Abstract
The acquisition of cell identity is associated with developmentally regulated changes in the cellular histone methylation signatures. For instance, commitment to neural differentiation relies on the tightly controlled gain or loss of H3K27me3, a hallmark of polycomb-mediated transcriptional gene silencing, at specific gene sets. The KDM6B demethylase, which removes H3K27me3 marks at defined promoters and enhancers, is a key factor in neurogenesis. Therefore, to better understand the epigenetic regulation of neural fate acquisition, it is important to determine how Kdm6b expression is regulated. Here, we investigated the molecular mechanisms involved in the induction of Kdm6b expression upon neural commitment of mouse embryonic stem cells. We found that the increase in Kdm6b expression is linked to a rearrangement between two 3D configurations defined by the promoter contact with two different regions in the Kdm6b locus. This is associated with changes in 5-hydroxymethylcytosine (5hmC) levels at these two regions, and requires a functional ten-eleven-translocation (TET) 3 protein. Altogether, our data support a model whereby Kdm6b induction upon neural commitment relies on an intronic enhancer the activity of which is defined by its TET3-mediated 5-hmC level. This original observation reveals an unexpected interplay between the 5-hmC and H3K27me3 pathways during neural lineage commitment in mammals. It also questions to which extent KDM6B-mediated changes in H3K27me3 level account for the TET-mediated effects on gene expression.





Similar content being viewed by others
References
Conway E, Healy E, Bracken AP (2015) PRC2 mediated H3K27 methylations in cellular identity and cancer. Curr Opin Cell Biol 37:42–48. https://doi.org/10.1016/j.ceb.2015.10.003
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, Fry B, Meissner A, Wernig M, Plath K, Jaenisch R, Wagschal A, Feil R, Schreiber SL, Lander ES (2006) A bivalent chromatin structure marks key developmental genes inembryonic stem cells. Cell 125:315–326
Lien WH, Guo X, Polak L, Lawton LN, Young RA, Zheng D, Fuchs E (2011) Genome-wide maps of histone modifications unwind in vivo chromatin states of the hair follicle lineage. Cell Stem Cell 9:219–232. https://doi.org/10.1016/j.stem.2011.07.015
Voigt P, Tee WW, Reinberg D (2013) A double take on bivalent promoters. Genes Dev 27:1318–1338. https://doi.org/10.1101/gad.219626.113
Azuara V, Perry P, Sauer S, Spivakov M, Jørgensen HF, John RM, Gouti M, Casanova M, Warnes G, Merkenschlager M, Fisher AG (2006) Chromatin signatures of pluripotent cell lines. Nat Cell Biol 8:532–538
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K (2007) High-resolution profiling of histone methylations in the human genome. Cell 129:823–837
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP, Lee W, Mendenhall E, O'Donovan A, Presser A, Russ C, Xie X, Meissner A, Wernig M, Jaenisch R, Nusbaum C, Lander ES, Bernstein BE (2007) Genome wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448:553–560
Pan G, Tian S, Nie J, Yang C, Ruotti V, Wei H, Jonsdottir GA, Stewart R, Thomson JA (2007) Whole-genome analysis of histone H3 lysine 4 and lysine 27 methylation in human embryonic stem cells. Cell Stem Cell 1:299–312
Mohn F, Weber M, Rebhan M, Roloff TC, Richter J, Stadler MB, Bibel M, Schübeler D (2008) Lineage-specific polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol Cell 30:755–766. https://doi.org/10.1016/j.molcel.2008.05.007
Sanz LA, Chamberlain S, Sabourin JC, Henckel A, Magnuson T, Hugnot JP, Feil R, Arnaud P (2008) A mono-allelic bivalent chromatin domain controls tissue-specific imprinting at Grb10. EMBO J 27:2523–2532. https://doi.org/10.1038/emboj.2008.142
