Abstract
Misfolded F508del-CFTR, the main molecular cause of the recessive disorder cystic fibrosis, is recognized by the endoplasmic reticulum (ER) quality control (ERQC) resulting in its retention and early degradation. The ERQC mechanisms rely mainly on molecular chaperones and on sorting motifs, whose presence and exposure determine CFTR retention or exit through the secretory pathway. Arginine-framed tripeptides (AFTs) are ER retention motifs shown to modulate CFTR retention. However, the interactions and regulatory pathways involved in this process are still largely unknown. Here, we used proteomic interaction profiling and global bioinformatic analysis to identify factors that interact differentially with F508del-CFTR and F508del-CFTR without AFTs (F508del-4RK-CFTR) as putative regulators of this specific ERQC checkpoint. Using LC–MS/MS, we identified kinesin family member C1 (KIFC1) as a stronger interactor with F508del-CFTR versus F508del-4RK-CFTR. We further validated this interaction showing that decreasing KIFC1 levels or activity stabilizes the immature form of F508del-CFTR by reducing its degradation. We conclude that the current approach is able to identify novel putative therapeutic targets that can be ultimately used to the benefit of CF patients.





Similar content being viewed by others
Notes
R program, https://www.r-project.org/ (Last accessed March 09, 2018).
GSEA database, http://software.broadinstitute.org/gsea/index.jsp (Last accessed February 18, 2017).
DAVID database, https://david.ncifcrf.gov/ (Last accessed March 08, 2018).
Reactome Pathway database, https://reactome.org/ (Last accessed September 30, 2017).
APID database, http://cicblade.dep.usal.es:8080/APID/init.action (Last accessed January 26, 2018).
Cytoscape, http://cytoscape.org (Last accessed January 26, 2018).
Harvard PrimerBank database, https://pga.mgh.harvard.edu/primerbank/ (Last accessed November 06, 2017).
Abbreviations
- ABC:
-
ATP-binding cassette
- AFTs:
-
Arginine-framed tripeptides
- APID:
-
Agile protein interactome dataserved
- ATP:
-
Adenosine triphosphate
- BHK:
-
Baby hamster kidney
- BP:
-
Biologic process
- cAMP:
-
cyclic adenosine monophosphate
- CC:
-
Cellular compartment
- CFBE41o-:
-
Cystic fibrosis bronchial epithelial cells
- CF:
-
Cystic fibrosis
- CFTR:
-
Cystic fibrosis transmembrane conductance regulator
- DAVID:
-
Database for annotation: visualization and integrated discovery
- DMP:
-
Dimethyl pimelimidate.2 HCl
- DSP:
-
Dithiobis(succinimidylpropinate)
- EMT:
-
Epithelial-to-mesenchymal transition
- ENaC:
-
Epithelial Na+ channel
- ER:
-
Endoplasmic reticulum
- ES:
-
Enrichment score
- ERQC:
-
Endoplasmic reticulum quality control
- GSEA:
-
Gene set enrichment analysis
- GO:
-
Gene ontology
- HDACi:
-
Histone deacetylase inhibitor
- HEK293:
-
Human embryonic kidney 293
- iBAQ:
-
Intensity-based absolute quantification
- MSD:
-
Membrane-spanning domain
- NBD:
-
Nucleotide-binding domain
- NT:
-
Non-treated
- PM:
-
Plasma membrane
- PPIs:
-
Protein–protein interactions
- QC:
-
Quality control
- RD:
-
Regulatory domain
- SB:
-
Sample buffer
- SEM:
-
Standard error of the mean
- SUMO:
-
Small ubiquitin-like modifiers
- TM:
-
Transmembrane segment
- UPR:
-
Unfolded protein response
- WCL:
-
Whole-cell lysate
- Wt:
-
Wild-type
- 16HBE14o-:
-
Human bronchial epithelial cells
References
