival. J Neurosci 34(8):2860–2870
Hirayama T et al (2012) CTCF is required for neural development and stochastic expression of clustered Pcdh genes in neurons. Cell Rep 2(2):345–357
Deardorff MA et al (2007) Mutations in cohesin complex members SMC3 and SMC1A cause a mild variant of cornelia de Lange syndrome with predominant mental retardation. Am J Hum Genet 80(3):485–494
Deardorff MA et al (2012b) RAD21 mutations cause a human cohesinopathy. Am J Hum Genet 90(6):1014–1027
Krantz ID et al (2004) Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. Nat Genet 36(6):631–635
Vega H et al (2005) Roberts syndrome is caused by mutations in ESCO2, a human homolog of yeast ECO1 that is essential for the establishment of sister chromatid cohesion. Nat Genet 37(5):468–470
White JK et al (2013) Genome-wide Generation and Systematic Phenotyping of Knockout Mice Reveals New Roles for Many Genes. Cell 154(2):452–464
Lai AY, Wade PA (2011) Cancer biology and NuRD: a multifaceted chromatin remodelling complex. Nat Rev Cancer 11(8):588–596
Woodage T et al (1997) Characterization of the CHD family of proteins. Proc Natl Acad Sci USA 94(21):11472–11477
Guzman-Ayala M et al (2015) Chd1 is essential for the high transcriptional output and rapid growth of the mouse epiblast. Development 142(1):118–127
Gaspar-Maia A et al (2009) Chd1 regulates open chromatin and pluripotency of embryonic stem cells. Nature 460(7257):863–868
Pilarowski GO et al (2018) Missense variants in the chromatin remodeler CHD1 are associated with neurodevelopmental disability. J Med Genet 55(8):561–566
Shen T et al (2015) CHD2 is required for embryonic neurogenesis in the developing cerebral cortex. Stem Cells 33(6):1794–1806
Kim YJ et al (2018) Chd2 is necessary for neural circuit development and long-term memory. Neuron 0:1–14
Homem CC, Repic M, Knoblich JA (2015) Proliferation control in neural stem and progenitor cells. Nat Rev Neurosci 16(11):647–659
Thomas RH et al (2015) CHD2 myoclonic encephalopathy is frequently associated with self-induced seizures. Neurology 84(9):951–958
Study, D.D.D. (2017) Prevalence and architecture of de novo mutations in developmental disorders. Nature 542(7642):433–438
Chénier S et al (2014) CHD2 haploinsufficiency is associated with developmental delay, intellectual disability, epilepsy and neurobehavioural problems. J Neurodev Disord 6(1):9
Lamar KJ, Carvill GL (2018) Chromatin remodeling proteins in epilepsy: lessons from CHD2-associated epilepsy. Front Mol Neurosci 11:208
Nitarska J et al (2016) A functional switch of NuRD chromatin remodeling complex subunits regulates mouse cortical development. Cell Rep 17(6):1683–1698
Snijders Blok L et al (2018) CHD3 helicase domain mutations cause a neurodevelopmental syndrome with macrocephaly and impaired speech and language. Nat Commun 9(1):4619
Drivas TG et al (2020) A second cohort of CHD3 patients expands the molecular mechanisms known to cause Snijders Blok-Campeau syndrome. Eur J Hum Genet 28(10):1422–1431
Weiss K et al (2016) De novo mutations in CHD4, an ATP-dependent chromatin remodeler gene, cause an intellectual disability syndrome with distinctive dysmorphisms. Am J Hum Genet 99(4):934–941
Weiss K et al (2020) The CHD4-related syndrome: a comprehensive investigation of theclinical spectrum, genotype–phenotype correlations, and molecular basis. Genet Med 22(2):389–397
Kovač K et al (2018) Tumour-associated missense mutations in the dMi-2 ATPase alters nucleosome remodelling properties in a mutation-specific manner. Nat Commun 9(1):2112
Ragheb R et al (2020) Differential regulation of lineage commitment in human and mouse primed pluripotent stem cells by NuRD. bioRxiv 2020:2020.02.05.935544
Egan CM et al (2013) CHD5 is required for neurogenesis and has a dual role in facilitating gene expression and polycomb gene repression. Dev Cell 26(3):223–236
Thompson PM et al (2003) CHD5, a new member of the chromodomain gene family, is preferentially expressed in the nervous system. Oncogene 22(7):1002–1011
Potts RC et al (2011) CHD5, a brain-specific paralog of Mi2 chromatin remodeling enzymes, regulates expression of neuronal genes. PLoS ONE 6(9):e24515
Pisansky MT et al (2017) Mice lacking the chromodomain helicase DNA-binding 5 chromatin remodeler display autism-like characteristics. Transl Psychiatry 7(6):e1152
Mills AA (2017) The chromodomain helicase DNA-binding chromatin remodelers: family traits that protect from and promote cancer. Cold Spring Harb Perspect Med 7(4):a026450
Kalscheuer VM et al (2008) Disruption of the TCF4 gene in a girl with mental retardation but without the classical Pitt-Hopkins syndrome. Am J Med Genet A 146A(16):2053–2059
Yamada K et al (2010) Characterization of a de novo balanced t(4;20)(q33;q12) translocation in a patient with mental retardation. Am J Med Genet A 152A(12):3057–3067
Douet-Guilbert N et al (2015) A novel translocation (6;20)(q13;q12) in acute myeloid leukemia likely results in LMBRD1–CHD6 fusion. Leukemia Lymphoma 56(2):527–528
Kargapolova Y et al (2020) Overarching control of autophagy and DNA damage response by CHD6 revealed by modeling a rare human pathology. bioRxiv 2020:2020.01.27.921171
Hurd EA et al (2007) Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm Genome 18(2):94–104
Yao H et al (2020) CHD7 promotes neural progenitor differentiation in embryonic stem cells via altered chromatin accessibility and nascent gene expression. Sci Rep 10(1):17445
Marie C et al (2018) Oligodendrocyte precursor survival and differentiation requires chromatin remodeling by Chd7 and Chd8. Proc Natl Acad Sci USA 115(35):E8246-e8255
He D et al (2016) Chd7 cooperates with Sox10 and regulates the onset of CNS myelination and remyelination. Nat Neurosci 19(5):678–689
Feng W et al (2017) Chd7 is indispensable for mammalian brain development through activation of a neuronal differentiation programme. Nat Commun 8(1):14758
Jiang X et al (2012) The mutation in Chd7 causes misexpression of Bmp4 and developmental defects in telencephalic midline. Am J Pathol 181(2):626–641
Layman WS et al (2009) Defects in neural stem cell proliferation and olfaction in Chd7 deficient mice indicate a mechanism for hyposmia in human CHARGE syndrome. Hum Mol Genet 18(11):1909–1923
Lin AE, Siebert JR, Graham JM Jr (1990) Central nervous system malformations in the CHARGE association. Am J Med Genet 37(3):304–310
Yu T et al (2013) Deregulated FGF and homeotic gene expression underlies cerebellar vermis hypoplasia in CHARGE syndrome. Elife 2:e01305
Sanlaville D et al (2006) Phenotypic spectrum of CHARGE syndrome in fetuses with CHD7 truncating mutations correlates with expression during human development. J Med Genet 43(3):211–217