Cui K, Zang C, Roh TY, Schones DE, Childs RW, Peng W, Zhao K (2009) Chromatin signatures in multipotent human hematopoietic stem cells indicate the fate of bivalent genes during differentiation. Cell Stem Cell 4:80–93. https://doi.org/10.1016/j.stem.2008.11.011
Maupetit-Méhouas S, Montibus B, Nury D, Tayama C, Wassef M, Kota SK, Fogli A, Cerqueira Campos F, Hata K, Feil R, Margueron R, Nakabayashi K, Court F, Arnaud P (2016) Imprinting control regions (ICRs) are marked by mono-allelic bivalent chromatin when transcriptionally inactive. Nucleic Acids Res 44:621–635. https://doi.org/10.1093/nar/gkv960
Albert M, Kalebic N, Florio M, Lakshmanaperumal N, Haffner C, Brandl H, Henry I, Huttner WB (2017) Epigenome profiling and editing of neocortical progenitor cells during development. EMBO J 36:2642–2658. https://doi.org/10.15252/embj.201796764
Hirabayashi Y, Suzki N, Tsuboi M, Endo TA, Toyoda T, Shinga J, Koseki H, Vidal M, Gotoh Y (2009) Polycomb limits the neurogenic competence of neural precursor cells to promote astrogenic fate transition. Neuron 63:600–613. https://doi.org/10.1016/j.neuron.2009.08.021
Pereira JD, Sansom SN, Smith J, Dobenecker MW, Tarakhovsky A, Livesey FJ (2010) Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc Natl Acad Sci USA 107:15957–15962. https://doi.org/10.1073/pnas.1002530107
Morimoto-Suzki N, Hirabayashi Y, Tyssowski K, Shinga J, Vidal M, Koseki H, Gotoh Y (2014) The polycomb component Ring1B regulates the timed termination of subcerebral projection neuron production during mouse neocortical development. Development 141(4343):53. https://doi.org/10.1242/dev.112276
Yao M, Zhou X, Zhou J, Gong S, Hu G, Li J, Huang K, Lai P, Shi G, Hutchins AP, Sun H, Wang H, Yao H (2018) PCGF5 is required for neural differentiation of embryonic stem cells. Nat Commun 9:1463. https://doi.org/10.1038/s41467-018-03781-0
Agger K, Cloos PA, Christensen J, Pasini D, Rose S, Rappsilber J, Issaeva I, Canaani E, Salcini AE, Helin K (2007) UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449:731–734
De Santa F, Totaro MG, Prosperini E, Notarbartolo S, Testa G, Natoli G (2007) The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell 130:1083–1094
Jepsen K, Solum D, Zhou T, McEvilly RJ, Kim HJ, Glass CK, Hermanson O, Rosenfeld MG (2007) SMRT-mediated repression of an H3K27 demethylase in progression from neural stem cell to neuron. Nature 450:415–419
Burgold T, Spreafico F, De Santa F, Totaro MG, Prosperini E, Natoli G, Testa G (2008) The histone H3 lysine 27-specific demethylase Jmjd3 is required for neural commitment. PLoS ONE 3:e3034. https://doi.org/10.1371/journal.pone.0003034
Burgold T, Voituron N, Caganova M, Tripathi PP, Menuet C, Tusi BK, Spreafico F, Bévengut M, Gestreau C, Buontempo S, Simeone A, Kruidenier L, Natoli G, Casola S, Hilaire G, Testa G (2012) The H3K27 demethylase JMJD3 is required for maintenance of the embryonic respiratory neuronal network, neonatal breathing, and survival. Cell Rep 2:1244–1258. https://doi.org/10.1016/j.celrep.2012.09.013
Park DH, Hong SJ, Salinas RD, Liu SJ, Sun SW, Sgualdino J, Testa G, Matzuk MM, Iwamori N, Lim DA (2014) Activation of neuronal gene expression by the JMJD3 demethylase is required for postnatal and adult brain neurogenesis. Cell Rep 8:1290–1299. https://doi.org/10.1016/j.stem.2007.08.003
Gaspard N, Bouschet T, Herpoel A, Naeije G, van den Ameele J, Vanderhaeghen P (2009) Generation of cortical neurons from mouse embryonic stem cells. Nat Protoc 4:1454–1463. https://doi.org/10.1038/nprot.2009.157
Neely MD, Litt MJ, Tidball AM, Li GG, Aboud AA, Hopkins CR, Chamberlin R, Hong CC, Ess KC, Bowman AB (2012) DMH1, a highly selective small molecule BMP inhibitor promotes neurogenesis of hiPSCs: comparison of PAX6 and SOX1 expression during neural induction. ACS Chem Neurosci 3:482–491. https://doi.org/10.1021/cn300029t