Collins FS (1992) Cystic fibrosis: molecular biology and therapeutic implications. Science 256:774–779
Zhang Z, Liu F, Chen J (2017) Conformational changes of CFTR upon phosphorylation and ATP binding. Cell 170(483–491):e8. https://doi.org/10.1016/j.cell.2017.06.041
Farinha CM, Canato S (2017) From the endoplasmic reticulum to the plasma membrane: mechanisms of CFTR folding and trafficking. Cell Mol Life Sci 74:39–55. https://doi.org/10.1007/s00018-016-2387-7
Zerangue N, Schwappach B, Jan YN, Jan LY (1999) A new ER trafficking signal regulates the subunit stoichiometry of plasma membrane KATP channels. Neuron 22:537–548. https://doi.org/10.1016/S0896-6273(00)80708-4
Chang XB, Cui L, Hou YX et al (1999) Removal of multiple arginine-framed trafficking signals overcomes misprocessing of delta F508 CFTR present in most patients with cystic fibrosis. Mol Cell 4:137–142
Roxo-Rosa M, Xu Z, Schmidt A et al (2006) Revertant mutants G550E and 4RK rescue cystic fibrosis mutants in the first nucleotide-binding domain of CFTR by different mechanisms. Proc Natl Acad Sci USA 103:17891–17896. https://doi.org/10.1073/pnas.0608312103
Clarke LA, Sousa L, Barreto C, Amaral MD (2013) Changes in transcriptome of native nasal epithelium expressing F508del-CFTR and intersecting data from comparable studies. Respir Res 14:38. https://doi.org/10.1186/1465-9921-14-38
Rauniyar N, Gupta V, Balch WE, Yates JR (2014) Quantitative proteomic profiling reveals differentially regulated proteins in cystic fibrosis cells. J Proteome Res 13:4668–4675. https://doi.org/10.1021/pr500370g
Roxo-Rosa M, da Costa G, Luider TM et al (2006) Proteomic analysis of nasal cells from cystic fibrosis patients and non-cystic fibrosis control individuals: search for novel biomarkers of cystic fibrosis lung disease. Proteomics 6:2314–2325. https://doi.org/10.1002/pmic.200500273
Wang X, Venable J, LaPointe P et al (2006) Hsp90 cochaperone Aha1 downregulation rescues misfolding of CFTR in cystic fibrosis. Cell 127:803–815. https://doi.org/10.1016/j.cell.2006.09.043
Gomes-Alves P, Couto F, Pesquita C et al (2010) Rescue of F508del-CFTR by RXR motif inactivation triggers proteome modulation associated with the unfolded protein response. Biochim Biophys Acta 1804:856–865. https://doi.org/10.1016/j.bbapap.2009.12.013
Pankow S, Bamberger C, Calzolari D et al (2015) ∆F508 CFTR interactome remodelling promotes rescue of cystic fibrosis. Nature 528:510–516. https://doi.org/10.1038/nature15729
Reilly R, Mroz MS, Dempsey E et al (2017) Targeting the PI3 K/Akt/mTOR signalling pathway in cystic fibrosis. Sci Rep 7:7642. https://doi.org/10.1038/s41598-017-06588-z
Carvalho AS, Ribeiro H, Voabil P et al (2014) Global mass spectrometry and transcriptomics array based drug profiling provides novel insight into glucosamine induced endoplasmic reticulum stress. Mol Cell Proteom 13:3294–3307. https://doi.org/10.1074/mcp.M113.034363
Schwanhäusser B, Busse D, Li N et al (2011) Global quantification of mammalian gene expression control. Nature 473:337–342. https://doi.org/10.1038/nature10098
Arike L, Valgepea K, Peil L et al (2012) Comparison and applications of label-free absolute proteome quantification methods on Escherichia coli. J Proteom 75:5437–5448. https://doi.org/10.1016/j.jprot.2012.06.020
Fabre B, Lambour T, Bouyssié D et al (2014) Comparison of label-free quantification methods for the determination of protein complexes subunits stoichiometry. EuPA Open Proteom 4:82–86. https://doi.org/10.1016/j.euprot.2014.06.001