Gregory LC et al (2013) Structural pituitary abnormalities associated with CHARGE syndrome. J Clin Endocrinol Metab 98(4):E737–E743
Liu L et al (2014) A novel CHD7 mutation in a Chinese patient with CHARGE syndrome. Meta Gene 2:469–478
Vissers LELM et al (2004) Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet 36(9):955–957
Bernier R et al (2014) Disruptive CHD8 mutations define a subtype of autism early in development. Cell 158(2):263–276
Sugathan A et al (2014) CHD8 regulates neurodevelopmental pathways associated with autism spectrum disorder in neural progenitors. Proc Natl Acad Sci USA 111(42):E4468–E4477
Durak O et al (2016) Chd8 mediates cortical neurogenesis via transcriptional regulation of cell cycle and Wnt signaling. Nat Neurosci 19(11):1477–1488
Sood S et al (2020) CHD8 dosage regulates transcription in pluripotency and early murine neural differentiation. Proc Natl Acad Sci USA 117(36):22331–22340
Wang P et al (2017) CRISPR/Cas9-mediated heterozygous knockout of the autism gene CHD8 and characterization of its transcriptional networks in cerebral organoids derived from iPS cells. Mol Autism 8(1):11
Sakamoto I et al (2000) A novel beta-catenin-binding protein inhibits beta-catenin-dependent Tcf activation and axis formation. J Biol Chem 275(42):32871–32878
Zhao C et al (2018) Dual requirement of CHD8 for chromatin landscape establishment and histone methyltransferase recruitment to promote CNS myelination and repair. Dev Cell 45(6):753-768.e8
Kawamura A et al (2020) Oligodendrocyte dysfunction due to Chd8 mutation gives rise to behavioral deficits in mice. Hum Mol Genet 29(8):1274–1291
Nishiyama M et al (2009) CHD8 suppresses p53-mediated apoptosis through histone H1 recruitment during early embryogenesis. Nat Cell Biol 11(2):172–182
Platt RJ et al (2017) Chd8 mutation leads to autistic-like behaviors and impaired striatal circuits. Cell Rep 19(2):335–350
Suetterlin P et al (2018) Altered neocortical gene expression, brain overgrowth and functional over-connectivity in chd8 haploinsufficient mice. Cereb Cortex 28(6):2192–2206
Gompers AL et al (2017) Germline Chd8 haploinsufficiency alters brain development in mouse. Nat Neurosci 20(8):1062–1073
Katayama Y et al (2016) CHD8 haploinsufficiency results in autistic-like phenotypes in mice. Nature 537(7622):675–679
Jiménez JA et al (2020) Chd8 haploinsufficiency impairs early brain development and protein homeostasis later in life. Mol Autism 11(1):74
Jung H et al (2018) Sexually dimorphic behavior, neuronal activity, and gene expression in Chd8-mutant mice. Nat Neurosci 21(9):1218–1228
Takebayashi S et al (2013) Murine esBAF chromatin remodeling complex subunits BAF250a and Brg1 are necessary to maintain and reprogram pluripotency-specific replication timing of select replication domains. Epigenet Chromatin 6(1):42
Staahl BT, Crabtree GR (2013) Creating a neural specific chromatin landscape by npBAF and nBAF complexes. Curr Opin Neurobiol 23(6):903–913
Bachmann C et al (2016) mSWI/SNF (BAF) complexes are indispensable for the neurogenesis and development of embryonic olfactory epithelium. PLoS Genet 12(9):e1006274
Lessard J et al (2007) An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron 55(2):201–215
Hempel A et al (2016) Deletions and de novo mutations of SOX11 are associated with a neurodevelopmental disorder with features of Coffin-Siris syndrome. J Med Genet 53(3):152–162
Lee MG et al (2007) Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 318(5849):447–450
Tuoc TC et al (2013) Chromatin regulation by BAF170 controls cerebral cortical size and thickness. Dev Cell 25(3):256–269
Narayanan R et al (2015) Loss of BAF (mSWI/SNF) complexes causes global transcriptional and chromatin state changes in forebrain development. Cell Rep 13(9):1842–1854
Nguyen H et al (2018) Epigenetic regulation by BAF complexes limits neural stem cell proliferation by suppressing Wnt signaling in late embryonic development. Stem Cell Rep 10(6):1734–1750
Holdhof D et al (2020) hGFAP-Positive stem cells depend on Brg1 for proper formation of cerebral and cerebellar structures. Cereb Cortex 30(3):1382–1392
Chenn A, Walsh CA (2002) Regulation of cerebral cortical size by control of cell cycle exit in neural precursors. Science 297(5580):365–369
Hirabayashi Y et al (2004) The Wnt/beta-catenin pathway directs neuronal differentiation of cortical neural precursor cells. Development 131(12):2791–2801
Caricasole A et al (2005) Two sides of the same coin: Wnt signaling in neurodegeneration and neuro-oncology. Biosci Rep 25(5–6):309–327
Lazzaro MA, Picketts DJ (2001) Cloning and characterization of the murine Imitation Switch (ISWI) genes: differential expression patterns suggest distinct developmental roles for Snf2h and Snf2l. J Neurochem 77(4):1145–1156
Alvarez-Saavedra M et al (2014) Snf2h-mediated chromatin organization and histone H1 dynamics govern cerebellar morphogenesis and neural maturation. Nat Commun 5(1):4181
Stopka T, Skoultchi AI (2003) The ISWI ATPase Snf2h is required for early mouse development. Proc Natl Acad Sci USA 100(24):14097–14102
Yip DJ et al (2012) Snf2l regulates Foxg1-dependent progenitor cell expansion in the developing brain. Dev Cell 22(4):871–878
Kumamoto T et al (2013) Foxg1 coordinates the switch from nonradially to radially migrating glutamatergic subtypes in the neocortex through spatiotemporal repression. Cell Reports 3(3):931–945
Martynoga B et al (2005) Foxg1 is required for specification of ventral telencephalon and region-specific regulation of dorsal telencephalic precursor proliferation and apoptosis. Dev Biol 283(1):113–127
Alvarez-Saavedra M et al (2019) Snf2h drives chromatin remodeling to prime upper layer cortical neuron development. Front Mol Neurosci 12:243
López AJ, Hecking JK, White AO (2020) The emerging role of ATP-dependent chromatin remodeling in memory and substance use disorders. Int J Mol Sci 21(18):6816
Bögershausen N, Wollnik B (2018) Mutational landscapes and phenotypic spectrum of SWI/SNF-related intellectual disability Disorders. Front Mol Neurosci 11:252
Santen GWE et al (2013) Coffin-siris syndrome and the BAF complex: genotype-phenotype study in 63 patients. Hum Mutat 34(11):1519–1528
Schrier SA et al (2012) The Coffin-Siris syndrome: a proposed diagnostic approach and assessment of 15 overlapping cases. Am J Med Genet A 158A(8):1865–1876
Nicolaides P, Baraitser M (1993) An unusual syndrome with mental retardation and sparse hair. Clin Dysmorphol 2(3):232–236
Filatova A et al (2019) Mutations in SMARCB1 and in other Coffin-Siris syndrome genes lead to various brain midline defects. Nat Commun 10(1):2966
Halgren C et al (2012a) Corpus callosum abnormalities, intellectual disability, speech impairment, and autism in patients with haploinsufficiency of ARID1B. Clin Genet 82(3):248–255
