Abstract
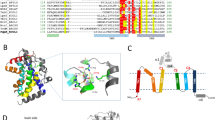
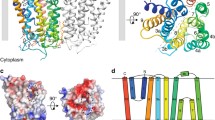
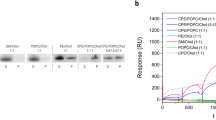
Type 2 phosphatidic acid phosphatases (PAP2s) can be either soluble or integral membrane enzymes. In bacteria, integral membrane PAP2s play major roles in the metabolisms of glycerophospholipids, undecaprenyl-phosphate (C55-P) lipid carrier and lipopolysaccharides. By in vivo functional experiments and biochemical characterization we show that the membrane PAP2 coded by the Bacillus subtilis yodM gene is the principal phosphatidylglycerol phosphate (PGP) phosphatase of B. subtilis. We also confirm that this enzyme, renamed bsPgpB, has a weaker activity on C55-PP. Moreover, we solved the crystal structure of bsPgpB at 2.25 Å resolution, with tungstate (a phosphate analog) in the active site. The structure reveals two lipid chains in the active site vicinity, allowing for PGP substrate modeling and molecular dynamic simulation. Site-directed mutagenesis confirmed the residues important for substrate specificity, providing a basis for predicting the lipids preferentially dephosphorylated by membrane PAP2s.




Similar content being viewed by others
References
Sigal YJ, McDermott MI, Morris AJ (2005) Integral membrane lipid phosphatases/phosphotransferases: common structure and diverse functions. Biochem J 387:281–293. doi:10.1042/BJ20041771
Manat G, Roure S, Auger R et al (2014) Deciphering the metabolism of undecaprenyl-phosphate: the bacterial cell-wall unit carrier at the membrane frontier. Microb Drug Resist 20:199–214. doi:10.1089/mdr.2014.0035
Lei KJ, Shelly LL, Pan CJ et al (1993) Mutations in the glucose-6-phosphatase gene that cause glycogen storage disease type 1a. Science 262:580–583
Cullen TW, Giles DK, Wolf LN et al (2011) Helicobacter pylori versus the host: remodeling of the bacterial outer membrane is required for survival in the gastric mucosa. PLoS Pathog 7:e1002454. doi:10.1371/journal.ppat.1002454
Fournier J-B, Rebuffet E, Delage L et al (2014) The Vanadium Iodoperoxidase from the marine flavobacteriaceae species Zobellia galactanivorans reveals novel molecular and evolutionary features of halide specificity in the vanadium haloperoxidase enzyme family. Appl Environ Microbiol 80:7561–7573. doi:10.1128/AEM.02430-14
Comba S, Menendez-Bravo S, Arabolaza A, Gramajo H (2013) Identification and physiological characterization of phosphatidic acid phosphatase enzymes involved in triacylglycerol biosynthesis in Streptomyces coelicolor. Microb Cell Fact 12:9. doi:10.1186/1475-2859-12-9
Touzé T, Tran AX, Hankins JV et al (2008) Periplasmic phosphorylation of lipid A is linked to the synthesis of undecaprenyl phosphate. Mol Microbiol 67:264–277. doi:10.1111/j.1365-2958.2007.06044.x
Chae M, Carman GM (2013) Characterization of the yeast actin patch protein app1p phosphatidate phosphatase. J Biol Chem 288:6427–6437. doi:10.1074/jbc.M112.449629
Peschel A (2002) How do bacteria resist human antimicrobial peptides? Trends Microbiol 10:179–186. doi:10.1016/S0966-842X(02)02333-8
Nishi H, Komatsuzawa H, Fujiwara T et al (2004) Reduced content of lysyl-phosphatidylglycerol in the cytoplasmic membrane affects susceptibility to moenomycin, as well as vancomycin, gentamicin, and antimicrobial peptides, in Staphylococcus aureus. Antimicrob Agents Chemother 48:4800–4807. doi:10.1128/AAC.48.12.4800-4807.2004
Kuhn S, Slavetinsky CJ, Peschel A (2015) Synthesis and function of phospholipids in Staphylococcus aureus. Int J Med Microbiol 305:196–202. doi:10.1016/j.ijmm.2014.12.016
Raetz CR (1986) Molecular genetics of membrane phospholipid synthesis. Annu Rev Genet 20:253–295. doi:10.1146/annurev.ge.20.120186.001345
Ratledge C, Wilkinson SG (1988) Microbial lipids vol. 1. Academic Presss, London
Goldberg DE, Rumley MK, Kennedy EP (1981) Biosynthesis of membrane-derived oligosaccharides: a periplasmic phosphoglyceroltransferase. Proc Natl Acad Sci USA 78:5513–5517