Matthiesen R (2007) Mass spectrometry data analysis in proteomics. Humana Press, Totowa
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26:1367–1372. https://doi.org/10.1038/nbt.1511
Matthiesen R, Prieto G, Amorim A et al (2012) SIR: deterministic protein inference from peptides assigned to MS data. J Proteom 75:4176–4183. https://doi.org/10.1016/j.jprot.2012.05.010
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. PNAS 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Mootha VK, Lindgren CM, Eriksson K-F et al (2003) PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273. https://doi.org/10.1038/ng1180
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37:1–13. https://doi.org/10.1093/nar/gkn923
Fabregat A, Jupe S, Matthews L et al (2018) The reactome pathway knowledgebase. Nucleic Acids Res 46:D649–D655. https://doi.org/10.1093/nar/gkx1132
Croft D, Mundo AF, Haw R et al (2014) The reactome pathway knowledgebase. Nucleic Acids Res 42:D472–D477. https://doi.org/10.1093/nar/gkt1102
Yu G, He Q-Y (2016) ReactomePA: an R/bioconductor package for reactome pathway analysis and visualization. Mol BioSyst 12:477–479. https://doi.org/10.1039/c5mb00663e
Alonso-López D, Gutiérrez MA, Lopes KP et al (2016) APID interactomes: providing proteome-based interactomes with controlled quality for multiple species and derived networks. Nucleic Acids Res 44:W529–W535. https://doi.org/10.1093/nar/gkw363
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Spandidos A, Wang X, Wang H, Seed B (2010) PrimerBank: a resource of human and mouse PCR primer pairs for gene expression detection and quantification. Nucleic Acids Res 38:D792–D799. https://doi.org/10.1093/nar/gkp1005
Mendes AI, Matos P, Moniz S et al (2011) Antagonistic regulation of cystic fibrosis transmembrane conductance regulator cell surface expression by protein kinases WNK4 and spleen tyrosine kinase. Mol Cell Biol 31:4076–4086. https://doi.org/10.1128/MCB.05152-11
Lobo MJ, Amaral MD, Zaccolo M, Farinha CM (2016) EPAC1 activation by cAMP stabilizes CFTR at the membrane by promoting its interaction with NHERF1. J Cell Sci 129:2599–2612. https://doi.org/10.1242/jcs.185629
Farinha CM, King-Underwood J, Sousa M et al (2013) Revertants, low temperature, and correctors reveal the mechanism of F508del-CFTR rescue by VX-809 and suggest multiple agents for full correction. Chem Biol 20:943–955. https://doi.org/10.1016/j.chembiol.2013.06.004
Simpson JC, Joggerst B, Laketa V et al (2012) Genome-wide RNAi screening identifies human proteins with a regulatory function in the early secretory pathway. Nat Cell Biol 14:764–774. https://doi.org/10.1038/ncb2510
Mellacheruvu D, Wright Z, Couzens AL et al (2013) The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat Methods 10:730–736. https://doi.org/10.1038/nmeth.2557
Gaulton A, Hersey A, Nowotka M et al (2017) The ChEMBL database in 2017. Nucleic Acids Res 45:D945–D954. https://doi.org/10.1093/nar/gkw1074
Van Goor F, Hadida S, Grootenhuis PDJ et al (2011) Correction of the F508del-CFTR protein processing defect in vitro by the investigational drug VX-809. Proc Natl Acad Sci USA 108:18843–18848. https://doi.org/10.1073/pnas.1105787108
Wu J, Mikule K, Wang W et al (2013) Discovery and mechanistic study of a small molecule inhibitor for motor protein KIFC1. ACS Chem Biol 8:2201–2208. https://doi.org/10.1021/cb400186w