Kosho T, Okamoto N (2014) Genotype-phenotype correlation of Coffin-Siris syndrome caused by mutations in SMARCB1, SMARCA4, SMARCE1, and ARID1A. Am J Med Genet C Semin Med Genet 166c(3):262–275
Santen GWE et al (2012) Mutations in SWI/SNF chromatin remodeling complex gene ARID1B cause Coffin-Siris syndrome. Nat Genet 44(4):379–380
Sousa SB, Hennekam RC, t.N.B.S.I. Consortium (2014) Phenotype and genotype in Nicolaides-Baraitser syndrome. Am J Med Genet C 166(3):302–314
Celen C et al (2017) Arid1b haploinsufficient mice reveal neuropsychiatric phenotypes and reversible causes of growth impairment. eLife 6:1–22
Tsurusaki Y et al (2012) Mutations affecting components of the SWI/SNF complex cause Coffin-Siris syndrome. Nat Genet 44(4):376–378
Vasileiou G et al (2018) Mutations in the BAF-complex subunit DPF2 are associated with coffin-siris syndrome. Am J Hum Genet 102(3):468–479
Bramswig NC et al (2017) Heterozygosity for ARID2 loss-of-function mutations in individuals with a Coffin-Siris syndrome-like phenotype. Hum Genet 136(3):297–305
Campeau PM, Hennekam RC (2014) DOORS syndrome: phenotype, genotype and comparison with Coffin-Siris syndrome. Am J Med Genet C Semin Med Genet 166c(3):327–332
Kleefstra T et al (2012) Disruption of an EHMT1-Associated chromatin-modification module causes intellectual disability. Am J Hum Genet 91(1):73–82
Yoon K-J et al (2018) Epigenetics and epitranscriptomics in temporal patterning of cortical neural progenitor competence. J Cell Biol 217(6):1901–1914
Sokpor G et al (2018) ATP-dependent chromatin remodeling during cortical neurogenesis. Front Neurosci 12:226
Mizuguchi G et al (2004) ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303(5656):343–348
Ruhl DD et al (2006) Purification of a human SRCAP complex that remodels chromatin by incorporating the histone variant H2A.Z into nucleosomes. Biochemistry 45(17):5671–5677
Keil JM et al (2020) Symmetric neural progenitor divisions require chromatin-mediated homologous recombination DNA repair by Ino80. Nat Commun 11(1):3839
Alazami AM et al (2015) Accelerating novel candidate gene discovery in neurogenetic disorders via whole-exome sequencing of prescreened multiplex consanguineous families. Cell Rep 10(2):148–161
Papamichos-Chronakis M et al (2011) Global regulation of H2A.Z localization by the INO80 chromatin-remodeling enzyme is essential for genome integrity. Cell 144(2):200–213
Robinson PL et al (1988) A unique association of short stature, dysmorphic features, and speech impairment (Floating-Harbor syndrome). J Pediatr 113(4):703–706
Hood RL et al (2012) Mutations in SRCAP, encoding SNF2-related CREBBP activator protein, cause Floating-Harbor syndrome. Am J Hum Genet 90(2):308–313
Fryns JP et al (1996) The floating-harbor syndrome: two affected siblings in a family. Clin Genet 50(4):217–219
Patton MA et al (1991) Floating-harbor syndrome. J Med Genet 28(3):201–204
Xue Y et al (2003) The ATRX syndrome protein forms a chromatin-remodeling complex with Daxx and localizes in promyelocytic leukemia nuclear bodies. Proc Natl Acad Sci USA 100(19):10635–10640
Tang J et al (2004) A novel transcription regulatory complex containing death domain-associated protein and the ATR-X syndrome protein. J Biol Chem 279(19):20369–20377
Lewis PW et al (2010) Daxx is an H3.3-specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc Natl Acad Sci USA 107(32):14075–14080
Goldberg AD et al (2010) Distinct factors control histone variant H33 localization at specific genomic regions. Cell 140(5):678–691
Kernohan KD et al (2010) ATRX partners with cohesin and MeCP2 and contributes to developmental silencing of imprinted genes in the brain. Dev Cell 18(2):191–202
Garrick D et al (2006) Loss of Atrx affects trophoblast development and the pattern of X-inactivation in extraembryonic tissues. PLoS Genet 2(4):e58
Bérubé NG et al (2005) The chromatin-remodeling protein ATRX is critical for neuronal survival during corticogenesis. J Clin Investig 115(2):258–267
Seah C et al (2008) Neuronal death resulting from targeted disruption of the Snf2 protein ATRX Is mediated by p53. J Neurosci 28(47):12570–12580
Watson LA et al (2013) Atrx deficiency induces telomere dysfunction, endocrine defects, and reduced life span. J Clin Invest 123(5):2049–2063
Gibbons RJ et al (1995) Mutations in a putative global transcriptional regulator cause X-linked mental retardation with α-thalassemia (ATR-X syndrome). Cell 80(6):837–845
Fame RM, MacDonald JL, Macklis JD (2011) Development, specification, and diversity of callosal projection neurons. Trends Neurosci 34:41–50
Lodato S, Shetty AS, Arlotta P (2015) Cerebral cortex assembly: generating and reprogramming projection neuron diversity. Trends Neurosci 38:117–125
Margari L et al (2016) Clinical manifestations in children and adolescents with corpus callosum abnormalities. J Neurol 263:1939–1945
Halgren C et al (2012b) Corpus callosum abnormalities, intellectual disability, speech impairment, and autism in patients with haploinsufficiency of ARID1B. Clin Genet 82:248–255
Shu T et al (2003) Abnormal development of forebrain midline glia and commissural projections in Nfia knock-out mice. J Neurosci 23:203–212
Alcamo EA et al (2008) Satb2 regulates callosal projection neuron identity in the developing cerebral cortex. Neuron 57:364–377
Britanova O et al (2008) Satb2 Is a postmitotic determinant for upper-layer neuron specification in the neocortex. Neuron 57:378–392
Leone DP et al (2015) Satb2 regulates the differentiation of both callosal and subcerebral projection neurons in the developing cerebral cortex. Cereb Cortex 25:3406–3419
Cederquist GY et al (2013) Lmo4 establishes rostral motor cortex projection neuron subtype diversity. J Neurosci 33:6321–6332
Harb K et al (2016) Area-specific development of distinct projection neuron subclasses is regulated by postnatal epigenetic modifications. eLife 5:1–25
Schoof M et al (2019) The transcriptional coactivator and histone acetyltransferase CBP regulates neural precursor cell development and migration. Acta Neuropathol Commun 7:199
Oshiro H et al (2015) Up-regulation of HP1γ expression during neuronal maturation promotes axonal and dendritic development in mouse embryonic neocortex. Genes Cells 20:108–120
Ross SE et al (2012) Bhlhb5 and Prdm8 form a repressor complex involved in neuronal circuit assembly. Neuron 73:292–303
Xu Q et al (2018) Autism-associated CHD8 deficiency impairs axon development and migration of cortical neurons. Mol Autism 9:1–17
Mignot C et al (2016) ARID1B mutations are the major genetic cause of corpus callosum anomalies in patients with intellectual disability. Brain 139:e64