Koch HU, Haas R, Fischer W (1984) The role of lipoteichoic acid biosynthesis in membrane lipid metabolism of growing Staphylococcus aureus. Eur J Biochem 138:357–363
Icho T, Raetz CRH (1983) Multiple genes for membrane-bound phosphatases in Escherichia coli and their action on phospholipid precursors. J Bacteriol 153:722–730
Lu Y-H, Guan Z, Zhao J, Raetz CRH (2011) Three phosphatidylglycerol-phosphate phosphatases in the inner membrane of Escherichia coli. J Biol Chem 286:5506–5518. doi:10.1074/jbc.M110.199265
Cantagrel V, Lefeber DJ (2011) From glycosylation disorders to dolichol biosynthesis defects: a new class of metabolic diseases. J Inherit Metab Dis 34:859–867. doi:10.1007/s10545-011-9301-0
Apfel CM, Takács B, Fountoulakis M et al (1999) Use of genomics to identify bacterial undecaprenyl pyrophosphate synthetase: cloning, expression, and characterization of the essential uppS gene. J Bacteriol 181:483–492
Siewert G, SJ (1967) Bacitracin: an inhibitor of the dephosphorylation of lipid pyrophosphate, an intermediate in the biosynthesis of the peptidoglycan of bacterial cell wall. Proc Natl Acad Sci USA 57:767–773
El Ghachi M, Bouhss A, Blanot D, Mengin-Lecreulx D (2004) The bacA gene of Escherichia coli encodes an undecaprenyl pyrophosphate phosphatase activity. J Biol Chem 279:30106–30113. doi:10.1074/jbc.M401701200
El Ghachi M, Derbise A, Bouhss A, Mengin-Lecreulx D (2005) Identification of multiple genes encoding membrane proteins with undecaprenyl pyrophosphate phosphatase (UppP) activity in Escherichia coli. J Biol Chem 280:18689–18695. doi:10.1074/jbc.M412277200
Bernard R, El Ghachi M, Mengin-Lecreulx D et al (2005) BcrC from Bacillus subtilis acts as an undecaprenyl pyrophosphate phosphatase in bacitracin resistance. J Biol Chem 280:28852–28857. doi:10.1074/jbc.M413750200
Zhao H, Sun Y, Peters JM et al (2016) Depletion of undecaprenyl pyrophosphate phosphatases (UPP-Pases) disrupts cell envelope biogenesis in Bacillus subtilis. J Bacteriol. doi:10.1128/JB.00507-16
Fan J, Jiang D, Zhao Y et al (2014) Crystal structure of lipid phosphatase Escherichia coli phosphatidylglycerophosphate phosphatase B. Proc Natl Acad Sci USA 111:7636–7640. doi:10.1073/pnas.1403097111
Tong S, Lin Y, Lu S et al (2016) Structural insight into substrate selection and catalysis of lipid phosphate phosphatase PgpB in the cell membrane. J Biol Chem. doi:10.1074/jbc.M116.737874
Isupov MN, Dalby AR, Brindley AA et al (2000) Crystal structure of dodecameric vanadium-dependent bromoperoxidase from the red algae Corallina officinalis. J Mol Biol 299:1035–1049. doi:10.1006/jmbi.2000.3806
Makde RD, Mahajan SK, Kumar V (2007) Structure and mutational analysis of the PhoN protein of Salmonella typhimurium provide insight into mechanistic details. BioChemistry 46:2079–2090. doi:10.1021/bi062180g
Bernard R, Joseph P, Guiseppi A et al (2003) YtsCD and YwoA, two independent systems that confer bacitracin resistance to Bacillus subtilis. FEMS Microbiol Lett 228:93–97
Barreteau H, Magnet S, Ghachi M El et al (2009) Quantitative high-performance liquid chromatography analysis of the pool levels of undecaprenyl phosphate and its derivatives in bacterial membranes. J Chromatogr B Anal Technol Biomed Life Sci 877:213–220. doi:10.1016/j.jchromb.2008.12.010
Rost B, Yachdav G, Liu J (2004) The PredictProtein server. Nucleic Acids Res 32:W321–W326. doi:10.1093/nar/gkh377
Touzé T, Blanot D, Mengin-Lecreulx D (2008) Substrate specificity and membrane topology of Escherichia coli PgpB, an undecaprenyl pyrophosphate phosphatase. J Biol Chem 283:16573–16583. doi:10.1074/jbc.M800394200
Tatar LD, Marolda CL, Polischuk AN et al (2007) An Escherichia coli undecaprenyl-pyrophosphate phosphatase implicated in undecaprenyl phosphate recycling. Microbiology 153:2518–2529. doi:10.1099/mic.0.2007/006312-0
Pompeo F, Van Heijenoort J, Mengin-Lecreulx D (1998) Probing the role of cysteine residues in glucosamine-1-phosphate acetyltransferase activity of the bifunctional glmU protein from Escherichia coli: Site-directed mutagenesis and characterization of the mutant enzymes. J Bacteriol 180:4799–4803