Jin SG, Zhang ZM, Dunwell TL, Harter MR, Wu X, Johnson J, Li Z, Liu J, Szabó PE, Lu Q, Xu GL, Song J, Pfeifer GP (2016) Tet3 reads 5-carboxylcytosine through its CXXC domain and is a potential guardian against neurodegeneration. Cell Rep 14:493–505. https://doi.org/10.1016/j.celrep.2015.12.044
Arnaud P, Hata K, Kaneda M, Li E, Sasaki H, Feil R, Kelsey G (2006) Stochastic imprinting in the progeny of Dnmt3L−/− females. Hum Mol Genet 15:589–598
Santiago M, Antunes C, Guedes M, Iacovino M, Kyba M, Reik W, Sousa N, Pinto L, Branco MR, Marques CJ (2019) Tet3 regulates cellular identity and DNA methylation in neural progenitor cells. Cell Mol Life Sci. https://doi.org/10.1007/s00018-019-03335-7
Iacovino M, Bosnakovski D, Fey H, Rux D, Bajwa G, Mahen E, Mitanoska A, Xu Z, Kyba M (2011) Inducible cassette exchange: a rapid and efficient system enabling conditional gene expression in embryonic stem and primary cells. Stem Cells 29:1580–1588
Bibel M, Richter J, Lacroix E, Barde YA (2007) Generation of a defined and uniform population of CNS progenitors and neurons from mouse embryonic stem cells. Nat Protoc 2:1034–1043
Court F, Baniol M, Hagege H, Petit JS, Lelay-Taha MN, Carbonell F, Weber M, Cathala G, Forne T (2011) Long-range chromatin interactions at the mouse Igf2/H19 locus reveal a novel paternally expressed long non-coding RNA. Nucleic Acids Res 39:5893–5906. https://doi.org/10.1093/nar/gkr209
Wagschal A, Delaval K, Pannetier M, Arnaud P, Feil R (2007) Chromatin immunoprecipitation (ChIP) on unfixed chromatin from cells and tissues to analyze histone modifications. Cold Spring Harbor Protocols. https://doi.org/10.1101/pdb.prot4767(pdb.prot4767-prot4767)
Bouschet T, Dubois E, Reynès C, Kota SK, Rialle S, Maupetit-Méhouas S, Pezet M, Le Digarcher A, Nidelet S, Demolombe V, Cavelier P, Meusnier C, Maurizy C, Sabatier R, Feil R, Arnaud P, Journot L, Varrault A (2017) In vitro corticogenesis from embryonic stem cells recapitulates the in vivo epigenetic control of imprinted gene expression. Cereb Cortex 27:2418–2433. https://doi.org/10.1093/cercor/bhw102
Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, Hu M, Liu JS, Ren B (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485:376–380. https://doi.org/10.1038/nature11082
Bonev B, Mendelson Cohen N, Szabo Q, Fritsch L, Papadopoulos GL, Lubling Y, Xu X, Lv X, Hugnot JP, Tanay A, Cavalli G (2017) Multiscale 3D genome rewiring during mouse neural development. Cell 171:557–572. https://doi.org/10.1016/j.cell.2017.09.043
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, Agarwal S, Iyer LM, Liu DR, Aravind L, Rao A (2009) Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324:930–935. https://doi.org/10.1126/science.1170116
Kriaucionis S, Heintz N (2009) The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324:929–930. https://doi.org/10.1126/science.1169786
Sérandour AA, Avner S, Oger F, Bizot M, Percevault F, Lucchetti-Miganeh C, Palierne G, Gheeraert C, Barloy-Hubler F, Péron CL, Madigou T, Durand E, Froguel P, Staels B, Lefebvre P, Métivier R, Eeckhoute J, Salbert G (2012) Dynamic hydroxymethylation of deoxyribonucleic acid marks differentiation-associated enhancers. Nucleic Acids Res 40:8255–8265
Hahn MA, Qiu R, Wu X, Li AX, Zhang H, Wang J, Jui J, Jin SG, Jiang Y, Pfeifer GP, Lu Q (2013) Dynamics of 5-hydroxymethylcytosine and chromatin marks in mammalian neurogenesis. Cell Rep 3:291–300. https://doi.org/10.1016/j.celrep.2013.01.011
Booth MJ, Branco MR, Ficz G, Oxley D, Krueger F, Reik W, Balasubramanian S (2012) Quantitative sequencing of 5-methylcytosine and 5-hydroxymethylcytosine at single-base resolution. Science 336:934–937. https://doi.org/10.1126/science.1220671