Peter CJ et al (2019) In vivo epigenetic editing of Sema6a promoter reverses transcallosal dysconnectivity caused by C11orf46/Arl14ep risk gene. Nat Commun 10:1–14
Najmabadi H et al (2011) Deep sequencing reveals 50 novel genes for recessive cognitive disorders. Nature 478:57–63
Meyer HS et al (2010) Number and laminar distribution of neurons in a thalamocortical projection column of rat vibrissal cortex. Cereb Cortex 20:2277–2286
Sahara S et al (2012) The fraction of cortical GABAergic neurons is constant from near the start of cortical neurogenesis to adulthood. J Neurosci 32:4755–4761
Selten M, van Bokhoven H, Nadif Kasri N (2018) Inhibitory control of the excitatory/inhibitory balance in psychiatric disorders. F1000Research 7:23
Lee S et al (2010) The largest group of superficial neocortical GABAergic interneurons expresses ionotropic serotonin receptors. J Neurosci 30:16796–16808
Rudy B et al (2011) Three groups of interneurons account for nearly 100% of neocortical GABAergic neurons. Dev Neurobiol 71:45–61
Hodge RD et al (2019) Conserved cell types with divergent features in human versus mouse cortex. Nature 573:61–68
Tasic B et al (2018) Shared and distinct transcriptomic cell types across neocortical areas. Nature 563:72–78
Gouwens NW et al (2020) Toward an integrated classification of neuronal cell types: morphoelectric and transcriptomic characterization of individual GABAergic cortical neurons. bioRxiv 2020:2020.02.03.932244
Gala R et al (2020) Consistent cross-modal identification of cortical neurons with coupled autoencoders. BioRXiv
Huang ZJ, Paul A (2019) The diversity of GABAergic neurons and neural communication elements. Nat Rev Neurosci 20:563–572
Hansen DV et al (2013) Non-epithelial stem cells and cortical interneuron production in the human ganglionic eminences. Nat Neurosci 16:1576–1587
Laclef C, Métin C (2018) Conserved rules in embryonic development of cortical interneurons. Semin Cell Dev Biol 76:86–100
Lim L et al (2018a) Development and functional diversification of cortical interneurons. Neuron 100(2):294–313
Marin O (2013) Cellular and molecular mechanisms controlling the migration of neocortical interneurons. Eur J Neurosci 38(1):2019–2029
Gelman D et al (2011) A wide diversity of cortical GABAergic interneurons derives from the embryonic preoptic area. J Neurosci 31:16570–16580
Kessaris N et al (2014) Genetic programs controlling cortical interneuron fate. Curr Opin Neurobiol 26:79–87
Tremblay R, Lee S, Rudy B (2016) GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91:260–292
Zimmer-Bensch G (2018) Diverse facets of cortical interneuron migration regulation—implications of neuronal activity and epigenetics. Brain Res 1700:160–169
Thomas T et al (2000) Querkopf, a MYST family histone acetyltransferase, is required for normal cerebral cortex development. Development 127:2537–2548
Bjornsson HT (2015b) The Mendelian disorders of the epigenetic machinery. Genome Res 25:1473–1481
Campeau PM et al (2012) The KAT6B-related disorders genitopatellar syndrome and Ohdo/SBBYS syndrome have distinct clinical features reflecting distinct molecular mechanisms. Hum Mutat 33:1520–1525
Tham E et al (2015) Dominant mutations in KAT6A cause intellectual disability with recognizable syndromic features. Am J Hum Genet 96(3):507–513
Arboleda VA et al (2015) De novo nonsense mutations in KAT6A, a lysine acetyl-transferase gene, cause a syndrome including microcephaly and global developmental delay. Am J Hum Genet 96(3):498–506
Kennedy J et al (2019) KAT6A Syndrome: genotype-phenotype correlation in 76 patients with pathogenic KAT6A variants. Genet Med 21(4):850–860
Voss AK et al (2009) Moz and retinoic acid coordinately regulate H3K9 acetylation, Hox gene expression, and segment identity. Dev Cell 17(5):674–686
Carvill GL et al (2013) Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat Genet 45:825–830
Meganathan K et al (2017) Regulatory networks specifying cortical interneurons from human embryonic stem cells reveal roles for CHD2 in interneuron development. Proc Natl Acad Sci USA 114:E11180–E11189
Jung EM et al (2017) Arid1b haploinsufficiency disrupts cortical interneuron development and mouse behavior. Nat Neurosci 20:1694–1707
Smith AL et al (2020) Arid1b haploinsufficiency in parvalbumin- or somatostatin-expressing interneurons leads to distinct ASD-like and ID-like behavior. Sci Rep 10(1):7834
Valor L et al (2013) Lysine acetyltransferases CBP and p300 as therapeutic targets in cognitive and neurodegenerative disorders. Curr Pharm Des 19:5051–5064
Tsui D et al (2014) CBP regulates the differentiation of interneurons from ventral forebrain neural precursors during murine development. Dev Biol 385:230–241
Medrano-Fernández A et al (2019) The epigenetic factor CBP is required for the differentiation and function of medial ganglionic eminence-derived interneurons. Mol Neurobiol 56:4440–4454
Merk DJ et al (2018) Opposing effects of CREBBP mutations govern the phenotype of rubinstein-taybi syndrome and adult SHH medulloblastoma. Dev Cell 44(6):709–772 (e6)
Al-Qattan MM et al (2019) Rubinstein-Taybi syndrome in a Saudi boy with distinct features and variants in both the CREBBP and EP300 genes: a case report. BMC Med Genet 20:10–15
Jovanovic JN, Thomson AM (2011) Development of cortical GABAergic innervation. Front Cell Neurosci 5:14
Sultan KT et al (2018) Progressive divisions of multipotent neural progenitors generate late-born chandelier cells in the neocortex. Nat Commun 9(1):4595
Lim L et al (2018b) Optimization of interneuron function by direct coupling of cell migration and axonal targeting. Nat Neurosci 21(7):920–931
Pensold D et al (2017) The DNA methyltransferase 1 (DNMT1) controls the shape and dynamics of migrating poa-derived interneurons fated for the murine cerebral cortex. Cereb Cortex 27:5696–5714
Symmank J et al (2018) DNMT1 modulates interneuron morphology by regulating Pak6 expression through crosstalk with histone modifications. Epigenetics 13:536–556
Takesian AE et al (2018) Inhibitory circuit gating of auditory critical-period plasticity. Nat Neurosci 21:1
Kim Y et al (2017) Brain-wide Maps Reveal Stereotyped Cell-Type-Based Cortical Architecture and Subcortical Sexual Dimorphism. Cell 171:456-469.e22
Butt SJ et al (2017) A role for GABAergic interneuron diversity in circuit development and plasticity of the neonatal cerebral cortex. Curr Opin Neurobiol 43:149–155
Reh RK et al (2020) Critical period regulation across multiple timescales. Proc Natl Acad Sci USA 117(38):23242–23251
Kinnischtzke AK et al (2012) Postnatal maturation of somatostatin-expressing inhibitory cells in the somatosensory cortex of GIN mice. Front Neural Circuits 6:33
LeBlanc JJ, Fagiolini M (2011) Autism: a “critical period” disorder? Neural Plast 2011:921680