Pannu V, Rida PCG, Ogden A et al (2015) HSET overexpression fuels tumor progression via centrosome clustering-independent mechanisms in breast cancer patients. Oncotarget 6:6076–6091
Mittal K, Choi DH, Klimov S et al (2016) A centrosome clustering protein, KIFC1, predicts aggressive disease course in serous ovarian adenocarcinomas. J Ovarian Res 9:17. https://doi.org/10.1186/s13048-016-0224-0
Tomati V, Pesce E, Caci E et al (2018) High-throughput screening identifies FAU protein as a regulator of mutant cystic fibrosis transmembrane conductance regulator channel. J Biol Chem 293:1203–1217. https://doi.org/10.1074/jbc.M117.816595
Okiyoneda T, Veit G, Sakai R et al (2018) Chaperone-independent peripheral quality control of CFTR by RFFL E3 ligase. Dev Cell 44(694–708):e7. https://doi.org/10.1016/j.devcel.2018.02.001
Hirokawa N, Noda Y, Tanaka Y, Niwa S (2009) Kinesin superfamily motor proteins and intracellular transport. Nat Rev Mol Cell Biol 10:682–696. https://doi.org/10.1038/nrm2774
Ahner A, Gong X, Schmidt BZ et al (2013) Small heat shock proteins target mutant cystic fibrosis transmembrane conductance regulator for degradation via a small ubiquitin-like modifier-dependent pathway. Mol Biol Cell 24:74–84. https://doi.org/10.1091/mbc.E12-09-0678
Jensen TJ, Loo MA, Pind S et al (1995) Multiple proteolytic systems, including the proteasome, contribute to CFTR processing. Cell 83:129–135
Ward CL, Omura S, Kopito RR (1995) Degradation of CFTR by the ubiquitin-proteasome pathway. Cell 83:121–127. https://doi.org/10.1016/0092-8674(95)90240-6
Kwon M, Godinho SA, Chandhok NS et al (2008) Mechanisms to suppress multipolar divisions in cancer cells with extra centrosomes. Genes Dev 22:2189–2203. https://doi.org/10.1101/gad.1700908
Mountain V, Simerly C, Howard L et al (1999) The kinesin-related protein, HSET, opposes the activity of Eg5 and cross-links microtubules in the mammalian mitotic spindle. J Cell Biol 147:351–366
Zhang JT, Jiang XH, Xie C et al (2013) Downregulation of CFTR promotes epithelial-to-mesenchymal transition and is associated with poor prognosis of breast cancer. Biochimica et Biophysica Acta (BBA) Mol Cell Res 1833:2961–2969. https://doi.org/10.1016/j.bbamcr.2013.07.021
Maisonneuve P, Marshall BC, Knapp EA, Lowenfels AB (2013) Cancer risk in cystic fibrosis: a 20-year nationwide study from the United States. J Natl Cancer Inst 105:122–129. https://doi.org/10.1093/jnci/djs481
Xiao Y-X, Shen H-Q, She Z-Y et al (2017) C-terminal kinesin motor KIFC1 participates in facilitating proper cell division of human seminoma. Oncotarget 8:61373–61384. https://doi.org/10.18632/oncotarget.18139
Mukhopadhyay A, Quiroz JA, Wolkoff AW (2014) Rab1a regulates sorting of early endocytic vesicles. Am J Physiol Gastrointest Liver Physiol 306:G412–G424. https://doi.org/10.1152/ajpgi.00118.2013
Acknowledgements
Work supported by centre Grant UID/MULTI/04046/2013 to BioISI, Romain Pauwels Research Award to C.M.F, and Project MIMED PTDC/EEI-ESS/4923/2014 to A.O.F. SC and JDS are recipient of fellowships from BioSys PhD programme PD/BD/114393/2016 (SFRH/BD/52491/2014) and PD/BD/106084/2015 (SFRH(BD/106084/2015) from FCT, Portugal, respectively. Proteomics Core Facility-SGIKER is part of ProteoRed-ISCIII network and is partially funded by ERDF and ESF.