Wu H, Zhang Y (2011) Tet1 and 5-hydroxymethylation: a genome-wide view in mouse embryonic stem cells. Cell Cycle 10:2428–2436
Greco CM, Kunderfranco P, Rubino M, Larcher V, Carullo P, Anselmo A, Kurz K, Carell T, Angius A, Latronico MV, Papait R, Condorelli G (2016) DNA hydroxymethylation controls cardiomyocyte gene expression in development and hypertrophy. Nat Commun 7:12418. https://doi.org/10.1038/ncomms12418
Tsagaratou A, Äijö T, Lio CW, Yue X, Huang Y, Jacobsen SE, Lähdesmäki H, Rao A (2014) Dissecting the dynamic changes of 5-hydroxymethylcytosine in T-cell development and differentiation. Proc Natl Acad Sci USA 111:E3306–E3315. https://doi.org/10.1073/pnas.1412327111
Mahé EA, Madigou T, Sérandour AA, Bizot M, Avner S, Chalmel F, Palierne G, Métivier R, Salbert G (2017) Cytosine modifications modulate the chromatin architecture of transcriptional enhancers. Genome Res 27:947–958. https://doi.org/10.1101/gr.211466.116
Ren G, Jin W, Cui K, Rodrigez J, Hu G, Zhang Z, Larson DR, Zhao K (2017) CTCF Mediated enhancer–promoter interaction is a critical regulator of cell-to-cell variation of gene expression. Mol Cell 67:1049–1058. https://doi.org/10.1016/j.molcel.2017.08.026
Li T, Yang D, Li J, Tang Y, Yang J, Le W (2015) Critical role of Tet3 in neural progenitor cell maintenance and terminal differentiation. Mol Neurobiol 51:142–154. https://doi.org/10.1007/s12035014-8734-5
Koh KP, Yabuuchi A, Rao S, Huang Y, Cunniff K, Nardone J, Laiho A, Tahiliani M, Sommer CA, Mostoslavsky G, Lahesmaa R, Orkin SH, Rodig SJ, Daley GQ, Rao A (2011) Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8:200–213. https://doi.org/10.1016/j.stem.2011.01.008
Dawlaty MM, Ganz K, Powell BE, Hu YC, Markoulaki S, Cheng AW, Gao Q, Kim J, Choi SW, Page DC, Jaenisch R (2011) Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9:166–175. https://doi.org/10.1016/j.stem.2011.07.010
Williams K, Christensen J, Pedersen MT, Johansen JV, Cloos PA, Rappsilber J, Helin K (2011) TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473:343–348. https://doi.org/10.1038/nature10066
Dawlaty MM, Breiling A, Le T, Raddatz G, Barrasa MI, Cheng AW, Gao Q, Powell BE, Li Z, Xu M, Faull KF, Lyko F, Jaenisch R (2013) Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev Cell 24:310–323. https://doi.org/10.1016/j.devcel.2012.12.015
Reimer M Jr, Pulakanti K, Shi L, Abel A, Liang M, Malarkannan S, Rao S (2019) Deletion of Tet proteins results in quantitative disparities during ESC differentiation partially attributable to alterations in gene expression. BMC Dev Biol 19:16. https://doi.org/10.1186/s12861-019-0196-6
Dawlaty MM, Breiling A, Le T, Barrasa MI, Raddatz G, Gao Q, Powell BE, Cheng AW, Faull KF, Lyko F, Jaenisch R (2014) Loss of Tet enzymes compromises proper differentiation of embryonic stem cells. Dev Cell 29:102–111. https://doi.org/10.1016/j.devcel.2014.03.003
Wu H, D'Alessio AC, Ito S, Xia K, Wang Z, Cui K, Zhao K, Sun YE, Zhang Y (2011) Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473:389–393. https://doi.org/10.1038/nature09934
Wu H, D'Alessio AC, Ito S, Wang Z, Cui K, Zhao K, Sun YE, Zhang Y (2011) Genome wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev 25:679–684. https://doi.org/10.1101/gad.2036011
Karuppagounder SS, Kumar A, Shao DS, Zille M, Bourassa MW, Caulfield JT, Alim I, Ratan RR (2015) Metabolism and epigenetics in the nervous system: creating cellular fitness and resistance to neuronal death in neurological conditions via modulation of oxygen-, iron-, and 2-oxoglutarate-dependent dioxygenases. Brain Res 1628(Pt B):273–287. https://doi.org/10.1016/j.brainres.2015.07.030
Acknowledgements
We thank Claire Chazaud for advice in ES cell derivation and all members of P.A.’s team for critical reading of the manuscript.