Amir RE et al (1999) Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl- CpG-binding protein 2. Nat Genet 23:185–188
Krishnan K et al (2015) MeCP2 regulates the timing of critical period plasticity that shapes functional connectivity in primary visual cortex. Proc Natl Acad Sci 112:E4782–E4791
Krishnan K et al (2017) MECP2 regulates cortical plasticity underlying a learned behaviour in adult female mice. Nat Commun 8:14077
Patrizi A et al (2019) Accelerated hyper-maturation of parvalbumin circuits in the absence of MeCP2. Cereb Cortex 30(1):256–268
Picard N, Fagiolini M (2019) MeCP2: an epigenetic regulator of critical periods. Curr Opin Neurobiol 59:95–101
Mierau SB et al (2016) Cell-specific regulation of N-methyl-D-aspartate receptor maturation by Mecp2 in cortical circuits. Biol Psychiat 79:746–754
Chao HT et al (2010) Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468:263–269
Durand S et al (2012) NMDA receptor regulation prevents regression of visual cortical function in the absence of Mecp2. Neuron 76:1078–1090
Dehorter N et al (2015) Tuning of fast-spiking interneuron properties by an activity-dependent transcriptional switch. Science 349(6253):1216–1220
Lau BYB et al (2020) Maternal experience-dependent cortical plasticity in mice is circuit- and stimulus-specific and requires MECP2. J Neurosci 2020:1964–2019
Negwer M et al (2020) EHMT1 regulates Parvalbumin-positive interneuron development and GABAergic input in sensory cortical areas. Brain Struct Funct 225:2701–2716
Apulei J et al (2019) Non-cell autonomous OTX2 homeoprotein regulates visual cortex plasticity through Gadd45b/g. Cereb Cortex 29:2384–2395
Beurdeley M et al (2012) Otx2 binding to perineuronal nets persistently regulates plasticity in the mature visual cortex. J Neurosci 32:9429–9437
Lee HHC et al (2017) Genetic Otx2 mis-localization delays critical period plasticity across brain regions. Mol Psychiatry 22:680–688
Spatazza J et al (2013) Choroid-plexus-derived Otx2 homeoprotein constrains adult cortical plasticity. Cell Rep 3:1815–1823
Kessaris N et al (2006) Competing waves of oligodendrocytes in the forebrain and postnatal elimination of an embryonic lineage. Nat Neurosci 9(2):173–179
Marshall CA, Goldman JE (2002) Subpallial dlx2-expressing cells give rise to astrocytes and oligodendrocytes in the cerebral cortex and white matter. J Neurosci 22(22):9821–9830
Pinto L, Götz M (2007) Radial glial cell heterogeneity—the source of diverse progeny in the CNS. Prog Neurobiol 83(1):2–23
Levison SW, Goldman JE (1993) Both oligodendrocytes and astrocytes develop from progenitors in the subventricular zone of postnatal rat forebrain. Neuron 10(2):201–212
Hartline DK (2011) The evolutionary origins of glia. Glia 59(9):1215–1236
Sloan SA, Barres BA (2014) Mechanisms of astrocyte development and their contributions to neurodevelopmental disorders. Curr Opin Neurobiol 27:75–81
Freeman MR, Rowitch DH (2013) Evolving concepts of gliogenesis: a look way back and ahead to the next 25 years. Neuron 80(3):613–623
Zuchero JB, Barres BA (2015) Glia in mammalian development and disease. Development 142(22):3805–3809
Belanger M, Magistretti PJ (2009) The role of astroglia in neuroprotection. Dialogues Clin Neurosci 11(3):281–295
Maragakis NJ, Rothstein JD (2001) Glutamate transporters in neurologic disease. Arch Neurol 58(3):365–370
Baldwin KT, Eroglu C (2017) Molecular mechanisms of astrocyte-induced synaptogenesis. Curr Opin Neurobiol 45:113–120
Walz W (2000) Role of astrocytes in the clearance of excess extracellular potassium. Neurochem Int 36(4–5):291–300
Belanger M, Allaman I, Magistretti PJ (2011) Brain energy metabolism: focus on astrocyte-neuron metabolic cooperation. Cell Metab 14(6):724–738
Krencik R et al (2015) Dysregulation of astrocyte extracellular signaling in Costello syndrome. Sci Transl Med 7(286):286ra66
Hillen AEJ, Burbach JPH, Hol EM (2018) Cell adhesion and matricellular support by astrocytes of the tripartite synapse. Prog Neurobiol 165–167:66–86
Ferry L et al (2017) Methylation of DNA Ligase 1 by G9a/GLP Recruits UHRF1 to Replicating DNA and Regulates DNA Methylation. Mol Cell 67(4):550-565.e5
Fritsch L et al (2010) A subset of the histone H3 lysine 9 methyltransferases Suv39h1, G9a, GLP, and SETDB1 participate in a multimeric complex. Mol Cell 37(1):46–56
Sher F, Boddeke E, Copray S (2011) Ezh2 expression in astrocytes induces their dedifferentiation toward neural stem cells. Cell Reprogram 13(1):1–6
Hwang WW et al (2014) Distinct and separable roles for EZH2 in neurogenic astroglia. Elife 3:e02439
Tartaglia M et al (2001) Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat Genet 29(4):465–468
Hegedus B et al (2007) Neurofibromatosis-1 regulates neuronal and glial cell differentiation from neuroglial progenitors in vivo by both cAMP- and Ras-dependent mechanisms. Cell Stem Cell 1(4):443–457
Paquin A et al (2009) Costello syndrome H-Ras alleles regulate cortical development. Dev Biol 330(2):440–451
Urosevic J et al (2011) Constitutive activation of B-Raf in the mouse germ line provides a model for human cardio-facio-cutaneous syndrome. Proc Natl Acad Sci USA 108(12):5015–5020
Pacey LK, Doering LC (2007) Developmental expression of FMRP in the astrocyte lineage: implications for fragile X syndrome. Glia 55(15):1601–1609
Quinlan RA et al (2007) GFAP and its role in Alexander disease. Exp Cell Res 313(10):2077–2087
Uhlmann EJ et al (2002) Astrocyte-specific TSC1 conditional knockout mice exhibit abnormal neuronal organization and seizures. Ann Neurol 52(3):285–296
Ballas N et al (2009) Non-cell autonomous influence of MeCP2-deficient glia on neuronal dendritic morphology. Nat Neurosci 12(3):311–317
Maezawa I et al (2009) Rett syndrome astrocytes are abnormal and spread MeCP2 deficiency through gap junctions. J Neurosci 29(16):5051–5061
Okabe Y et al (2012) Alterations of gene expression and glutamate clearance in astrocytes derived from an MeCP2-null mouse model of Rett syndrome. PLoS ONE 7(4):e35354
Lioy DT et al (2011) A role for glia in the progression of Rett’s syndrome. Nature 475(7357):497–500
Fatemi SH et al (2008) Expression of astrocytic markers aquaporin 4 and connexin 43 is altered in brains of subjects with autism. Synapse 62(7):501–507
Nagy C et al (2016) Repression of astrocytic connexins in cortical and subcortical brain regions and prefrontal enrichment of H3K9me3 in depression and suicide. Int J Neuropsychopharmacol 20(1):50–57
Nagy C et al (2015) Astrocytic abnormalities and global DNA methylation patterns in depression and suicide. Mol Psychiatry 20(3):320–328
Matsumoto S et al (2006) Brg1 is required for murine neural stem cell maintenance and gliogenesis. Dev Biol 289(2):372–383