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917. doi:10.1139/o59-099
Mascher T, Margulis NG, Wang T et al (2003) Cell wall stress responses in Bacillus subtilis: the regulatory network of the bacitracin stimulon. Mol Microbiol 50:1591–1604. doi:10.1046/j.1365-2958.2003.03786.x
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645. doi:10.1073/pnas.120163297
Hamilton CM, Aldea M, Washburn BK et al (1989) New method for generating deletions and gene replacements in Escherichia coli. J Bacteriol 171:4617–4622
Baba T, Ara T, Hasegawa M et al (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2(2006):0008. doi:10.1038/msb4100050
Caffrey M, Cherezov V (2009) Crystallizing membrane proteins using lipidic mesophases. Nat Protoc 4:706–731. doi:10.1038/nprot.2009.31
Cheng A, Hummel B, Qiu H, Caffrey M (1998) A simple mechanical mixer for small viscous lipid-containing samples. Chem Phys Lipids 95:11–21. doi:10.1016/S0009-3084(98)00060-7
Li D, Boland C, Walsh K, Caffrey M (2012) Use of a robot for high-throughput crystallization of membrane proteins in lipidic mesophases. J Vis Exp e4000. doi:10.3791/4000
Li D, Boland C, Aragao D, et al (2012) Harvesting and cryo-cooling crystals of membrane proteins grown in lipidic mesophases for structure determination by macromolecular crystallography. J Vis Exp e4001. doi:10.3791/4001
Kabsch W (2010) XDS. Acta Crystallogr Sect D Biol Crystallogr 66:125–132. doi:10.1107/S0907444909047337
Terwilliger TC, Adams PD, Read RJ et al (2009) Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr Sect D Biol Crystallogr 65:582–601. doi:10.1107/S0907444909012098
Afonine P V., Grosse-Kunstleve RW, Echols N et al (2012) Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr Sect D Biol Crystallogr 68:352–367. doi:10.1107/S0907444912001308
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr Sect D Biol Crystallogr 66:486–501. doi:10.1107/S0907444910007493
Afonine PV, Moriarty NW, Mustyakimov M, et al (2015) FEM: feature-enhanced map. Acta Crystallogr Sect D Biol Crystallogr 71:646–666. doi:10.1107/S1399004714028132
Krieger E, Darden T, Nabuurs SB et al (2004) Making optimal use of empirical energy functions: force-field parameterization in crystal space. Proteins Struct Funct Genet 57:678–683. doi:10.1002/prot.20251
Best RB, Hummer G (2009) Optimized molecular dynamics force fields applied to the helix-coil transition of polypeptides. J Phys Chem B 113:9004–9015. doi:10.1021/jp901540t
Essmann U, Perera L, Berkowitz ML et al (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Acknowledgements
This work was supported by the Belgian program of Interuniversity Attraction Poles initiated by the Federal Office for Scientific Technical and Cultural Affairs (IAP no. P7/44), the FRS-FNRS (MIS F.4518.12, IISN 4.4503.11), the Tournesol/Hubert Curien partnership between Belgium and France (R.CFRA.1567), the Science Foundation Ireland (Grant Number 12/IA/1255), the Agence Nationale de la Recherche (Bactoprenyl project, ANR-11-BSV3-002), the Centre National de la Recherche Scientifique and the University of Paris-Sud (UMR 9198) and the Aix-Marseille University. The assistance and support of beamline scientists at the Advanced Photon Source (23-ID) are acknowledged.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Additional information
Meriem El Ghachi, Nicole Howe, Rodolphe Auger and Alexandre Lambion contributed equally.
Data deposition: coordinates of the bsPgpB crystal structure have been deposited in the Protein Data Bank (PDB id code 5JKI).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ghachi, M.E., Howe, N., Auger, R. et al. Crystal structure and biochemical characterization of the transmembrane PAP2 type phosphatidylglycerol phosphate phosphatase from Bacillus subtilis . Cell. Mol. Life Sci. 74, 2319–2332 (2017). https://doi.org/10.1007/s00018-017-2464-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2464-6