Author information
Authors and Affiliations
Contributions
SC designed and performed the experiments, analysed data and wrote the manuscript; JDS analysed data; ASC prepared the sample and performed mass spectrometry experiment; KA and RM performed the mass spectrometry experiment. MDA provided advice and comments on the manuscript; AOF guided the experiments design for bioinformatics analysis, secured funding and provided advice; CMF guided the project, secured funding, guided the experiments design and wrote the manuscript. All authors read, revised and approved the manuscript.
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
- Characterization of CFBE cells stably expressing F508del-4RK-CFTR. CFBE cells expressing F508del-4RK-CFTR (lane 1-6), wt-CFTR (lane 7) and F508del-CFTR (lane 8) were grown at 37 °C or 26 °C and in some cases incubated with VX-809 (3 µM) for 24 h or DMSO (vehicle compound). Western blot (top) showing both mature form band C (~ 170 kDa) and immature form band B (~ 140 kDa) of CFTR protein. Calnexin (90 kDa) was used as a loading control (bottom). (JPEG 66 kb)
Fig. S2 - Bioinformatics analysis of the proteins differentially interacting with F508del-CFTR and F508del-4RK-CFTR. (A)
Hallmark enrichment for F508del-4RK-CFTR and F508del-CFTR using gene set enrichment (GSEA). GSEA was performed for a dataset of 834 proteins. Using GSEA tool, the subset of genes that contribute mostly to the enrichment of each hallmark was identified and plotted according to their affinity to F508del-4RK-CFTR (black dots) versus F508del-CFTR (grey dots) - represented by log2 of the ratio F508del-4RK-CFTR/F508del-CFTR. (B) Biological process represented for F508del-4RK-CFTR and F508del-CFTR using DAVID. From the total of 834 proteins, 198 with more affinity to F508del-CFTR and 164 proteins with more affinity to F508del-4RK-CFTR were used to find the GO terms – biological process enriched in both subsets. p value < 0.05. (C) Cellular component represented for F508del-4RK-CFTR and F508del-CFTR using DAVID. From the total of 834 proteins, 198 with more affinity to F508del-CFTR and 164 proteins with more affinity to F508del-4RK-CFTR were used to find the GO terms – cellular compartment enriched in both subsets. Significant levels are represented by p value < 0.05. (PDF 371 kb)
Fig. S3 - Scatter plot representing the differential protein interactions for the 22 putative hits in F508del-CFTR versus F508del-4RK-CFTR and peptides corresponding to the AFT regions of CFTR.
Log2 plot of the protein abundance ratio for F508del-4RK-CFTR versus F508del-CFTR and Log2 plot of the protein abundance ratio for peptide K (corresponding to mutated AFTs with Lys replacement) versus peptide R (Arg containing). Proteins were identified by LC–MS/MS and the threshold (log2 = ±1). (JPEG 113 kb)
Fig. S4
- Force-directed network of CFTR versus top hits interactome. Network comprises the connection of CFTR interactome (depicted in green circle) and the interactome of the selected hits: FKBP4, HNRNPA2B1, KIFC1, YWHAE (depicted in orange circles). All components comprising the interactors are predicted as nodes (yellow circles) and all were identified by LS-MS/MS covered the 834 proteins. Straight white lines illustrate edges that define interactions based on the APID protein interaction database which was accessed using Cytoscape platform. Dashed red lines illustrate the edges in which interactions occur with one node distance and full orange lines the edges in which interactions occur with two or more nodes distance to CFTR. (JPEG 149 kb)
Fig. S5 - Effect of FKBP4, 14-3-3ε and HNRNPA2B1 knockdown on F508del-CFTR protein processing.