Kim SI et al (2009) BRG1 requirement for long-range interaction of a locus control region with a downstream promoter. Proc Natl Acad Sci USA 106(7):2259–2264
Ni Z, Bremner R (2007) Brahma-related gene 1-dependent STAT3 recruitment at IL-6-inducible genes. J Immunol 178(1):345–351
Ito K et al (2018) Gfap and Osmr regulation by BRG1 and STAT3 via interchromosomal gene clustering in astrocytes. Mol Biol Cell 29(2):209–219
Gao F et al (2019) Heterozygous mutations in SMARCA2 reprogram the enhancer landscape by global retargeting of SMARCA4. Mol Cell 75(5):891-904.e7
Telley L et al (2016) Sequential transcriptional waves direct the differentiation of newborn neurons in the mouse neocortex. Science 351(6280):1443
Benevento M et al (2016) Histone methylation by the Kleefstra syndrome protein EHMT1 mediates homeostatic synaptic scaling. Neuron 91(2):341–355
Tanenbaum ME et al (2014) A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159:635–646
Roux KJ et al (2012) A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J Cell Biol 196:801–810
Ummethum H, Hamperl S (2020) Proximity labeling techniques to study chromatin. Front Genet 11:1–13
Klingler E, Jabaudon D (2020) Do progenitors play dice? Elife 2020:9
Govindan S, Oberst P, Jabaudon D (2018) In vivo pulse labeling of isochronic cohorts of cells in the central nervous system using FlashTag. Nat Protoc 13:2297–2311
Ascoli GA et al (2008) Petilla terminology: nomenclature of features of GABAergic interneurons of the cerebral cortex. Nat Rev Neurosci 9(7):557–568
Tabata H (2015) Diverse subtypes of astrocytes and their development during corticogenesis. Front Neurosci 9:114
Tasic B et al (2016) Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 19(2):335–346
Bachoo RM et al (2004) Molecular diversity of astrocytes with implications for neurological disorders. Proc Natl Acad Sci USA 101(22):8384–8389
Doyle JP et al (2008) Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135(4):749–762
Gouwens NW et al (2019) Classification of electrophysiological and morphological neuron types in the mouse visual cortex. Nat Neurosci 22(7):1182–1195
Frega M et al (2020) Distinct pathogenic genes causing intellectual disability and autism exhibit a common neuronal network hyperactivity phenotype. Cell Rep 30(1):173–186
Poeta L et al (2019) Histone demethylase KDM5C is a SAHA-sensitive central hub at the crossroads of transcriptional axes involved in multiple neurodevelopmental disorders. Hum Mol Genet 28(24):4089–4102
Wade AA et al (2018) Common CHD8 genomic targets contrast with model-specific transcriptional impacts of CHD8 haploinsufficiency. Front Mol Neurosci 11:481
Siderius LE et al (1999) X-linked mental retardation associated with cleft lip/palate maps Xp11.3-q21.3. Am J Med Genet 85(3):216–220
Strømme P et al (2002) Mutations in the human ortholog of Aristaless cause X-linked mental retardation and epilepsy. Nat Genet 30(4):441–445
van der Werf IM et al (2017) Mutations in two large pedigrees highlight the role of ZNF711 in X-linked intellectual disability. Gene 605:92–98
Vallianatos CN et al (2020) Mutually suppressive roles of KMT2A and KDM5C in behaviour, neuronal structure, and histone H3K4 methylation. Commun Biol 3(1):278
Bjornsson HT et al (2014) Histone deacetylase inhibition rescues structural and functional brain deficits in a mouse model of Kabuki syndrome. Sci Transl Med 6(256):256ra135
Park J, Thomas S, Munster PN (2015) Epigenetic modulation with histone deacetylase inhibitors in combination with immunotherapy. Epigenomics 7(4):641–652
Alarcón JM et al (2004) Chromatin acetylation, memory, and LTP are impaired in CBP+/- mice: a model for the cognitive deficit in Rubinstein-Taybi syndrome and its amelioration. Neuron 42(6):947–959
Cenik B et al (2011) Suberoylanilide hydroxamic acid (vorinostat) up-regulates progranulin transcription: rational therapeutic approach to frontotemporal dementia. J Biol Chem 286(18):16101–16108
Penisson M et al (2019) Genes and mechanisms involved in the generation and amplification of basal radial glial cells. Front Cell Neurosci 13:1–21
Falcone C et al (2019) Cortical interlaminar astrocytes across the therian mammal radiation. J Comp Neurol 527(10):1654–1674
Boldog E et al (2018) Transcriptomic and morphophysiological evidence for a specialized human cortical GABAergic cell type. Nat Neurosci 21(9):1185–1195
Deneault E et al (2018) Complete disruption of autism-susceptibility genes by gene editing predominantly reduces functional connectivity of isogenic human neurons. Stem Cell Rep 11(5):1211–1225
Cederquist GY et al (2020) A multiplex human pluripotent stem cell platform defines molecular and functional subclasses of autism-related genes. Cell Stem Cell 27(1):35-49.e6
Trevino AE et al (2020) Chromatin accessibility dynamics in a model of human forebrain development. Science 367(6476):eaay1645
Mansour AAF, Schafer ST, Gage FH (2020) Cellular complexity in brain organoids: Current progress and unsolved issues. Semin Cell Dev Biol. https://doi.org/10.1016/j.semcdb.2020.05.013
Lancaster MA et al (2013) Cerebral organoids model human brain development and microcephaly. Nature 501(7467):373–379
Mariani J et al (2015) FOXG1-Dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 162(2):375–390
de Jong, J.O., et al., Cortical Overgrowth in a Preclinical Forebrain Organoid Model of CNTNAP2-Associated Autism Spectrum Disorder. Biorxiv, 2019: https://doi.org/https://doi.org/10.1101/739391
Urresti, J., et al., Cortical Organoids Model Early Brain Development Disrupted by 16p11.2 Copy Number Variants in Autism. bioRxiv, 2020: p. 2020.06.25.172262.
Zhang W et al (2020) Cerebral organoid and mouse models reveal a RAB39b-PI3K-mTOR pathway-dependent dysregulation of cortical development leading to macrocephaly/ autism phenotypes. Genes Dev 34:580–597
Pham MT et al (2018) Generation of human vascularized brain organoids. NeuroReport 29:588–593
Shi Y et al (2020) Vascularized human cortical organoids (vOrganoids) model cortical development in vivo. PLoS Biol 18:1–29
Mansour AA et al (2018) An in vivo model of functional and vascularized human brain organoids. Nat Biotechnol 36:432–441
Krasteva V et al (2012) The BAF53a subunit of SWI/SNF-like BAF complexes is essential for hemopoietic stem cell function. Blood 120(24):4720–4732
Bell S et al (2019) Mutations in ACTL6B Cause Neurodevelopmental Deficits and Epilepsy and Lead to Loss of Dendrites in Human Neurons. Am J Hum Genet 104(5):815–834
Gao X et al (2008) ES cell pluripotency and germ-layer formation require the SWI/SNF chromatin remodeling component BAF250a. Proc Natl Acad Sci U S A 105(18):6656–6661
Zhang, W., et al., The BAF and PRC2 Complex Subunits Dpf2 and Eed Antagonistically Converge on Tbx3 to Control ESC Differentiation. Cell Stem Cell, 2019. 24(1): p. 138–152 e8.