Funding
This research has been financed by the French government IDEX-ISITE initiative 16- IDEX-0001 (CAP 20–25) (Emergence, Challenge 3-recherche) (to F. C. and P. A.), the Ligue Contre le Cancer comité de l’Ardéche, du Puy de Dôme and du Cantal (to F. C. and P. A.) and “Conseil Régional d’Auvergne” (to PA). B. M. had a fellowship from Fondation pour la recherche médicale (FRM). C.J.M. is funded by the Portuguese Foundation for Science and Technology (FCT-CEECIND/00371/2017).
Author information
Authors and Affiliations
Contributions
PA initiated and supervised the study. BM, FC and PA designed the study. BM, JC, TB, AC, SMM, DN, CGG, SC, NA, CCH, CC, IV, CJM, FC and PA performed the experiments. FC performed the bioinformatics analyses. BM, JC, AC, DN, CGG, IV, JM, FC and PA analyzed the data. BM and FC produced the figures with JC’s and IV’s input. PA wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
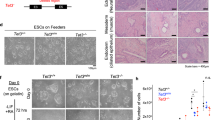
Supp Figure 1: Characterization of the Kdm6b locus. a
Pou5f1, Nestin and Pax6 expression levels in ES cells (n=6) and in NP cells at day 12 (n=6) of in vitro corticogenesis. b Kdm6b expression level in ES cells (n=4) and in NP cells (n=4) at day 12 of in vitro corticogenesis. Expression was assessed with primers located in exon 11 (left panel) and with primers between exon 21 and 23 (right panel). c Race-PCR mapping along the Kdm6b locus in ES cells, neonate brain, and at D8 of in vitro corticogenesis. d CGI1 and CGI2 bisulphite-based DNA methylation analyses by COBRA and/or sequencing in ES and NP cells. For each sequenced region, the methylation patterns are symbolized by lollipops (black: methylated CpG; white: unmethylated CpG). In a and b, statistical significance was determined with the unpaired t-test (p values in the figure). Data are presented as the mean ± SEM (TIF 2047 kb)
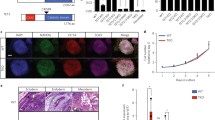
Supp Figure 2: Genomic features of the Kdm6b locus. a
Position of the Kdm6b locus relative to the topological associated domains (TAD), as defined by Dixon et al [34]. b The main genomic features are conserved at the mouse Kdm6b and human KDM6B loci, including the CpG island (CGI3) between exons 17 and 18. c Kdm6bos expression level in ES cells (n=3) and at D8 (n=3) and D12 (NP stage; n=3) of in vitro corticogenesis. Statistical significance was determined by one-way ANOVA (comparison of each differentiation point with ES cells). Data are presented as the mean ± SEM. There was no significant difference between ES and the differentiation points. d Data-mining analysis of H3K4me1, H3K27ac, and H3K4me3 at the Kdm6b locus in ES cells, whole brain, and NP cells. The GRO-seq signals obtained from ES cell are also shown. Signal are shown for Kdm6b sense (upper panel) and antisense (lower panel) transcription (TIF 3262 kb)
Rights and permissions
About this article
Cite this article
Montibus, B., Cercy, J., Bouschet, T. et al. TET3 controls the expression of the H3K27me3 demethylase Kdm6b during neural commitment. Cell. Mol. Life Sci. 78, 757–768 (2021). https://doi.org/10.1007/s00018-020-03541-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03541-8