CFBE cells expressing F508del-CFTR were transfected with siRNA against FKBP4, 14-3-3ε, HNRNPA2B1 or EGFP as non-targeting siRNA for 48 h. CT - transfection reagent only was used. Cells were also incubated with VX-809 (3 µM) or DMSO (vehicle control) are shown. Detection of F508del-CFTR protein expression (top) under (A) FKBP4, (B) 14-3-3ε or (C) HNRNPA2B1 knockdown. Equal amount of protein was loaded in each lane, as demonstrated by calnexin loading control (Bottom). (D) CFTR processing (band C divided by total CFTR) was normalized to siEGFP control. (E) Gene knockdown with siRNA in CFBE cells. Fold expression of FKBP4, 14-3-3ε or HNRNPA2B1 mRNA levels was obtained by relative quantification (ddCT method) and was normalized to an internal control (CAP-1). Data are shown as the mean ± SEM, n = 3. * p ≤ 0.05. Statistical analysis was performed using two-tailed unpaired Student’s t test. (JPEG 331 kb)
Fig. S6
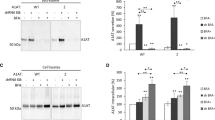
- Effect of KIFC1 inhibition on F508del-CFTR expression. (A) CFBE cells expressing F508del-CFTR were incubated with 0.4 µM KIFC1 inhibitor (AZ82 compound) or DMSO, vehicle control (non-treated - NT), for 24 h, 48 h or 72 h. As a control for KIFC1 levels, cells were transfected with siRNA pool for KIFC1 or EGFP as non-targeting siRNA for 48 h. Western blot for CFTR (Top). Equal amount of protein was loaded in each lane, as demonstrated by α-tubulin (~ 50 kDa) detection (Bottom). (B) Quantification of CFTR (band B and C) band intensity normalized to loading control and to NT or siEGFP. (C) CFTR processing (band C divided by total CFTR) normalized to NT or siEGFP. Data are shown as mean ± SEM, n = 3. * ρ ≤ 0.05. Statistical analysis was performed using two-tailed unpaired Student’s t test. (JPEG 144 kb)
Fig. S7 - Distance of F508del-CFTR interactors to KIFC1.
The 198 interactors with higher affinity to F508del-CFTR were subjected to APID1 (accounting for all known interactions) and APID2 (accounting for interactions proved by two or more experiments). Each circle represents the distance from the proteins to KIFC1 (from one to five edges). Inside of each circle is represented the number of proteins showing the associated distance. (JPEG 92 kb)
Table S1 - Total interactome identified for F508del-CFTR and F508del-4RK-CFTR.
All identified proteins from MS analysis. (XLSX 156 kb)
Table S2 - Protein fold change interaction for F508del-CFTR versus F508del-4RK-CFTR.
Fold change interaction was obtained by log2 of the ratio of the amount of the protein detected in association with F508del-CFTR versus F508del-4RK-CFTR. (XLSX 94 kb)
Table S3 - Protein targets and inhibitors listed in ChEMBL database.
The 198 proteins with higher affinity for F508del-CFTR (log2 below -1) were searched in ChEMBL. The interactors with available inhibitors are listed along the fold change score (regarding F508del- over F508del-4RK-CFTR interaction). (XLSX 27 kb)
Table S4 - APID interactions linking CFTR and KIFC1.
The 198 proteins with stronger affinity with F508del-CFTR were analysed in APID1 and APID2 to identify their possible link to KIFC1, one to five represent the number of edges. (XLSX 30 kb)
Rights and permissions
About this article
Cite this article
Canato, S., Santos, J.D., Carvalho, A.S. et al. Proteomic interaction profiling reveals KIFC1 as a factor involved in early targeting of F508del-CFTR to degradation. Cell. Mol. Life Sci. 75, 4495–4509 (2018). https://doi.org/10.1007/s00018-018-2896-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-018-2896-7