White AO et al (2016) BDNF rescues BAF53b-dependent synaptic plasticity and cocaine-associated memory in the nucleus accumbens. Nat Commun 7:11725
Chandler RL, Magnuson T (2016) The SWI/SNF BAF-A complex is essential for neural crest development. Dev Biol 411(1):15–24
Friocourt G et al (2008) Cell-Autonomous Roles of ARX in Cell Proliferation and Neuronal Migration during Corticogenesis. The Journal of Neuroscience 28(22):5794–5805
Laperuta C et al (2007) MRX87 family with Aristaless X dup24bp mutation and implication for polyAlanine expansions. BMC Med Genet 8:25–25
Poeta L et al (2013) A regulatory path associated with X-linked intellectual disability and epilepsy links KDM5C to the polyalanine expansions in ARX. Am J Hum Genet 92(1):114–125
Shoubridge C, Fullston T, Gécz J (2010) ARX spectrum disorders: making inroads into the molecular pathology. Hum Mutat 31(8):889–900
Medina CF et al (2008) Altered visual function and interneuron survival in Atrx knockout mice: inference for the human syndrome. Hum Mol Genet 18(5):966–977
Tamming RJ et al (2020) Atrx deletion in neurons leads to sexually dimorphic dysregulation of miR-137 and spatial learning and memory deficits. Cell Rep 31(13):107838
Tamming RJ et al (2017) Mosaic expression of Atrx in the mouse central nervous system causes memory deficits. Dis Models Mech 10(2):119–126
Smith RD et al (1980) Short stature, psychomotor retardation, and unusual facial appearance in two brothers. Am J Med Genet 7(1):5–9
Hori K, Hoshino M (2017) Neuronal migration and AUTS2 syndrome. Brain Sci 7(5):54
Lizarraga SB et al (2010) Cdk5rap2 regulates centrosome function and chromosome segregation in neuronal progenitors. Development 137(11):1907–1917
Lathrop MJ et al (2010) Deletion of the Chd6 exon 12 affects motor coordination. Mamm Genome 21(3–4):130–142
Alendar A et al (2020) Gene expression regulation by the Chromodomain helicase DNA-binding protein 9 (CHD9) chromatin remodeler is dispensable for murine development. PLoS ONE 15(5):e0233394
Subburaju S et al (2016) Toward dissecting the etiology of schizophrenia: HDAC1 and DAXX regulate GAD67 expression in an in vitro hippocampal GABA neuron model. Transl Psychiatry 6(1):e723–e723
Michod D et al (2012) Calcium-dependent dephosphorylation of the histone chaperone DAXX regulates H3.3 loading and transcription upon neuronal activation. Neuron 74(1):122–35
Takebayashi S et al (2007) Major and essential role for the DNA methylation mark in mouse embryogenesis and stable association of DNMT1 with newly replicated regions. Mol Cell Biol 27(23):8243–8258
Christian DL et al (2020) DNMT3A haploinsufficiency results in behavioral deficits and global epigenomic dysregulation shared across neurodevelopment disorders. bioRxiv 33(8):108416
Tarusawa E et al (2016) Establishment of high reciprocal connectivity between clonal cortical neurons is regulated by the Dnmt3b DNA methyltransferase and clustered protocadherins. BMC Biol 14(1):103
Toyoda S et al (2014) Developmental epigenetic modification regulates stochastic expression of clustered protocadherin genes, generating single neuron diversity. Neuron 82(1):94–108
Cohen ASA et al (2015) A novel mutation in EED associated with overgrowth. J Hum Genet 60(6):339–342
Liu P-P et al (2019) Polycomb protein EED regulates neuronal differentiation through targeting SOX11 in hippocampal dentate gyrus. Stem Cell Rep 13(1):115–131
Balemans MC et al (2010) Reduced exploration, increased anxiety, and altered social behavior: autistic-like features of euchromatin histone methyltransferase 1 heterozygous knockout mice. Behav Brain Res 208(1):47–55
Balemans MC et al (2013) Hippocampal dysfunction in the Euchromatin histone methyltransferase 1 heterozygous knockout mouse model for Kleefstra syndrome. Hum Mol Genet 22(5):852–866
Benevento M et al (2017) Haploinsufficiency of EHMT1 improves pattern separation and increases hippocampal cell proliferation. Sci Rep 7:40284
Frega M et al (2019) Neuronal network dysfunction in a model for Kleefstra syndrome mediated by enhanced NMDAR signaling. Nat Commun 10(1):4928
Iacono G et al (2018) Increased H3K9 methylation and impaired expression of Protocadherins are associated with the cognitive dysfunctions of the Kleefstra syndrome. Nucleic Acids Res 46(10):4950–4965
Kleefstra T et al (2006) Loss-of-function mutations in euchromatin histone methyl transferase 1 (EHMT1) cause the 9q34 subtelomeric deletion syndrome. Am J Hum Genet 79(2):370–377
Koemans TS et al (2017) Functional convergence of histone methyltransferases EHMT1 and KMT2C involved in intellectual disability and autism spectrum disorder. PLoS Genet 13(10):e1006864
Marchese G et al (2016) Kleefstra-variant syndrome with heterozygous mutations in EHMT1 and KCNQ2 genes: a case report. Neurol Sci 37:829–831
Martens MB et al (2016) Euchromatin histone methyltransferase 1 regulates cortical neuronal network development. Sci Rep 6:35756
Chopra A et al (2020) Hypoxia-inducible lysine methyltransferases: G9a and GLP hypoxic regulation, non-histone substrate modification, and pathological relevance. Front Genet 11:579636
Fiszbein A et al (2016) Alternative splicing of g9a regulates neuronal differentiation. Cell Rep 14(12):2797–2808
Maze I et al (2010) Essential role of the histone methyltransferase G9a in cocaine-induced plasticity. Science 327(5962):213–216
Henriquez B et al (2013) Ezh1 and Ezh2 differentially regulate PSD-95 gene transcription in developing hippocampal neurons. Mol Cell Neurosci 57:130–143
Imagawa E et al (2017) Mutations in genes encoding polycomb repressive complex 2 subunits cause Weaver syndrome. Hum Mutat 38(6):637–648
Lan F et al (2007) A histone H3 lysine 27 demethylase regulates animal posterior development. Nature 449(7163):689–694
Morris MJ et al (2013) Loss of histone deacetylase 2 improves working memory and accelerates extinction learning. J Neurosci 33(15):6401–6411
Haberland M et al (2009) Epigenetic control of skull morphogenesis by histone deacetylase 8. Genes Dev 23(14):1625–1630
Min J-N et al (2013) The mINO80 chromatin remodeling complex is required for efficient telomere replication and maintenance of genome stability. Cell Res 23(12):1396–1413
Herz HM et al (2010) The H3K27me3 demethylase dUTX is a suppressor of Notch- and Rb-dependent tumors in Drosophila. Mol Cell Biol 30(10):2485–2497
Shangguan H et al (2019) Kabuki syndrome: novel pathogenic variants, new phenotypes and review of literature. Orphanet J Rare Dis 14(1):255
Tunovic S et al (2014) De novo ANKRD11 and KDM1A gene mutations in a male with features of KBG syndrome and Kabuki syndrome. Am J Med Genet A 164a(7):1744–9
Sun G et al (2010) Histone demethylase LSD1 regulates neural Stem cell proliferation. Mol Cell Biol 30(8):1997–2005
Jakovcevski M et al (2015) Neuronal Kmt2a/Mll1 histone methyltransferase is essential for prefrontal synaptic plasticity and working memory. J Neurosci 35(13):5097–5108
Jones WD et al (2012) De Novo Mutations in MLL Cause Wiedemann-Steiner Syndrome. Am J Hum Genet 91(2):358–364
Camarena V et al (2014) Disruption of Mbd5 in mice causes neuronal functional deficits and neurobehavioral abnormalities consistent with 2q23.1 microdeletion syndrome. EMBO Mol Med 6(8):1003–15
Antoine MW et al (2019) Increased excitation-inhibition ratio stabilizes synapse and circuit excitability in four autism mouse models. Neuron 101:648-661.e4
Banerjee A et al (2016) Jointly reduced inhibition and excitation underlies circuit-wide changes in cortical processing in Rett syndrome. Proc Natl Acad Sci USA 113:E7287–E7296
Nelson SB, Valakh V (2015) Excitatory/inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron 87:684–698
Kawauchi S et al (2009) Multiple organ system defects and transcriptional dysregulation in the Nipbl+/− Mouse, a model of cornelia de lange syndrome. PLoS Genet 5(9):e1000650
Boussadia B et al (2016) Lack of CAR impacts neuronal function and cerebrovascular integrity in vivo. Exp Neurol 283(Pt A):39–48
Krasteva V, Crabtree GR, Lessard JA (2017) The BAF45a/PHF10 subunit of SWI/SNF-like chromatin remodeling complexes is essential for hematopoietic stem cell maintenance. Exp Hematol 48:58-71.e15
Franzoni E et al (2015) miR-128 regulates neuronal migration, outgrowth and intrinsic excitability via the intellectual disability gene Phf6. eLife 4:e04263
Zhang C et al (2013) The X-linked intellectual disability protein PHF6 associates with the PAF1 complex and regulates neuronal migration in the mammalian brain. Neuron 78(6):986–993
Zweier C et al (2014) Females with de novo aberrations in PHF6: Clinical overlap of Borjeson–Forssman–Lehmann with Coffin-Siris syndrome. Am J Med Genet C 166(3):290–301
Chen X et al (2018) Phf8 histone demethylase deficiency causes cognitive impairments through the mTOR pathway. Nat Commun 9(1):114
Riveiro AR et al (2017) JMJD-12/PHF8 controls axon guidance by regulating Hedgehog-like signaling. Development 144(5):856–865
Turnbull J et al (2012) Early-onset Lafora body disease. Brain 135(9):2684–2698
Pierce SB et al (2018) De novo mutation in RING1 with epigenetic effects on neurodevelopment. Proc Natl Acad Sci USA 115(7):1558–1563
Du TT et al (2014) Setdb2 controls convergence and extension movements during zebrafish gastrulation by transcriptional regulation of dvr1. Dev Biol 392(2):233–244
Falandry C et al (2010) CLLD8/KMT1F is a lysine methyltransferase that is important for chromosome segregation. J Biol Chem 285(26):20234–20241
Xu PF et al (2010) Setdb2 restricts dorsal organizer territory and regulates left-right asymmetry through suppressing fgf8 activity. Proc Natl Acad Sci USA 107(6):2521–2526
Kuechler A et al (2015) Loss-of-function variants of SETD5 cause intellectual disability and the core phenotype of microdeletion 3p25.3 syndrome. Eur J Hum Genet 23(6):753–60
Moore SM et al (2019) Setd5 haploinsufficiency alters neuronal network connectivity and leads to autistic-like behaviors in mice. Transl Psychiatry 9(1):24
Koga M et al (2009) Involvement of SMARCA2/BRM in the SWI/SNF chromatin-remodeling complex in schizophrenia. Hum Mol Genet 18(13):2483–2494
Battaglioli E et al (2002) REST repression of neuronal genes requires components of the hSWISNF complex. J Biol Chem 277(43):41038–45
Machol K et al (2019) Expanding the spectrum of BAF-related disorders: de novo variants in smarcc2 cause a syndrome with intellectual disability and developmental delay. Am J Hum Genet 104(1):164–178
Matsumoto S et al (2016) Brg1 directly regulates Olig2 transcription and is required for oligodendrocyte progenitor cell specification. Dev Biol 413(2):173–187
Diets IJ et al (2019) A recurrent de novo missense pathogenic variant in SMARCB1 causes severe intellectual disability and choroid plexus hyperplasia with resultant hydrocephalus. Genet Med 21(3):572–579
Al Mutairi F et al (2018) A mendelian form of neural tube defect caused by a de novo null variant in SMARCC1 in an identical twin. Ann Neurol 83(2):433–436
Tuoc T et al (2017) Ablation of BAF170 in developing and postnatal dentate gyrus affects neural stem cell proliferation, differentiation, and learning. Mol Neurobiol 54(6):4618–4635
Harmacek L et al (2014) A unique missense allele of BAF155, a core BAF chromatin remodeling complex protein, causes neural tube closure defects in mice. Dev Neurobiol 74(5):483–497
Nixon KCJ et al (2019) A syndromic neurodevelopmental disorder caused by mutations in SMARCD1, a Core SWI/SNF subunit needed for context-dependent neuronal gene regulation in flies. Am J Hum Genet 104(4):596–610
Fujita Y et al (2017) Decreased cohesin in the brain leads to defective synapse development and anxiety-related behavior. J Exp Med 214(5):1431–1452
Wang Y et al (2013) Transcription factor Sox11 is essential for both embryonic and adult neurogenesis. Dev Dyn 242(6):638–653
Miró X et al (2009) Haploinsufficiency of the murine polycomb gene Suz12 results in diverse malformations of the brain and neural tube. Dis Models Mech 2(7–8):412–418
Donohoe ME et al (1999) Targeted disruption of mouse Yin Yang 1 transcription factor results in peri-implantation lethality. Mol Cell Biol 19(10):7237–7244
Gabriele M et al (2017) YY1 Haploinsufficiency causes an intellectual disability syndrome featuring transcriptional and chromatin Dysfunction. Am J Hum Genet 100(6):907–925
Wu T, Donohoe ME (2019) Yy1 regulates Senp1 contributing to AMPA receptor GluR1 expression following neuronal depolarization. J Biomed Sci 26(1):79
Weymann S et al (2001) Severe arterial occlusive disorder and brachysyndactyly in a boy: a further case of Grange syndrome? Am J Med Genet 99(3):190–195
Shi Y et al (2011) Exome sequencing identifies ZNF644 mutations in high myopia. PLoS Genet 7(6):e1002084
Tarpey PS et al (2009) A systematic, large-scale resequencing screen of X-chromosome coding exons in mental retardation. Nat Genet 41(5):535–543
Kleine-Kohlbrecher D et al (2010) A functional link between the histone demethylase PHF8 and the transcription factor ZNF711 in X-linked mental retardation. Mol Cell 38(2):165–178
Acknowledgements
This work was supported by grants from: The Netherlands Organization for Scientific Research, NWO-CAS grant 012.200.001 (to N.N.K); the Netherlands Organization for Health Research and Development ZonMw grant 91217055 (to N.N.K); SFARI grant 610264 (to N.N.K); ERA-NET NEURON-102 SYNSCHIZ grant (NWO) 013-17-003 4538 (to D.S) and ERA-NET NEURON DECODE! grant (NWO) 013.18.001 (to N.N.K).
Author information
Authors and Affiliations
Contributions
BM, MN, and NNK conceived and wrote the manuscript. DS provided resources.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mossink, B., Negwer, M., Schubert, D. et al. The emerging role of chromatin remodelers in neurodevelopmental disorders: a developmental perspective. Cell. Mol. Life Sci. 78, 2517–2563 (2021). https://doi.org/10.1007/s00018-020-03714-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03714-5